Time-resolved proximity proteomics uncovers a membrane tension-sensitive caveolin-1 interactome at the rear of migrating cells
Curation statements for this article:-
Curated by eLife
eLife assessment
This important study uses convincing time-resolved proximity proteomics, validated with proximity ligation assays, to provide new insight into mechanical regulation of caveolin-1 complexes that form in migrating cells. Solid follow up experiments reveal a reciprocal relationship between mechanosensitive caveolae and RhoGTPase signalling in migrating cells, but evidence supporting a direct link between the newly identified factors with a specific caveolae subpopulation remains incomplete at this stage.
This article has been Reviewed by the following groups
Discuss this preprint
Start a discussion What are Sciety discussions?Listed in
- Evaluated articles (eLife)
Abstract
Caveolae are small membrane pits with fundamental roles in mechanotransduction. Several studies have shown that caveolae flatten out in response to increased membrane tension, thereby acting as a mechanosensitive membrane reservoir that buffers acute mechanical stress. Caveolae have also been implicated in the control of RhoA/ROCK-mediated actomyosin contractility at the rear of migrating cells. However, how membrane tension controls the organisation of caveolae and their role in mechanotransduction remains unclear. To address this, we systematically quantified protein–protein interactions of caveolin-1 in migrating RPE1 cells at steady state and in response to an acute increase in membrane tension using biotin-based proximity labelling and quantitative mass spectrometry. Our data show that caveolae are highly enriched at the rear of migrating RPE1 cells and that membrane tension rapidly and reversibly disrupts the caveolar protein coat. Membrane tension also detaches caveolin-1 from focal adhesion proteins and several mechanosensitive regulators of cortical actin including filamins and cortactin. In addition, we present evidence that ROCK and the RhoGAP ARHGAP29 associate with caveolin-1 in a manner dependent on membrane tension, with ARHGAP29 influencing caveolin-1 Y14 phosphorylation, caveolae rear localisation, and RPE1 cell migration. Taken together, our work uncovers a membrane tension-sensitive coupling between caveolae and the rear-localised F-actin cytoskeleton. This provides a framework for dissecting the molecular mechanisms underlying caveolae-regulated mechanotransduction pathways.
Article activity feed
-

-

Author response:
Reviewer #1 (Public Review):
In this study, Girardello et al. use proteomics to reveal the membrane tension sensitive caveolin-1 interactome in migrating cells. The authors use EM and surface rendering to demonstrate that caveolae formed at the rear of migrating cells are complex membrane-linked multilobed structures, and they devise a robust strategy to identify caveolin-1 associated proteins using APEX2-mediated proximity biotinylation. This important dataset is further validated using proximity ligation assays to confirm key interactions, and follows up with an interrogation of a surprising relationship between caveolae and RhoGTPase signalling, where caveolin-1 recruits ROCK1 under high membrane tension conditions, and ROCK1 activity is required to reform caveolae upon reversion to isotonic solution. However, …
Author response:
Reviewer #1 (Public Review):
In this study, Girardello et al. use proteomics to reveal the membrane tension sensitive caveolin-1 interactome in migrating cells. The authors use EM and surface rendering to demonstrate that caveolae formed at the rear of migrating cells are complex membrane-linked multilobed structures, and they devise a robust strategy to identify caveolin-1 associated proteins using APEX2-mediated proximity biotinylation. This important dataset is further validated using proximity ligation assays to confirm key interactions, and follows up with an interrogation of a surprising relationship between caveolae and RhoGTPase signalling, where caveolin-1 recruits ROCK1 under high membrane tension conditions, and ROCK1 activity is required to reform caveolae upon reversion to isotonic solution. However, caveolin-1 recruits the RhoA inactivator ARHGAP29 when membrane tension is low and ARHGAP29 overexpression leads to disassembly of caveolae and reduced cell motility. This study builds on previous findings linking caveolae to positive feedback regulation of RhoA signalling, and provides further evidence that caveolae serve to drive rear retraction in migration but also possess an intrinsic brake to limit RhoA activation, leading the authors to suggest that cycles of caveolae assembly and disassembly could thereby be central to establish a stable cell rear for persistent cell migration
A major strength of the manuscript is the robust proteomic dataset. The experimental set up is well defined and mostly well controlled, and there is good internal validation in that the high abundance of core caveolar proteins in low membrane tension (isotonic) conditions, and absence under high membrane tension (brief hypo-osmotic shock) conditions, correlating very well with previous finding. The data could however be better presented to show where statically robust changes occur, and supplementary information should include a table of showing abundance. It's very good to see a link to PRIDE, providing a useful resource for the community.
We thank the reviewer for the positive feedback. We have included the outputs from the search engine in Supplementary File 1.
The authors detail several known interactions and their mechanosensitivty, but also report new interactors of caveolin-1. Several mechanosensitive interactions of caveolin-1 take place at the cell rear, but others are more diffuse across the cell looking at the PLA data (e.g FLN1, CTTN, HSPB1; Figure 4A-F and Figure 4 supplement 1). It is interesting to speculate that those at the cell rear are involved in caveolae, whilst others are linked specifically to caveolin-1 (e.g. dolines). PLA or localisation analysis with Cavin1/PTRF may be able to resolve this and further specify caveolae versus non-caveolae mechanosensitive interactions.
We thank the reviewer for this interesting idea. It is true that many if not most proteins we identified to be associated with Cav1 are not restricted to the cell rear. To analyse to what extent the identified proteins interact with Cav1 at the rear we reanalysed our PLA data for some of the antibody combinations we looked at. This new analysis is now shown in Fig 5G. As expected, for Cav1/PTRF and Cav1/EHD2 most PLA dots (70-80%) were found at the rear. This rear bias is also evident from the representative images we show in the Figure panels 5A and 5E. On the contrary, much fewer PLA dots (~40%) were rear-localised for Cav1/CTTN and Cav1/FLNA antibody combinations. This reflects the much broader cellular distribution of these proteins compared to the core caveolae proteins, and might suggest that there are generally few links between caveolae and cortical actin. However, it is also possible that such links/interactions are more difficult to detect using PLA (because of the extended distance between caveolae and the actin cortex, or because of steric constraints).
The Cav1/ARHGAP29 influence on YAP signalling is interesting, but appear to be quite isolated from the rest of the manuscript. Does overexpression of ARHGAP29 influence YAP signalling and/or caveolar protein expression/Cav1pY14?
Our data and published work originally prompted us to speculate that there is a potential functional link between Cav1, YAP, and ARHGAP29. In an attempt to address this we have performed several Western blots on cell lysates from cells overexpressing ARHGAP29. We did not see major changes in Cav1 Y14 phosphorylation levels in cells overexpressing ARHGAP29, and YAP and pYAP levels also remained unchanged (not shown). In addition, based on previous literature 1,2 we expected to see an effect on ARHGAP29 mRNA levels and YAP target gene transcripts in Cav1 siRNA transfected cells. To our surprise, the mRNA levels of three independent YAP target genes and ARHGAP29 were unchanged in Cav1 siRNA treated cells (this is now shown in Figure 6 Figure Supplement 1). Our data therefore suggest that in RPE1 cells, the connection between Cav1 and ARHGAP29 is independent of YAP signalling, and that the increase in ARHGAP29 protein levels observed in Cav1 siRNA cells is due to some unknown post-translational mechanism.
ARHGAP29 and RhoA/ROCK1 related observations are very interesting and potentially really important. However, the link between ARHGAP29 and caveolae is not well established (other than in proteomic data). PLA or FRET could help establish this.
We agree that the physical and functional link between caveolae (or Cav1) and ARHGAP29 was not well worked out in the original manuscript. In an attempt to address this we have performed PLA assays in GFP-ARHGAP29 transfected cells (as we did not find a suitable ARHGAP29 antibody that works reliably in IF) using anti-Cav1 and anti-GFP antibodies. The PLA signal we obtained for Cav1 and ARHGAP29 was not significantly different to control PLA experiments. There was very little PLA signal to start with. This is not surprising given that ARHGAP29 localisation is mostly diffuse in the cytoplasm, whilst Cav1 is concentrated at the rear. In addition, in cases where we do see ARHGAP29 localisation at the cell cortex, Cav1 tends to be absent (this is now shown in Figure 6 – Figure Supplement 2E). In other words, with the tools we have available, we see little colocalization between Cav1 and ARHGAP29 at steady state. Altogether we speculate that ARHGAP29, through its negative effect on RhoA, flattens caveolae at the membrane or interferes with caveolae assembly at these sites.
This of course prompts the question why ARHGAP29 was identified in the Cav1 proteome with such specificity and reproducibility in the first place? This can be explained by the way APEX2 labeling works. Proximity biotinylation with APEX2 is extremely sensitive and restricted to a labelling radius of ~20 nm 3. The labeling reaction is conducted on live and intact cells at room temperature for 1 min. Although 1 min appears short, dynamic cellular processes occur at the time scale of seconds and are ongoing during the labelling reaction. It is conceivable that within this 1 min time frame, ARHGAP29 cycles on and off the rear membrane (kiss and run). This allows ARHGAP29 to be biotinylated by Cav1-APEX2, resulting in its identification by MS. We have included this in the discussion section.
The relationship between ARHGAP29 and RhoA signalling is not well defined. Is GAP activity important in determining the effect on migration and caveolae formation? What is the effect on RhoA activity? Alternatively, the authors could investigate YAP dependent transcriptional regulation downstream of overexpression.
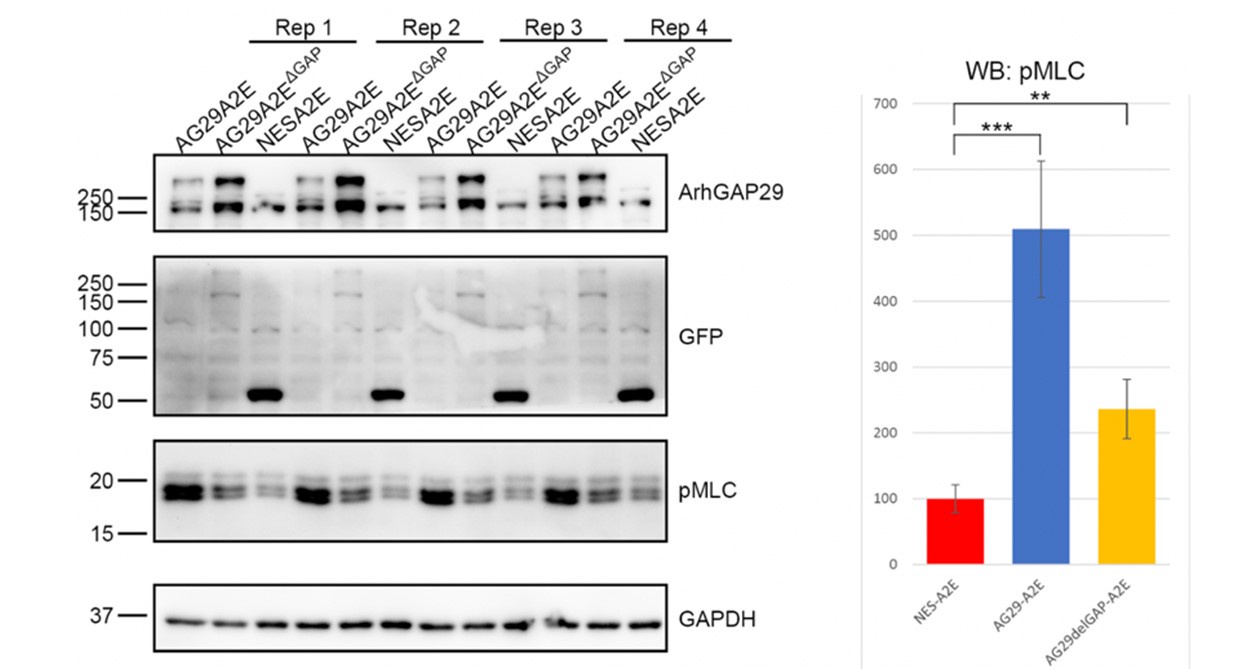
We have addressed this point using overexpression and siRNA transfections. We overexpressed ARHGAP29 or ARHGAP29 lacking its GAP domain and performed WB analysis against pMLC (which is a commonly used and reliable readout for RhoA and myosin-II activity). Much to our surprise, overexpression of ARHGAP29 increased (rather than decreased) pMLC levels, partially in a GAP-dependent manner (see Author response image 1). This is puzzling, as ARHGAP29 is expected to reduce RhoA-GTP levels, which in turn is expected to reduce ROCK activity and hence pMLC levels. In addition, and also surprisingly, siRNA-mediated silencing of ARHGAP29 did not significantly change pMLC levels. By contrast, pMLC levels were strongly reduced in Cav1 siRNA treated cells (this is shown in Fig. 6A and 6B in the revised manuscript). These new data underscore the important role of caveolae in the control of myosin-II activity, but do not allow us to draw any firm conclusions about the role of ARHGAP29 at the cell rear.
Author response image 1.
Overexpression of ARHGAP29 reduces, rather than increases pMLC in RPE1 cells.
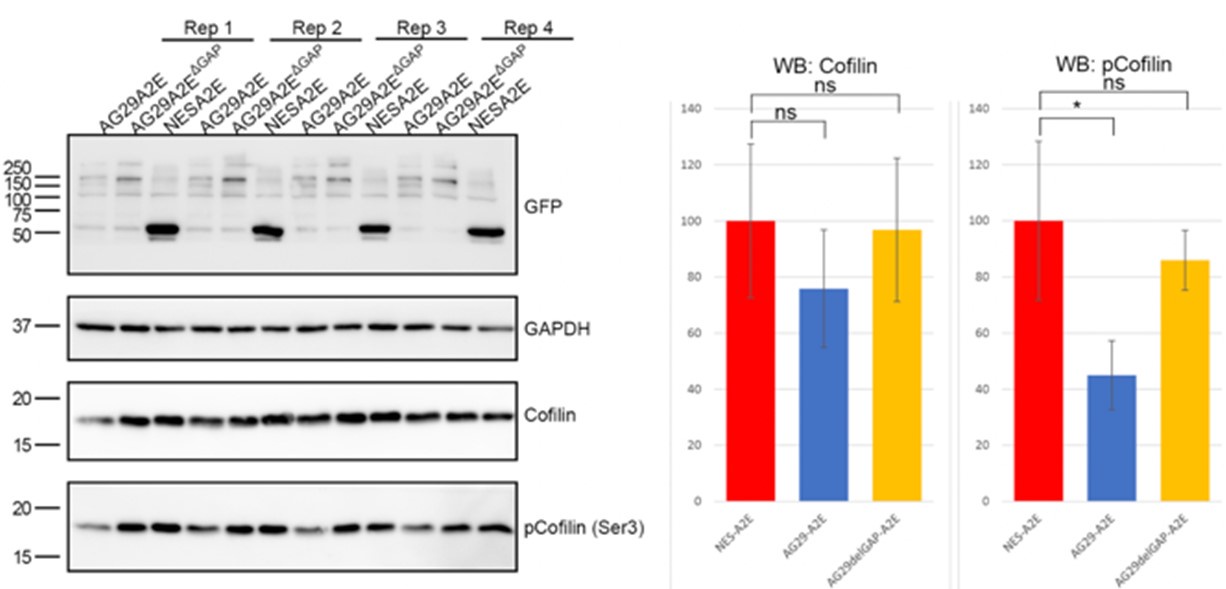
We are uncertain as to how to interpret the ARHGAP29 overexpression data presented in Author response image 1 and therefore decided not to include it in the manuscript. One possibility is that inactivation of RhoA below a certain critical threshold causes other mechanisms to compensate. For instance, the activity of alternative MLC kinases such as MLCK could be enhanced under these conditions. Another possibility is that ARHGAP29 controls MLC phosphorylation indirectly. For instance, it has been shown that ARHGAP29 promotes actin destabilization through inactivating LIMK/cofilin signalling 1. In agreement with this, we find that overexpression of ARHGAP29 reduces p-cofilin (serine 3) levels (see Author response image 2). Since cofilin and MLC crosstalk 4, it is possible that increased pMLC levels are the result of a feedback loop that compensates for the effect of actin depolymerisation. This is now discussed in the discussion section. Whichever the case, we hope the reviewers understand that deeper mechanistic insight into the intricate mechanisms of Rho signalling at the cell rear are beyond the scope of this manuscript.
Author response image 2.
Overexpression of ARHGAP29 reduces p-cofilin levels in RPE1.
Reviewer #2 (Public Review):
Girardello et al investigated the composition of the molecular machinery of caveolae governing their mechano-regulation in migrating cells. Using live cell imaging and RPE1 cells, the authors provide a spatio-temporal analysis of cavin-3 distribution during cell migration and reveal that caveolae are preferentially localized at the rear of the cell in a stable manner. They further characterize these structures using electron tomography and reveal an organization into clusters connected to the cell surface. By performing a proteomic approach, they address the interactome of caveolin-1 proteins upon mechanical stimulation by exposing RPE1 cells to hypo-osmotic shock (which aims to increase cell membrane tension) or not as a control condition. The authors identify over 300 proteins, notably proteins related to actin cytoskeleton and cell adhesion. These results were further validated in cellulo by interrogating protein-protein interactions using proximity ligation assays and hypo-osmotic shock. These experiments confirmed previous data showing that high membrane tension induces caveolae disassembly in a reversible manner. Eventually, based on literature and on the results collected by the proteomic analysis, authors investigated more deeply the molecular signaling pathway controlling caveolae assembly upon mechanical stimuli. First, they confirm the targeting of ROCK1 with Caveolin-1 and the implication of the kinase activity for caveolae formation (at the rear of the cell). Then, they show that RhoGAP ARHGAP29, a factor newly identified by the proteomic analysis, is also implicated in caveolae mechano-regulation likely through YAP protein and found that overexpression of RhoGAP ARHGAP29 affects cell motility. Overall, this paper interrogated the role of membrane tension in caveolae located at the rear of the cell and identified a new pathway controlling cell motility.
Strengths:
Using a proximity-based proteomic assay, the authors reveal the protein network interacting with caveolae upon mechanical stimuli. This approach is elegant and allows to identify a substantial new set of factors involved in the mechano-regulation of caveolin-1, some of which have been verified directly in the cell by PLA. This study provides a compelling set of data on the interactions between caveolae and its cortical network which was so far ill-characterized.
We thank the reviewer for this positive feedback.
Weaknesses:
The methodology demonstrating an impact of membrane tension is not precise enough to directly assess a direct role on caveolae at a subcellular scale, that is between the front and the rear of the cell. First, a better characterization of the "front-rear" cellular model is encouraged.
We agree with the reviewer that a quantitative analysis of the caveolae front-rear polarity would strengthen our conclusions. To address this, we have analysed the localisation of Cav1 and cavins in detail and in a large pool of cells, both in fixed and live cells. Our quantification clearly shows that Cav1 and cavins are enriched at the cell rear. This is now shown in Figure 1 and Figure 1 - Figure Supplement 1. To demonstrate that Cav1/cavins are truly rear-localised we analysed live migrating cells expressing tagged Cav1 or cavins. This analysis, which was performed on several individual time lapse movies, showed that caveolae rear localisation is remarkably stable (e.g. Figure 1C and 1D). We also present novel data panels and movies showing caveolae dynamics during rear retractions, in dividing cells, and in cells that polarise de novo. This new data is now described in the first paragraph of the results section.
Secondly, authors frequently present osmotic shock as "high membrane tension" stimuli. While osmotic shock is widely used in the field, this study is focused only on caveolae localized at the rear of cell and it remains unclear how the level of a global mechanical stimuli triggered by an osmotic shock could mimic a local stimuli.
We agree with the reviewer that osmotic shock will cause a global increase in membrane tension and therefore is only of limited value to understand how membrane tension is regulated at the rear, and how caveolae respond to such a local stimulus. It was not our aim nor is it our expertise to address such questions. To answer this sophisticated optogenetic approaches or localised membrane tension measurements (e.g. through the use of the Flipper-TR probe) are needed. It is beyond the scope of this manuscript to perform such experiments. However, given the strong enrichment of caveolae at the cell rear, we believe it is justified to propose that the changes we observe in the proteome do (mostly) reflect changes in caveolae at the rear. We have now included several quantifications on fixed cells, live cells, and PLA assays to support that caveolae are highly enriched at the rear. In addition, and importantly, a recent preprint by the Roux lab shows that membrane tension gradients indeed exist in many migrating and non-migrating cells 5. Using very similar hypotonic shock assays, the Caswell lab also showed that low membrane tension at the rear is required for caveolae formation 6. We have included a section in the discussion in which we elaborate on how membrane tension is controlled in migrating cells, and how it might regulate caveolae rear localisation.
In the present case, it remains unknown the extent to which this mechanical stress is physiologically relevant to mimic mechanical forces applied at the rear of a migrating cell.
This is true. Our study does not address the nature of mechanical forces at the cell rear. This a complex subject that is technically challenging to address, and therefore is beyond the scope of this manuscript.
Some images are not satisfying to fully support the conclusions of the article.
We agree that some of the images, in particular the ones presented for the PLA assays, do not always show a clear rear localisation of caveolae. We have explained above why this is the case. We hope that our new quantitative measurements, movies and figure panels, addresses the reviewer’s concern.
At this stage, the lack of an unbiased quantitative analysis of the spatio-temporal analysis of caveolae upon well-defined mechanical stimuli is also needed.
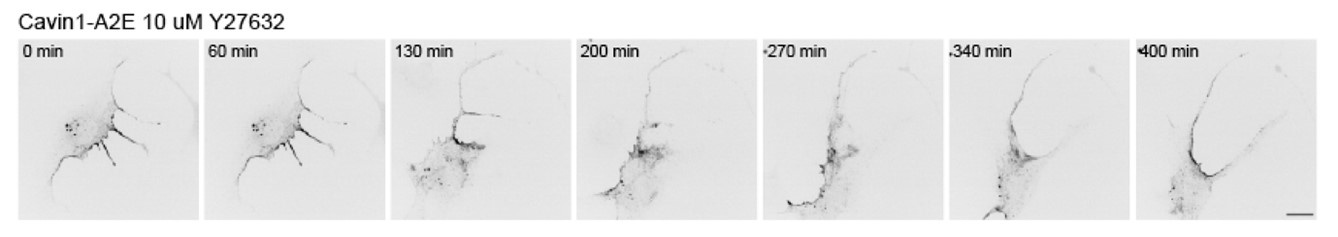
These are all very good points that were previously addressed beautifully by the Caswell group 6. To address this in part in our RPE1 cell system, we imaged RPE1 cells exposed to the ROCK inhibitor Y27632 (see Author response image 3). The data shows that cell rear retraction is impeded in response to ROCK inhibition, which is in line with several previous reports. Cavin-1 remained mostly associated with the cell rear, although the distribution appeared more diffuse. We believe this data does not add much new insight into how caveolae function at the rear, and hence was not included in the manuscript.
Author response image 3.
Effect of ROCK inhibition on cavin1 rear localisation and rear retraction. Cells were imaged one hour after the addition of Y27632.
Cells on images, in particular Figure 1, are difficult to see. Signal-to noise ratio in different cell area could generate a biased. Since there is inconsistency between caveolae density and localization between Figures, more solid illustrations are needed along quantitative analysis.
As mentioned above, we have carefully analysed the localisation of caveolae in fixed cells (using Cav1 and cavin1 antibodies as well as Cav1 and cavin fusion proteins) and in live cells transfected with various different caveolae proteins. The analysis clearly demonstrates an enrichment of caveolae at the rear (Figure 1 and Figure 1 – Figure Supplement 1). Our tomography and TEM data supports this as well (Figure 2).
References:
Qiao Y, Chen J, Lim YB, et al. YAP Regulates Actin Dynamics through ARHGAP29 and Promotes Metastasis. Cell reports. 2017;19(8):1495-1502.
Rausch V, Bostrom JR, Park J, et al. The Hippo Pathway Regulates Caveolae Expression and Mediates Flow Response via Caveolae. Curr Biol. 2019;29(2):242-255 e246.
Hung V, Udeshi ND, Lam SS, et al. Spatially resolved proteomic mapping in living cells with the engineered peroxidase APEX2. Nat Protoc. 2016;11(3):456-475.
Wiggan O, Shaw AE, DeLuca JG, Bamburg JR. ADF/cofilin regulates actomyosin assembly through competitive inhibition of myosin II binding to F-actin. Dev Cell. 2012;22(3):530-543.
Juan Manuel García-Arcos AM, Julissa Sánchez Velázquez, Pau Guillamat, Caterina Tomba, Laura Houzet, Laura Capolupo, Giovanni D’Angelo, Adai Colom, Elizabeth Hinde, Charlotte Aumeier, Aurélien Roux. Actin dynamics sustains spatial gradients of membrane tension in adherent cells. bioRxiv 20240715603517. 2024.
Hetmanski JHR, de Belly H, Busnelli I, et al. Membrane Tension Orchestrates Rear Retraction in Matrix-Directed Cell Migration. Dev Cell. 2019;51(4):460-475 e410.
Tsai TY, Collins SR, Chan CK, et al. Efficient Front-Rear Coupling in Neutrophil Chemotaxis by Dynamic Myosin II Localization. Dev Cell. 2019;49(2):189-205 e186.
Mueller J, Szep G, Nemethova M, et al. Load Adaptation of Lamellipodial Actin Networks. Cell. 2017;171(1):188-200 e116.
De Belly H, Yan S, Borja da Rocha H, et al. Cell protrusions and contractions generate long-range membrane tension propagation. Cell. 2023.
Matthaeus C, Sochacki KA, Dickey AM, et al. The molecular organization of differentially curved caveolae indicates bendable structural units at the plasma membrane. Nat Commun. 2022;13(1):7234.
Sinha B, Koster D, Ruez R, et al. Cells respond to mechanical stress by rapid disassembly of caveolae. Cell. 2011;144(3):402-413.
Lieber AD, Schweitzer Y, Kozlov MM, Keren K. Front-to-rear membrane tension gradient in rapidly moving cells. Biophysical journal. 2015;108(7):1599-1603.
Shi Z, Graber ZT, Baumgart T, Stone HA, Cohen AE. Cell Membranes Resist Flow. Cell. 2018;175(7):1769-1779 e1713.
Grande-Garcia A, Echarri A, de Rooij J, et al. Caveolin-1 regulates cell polarization and directional migration through Src kinase and Rho GTPases. The Journal of cell biology. 2007;177(4):683-694.
Grande-Garcia A, del Pozo MA. Caveolin-1 in cell polarization and directional migration. Eur J Cell Biol. 2008;87(8-9):641-647.
Ludwig A, Howard G, Mendoza-Topaz C, et al. Molecular composition and ultrastructure of the caveolar coat complex. PLoS biology. 2013;11(8):e1001640.
-

eLife assessment
This important study uses convincing time-resolved proximity proteomics, validated with proximity ligation assays, to provide new insight into mechanical regulation of caveolin-1 complexes that form in migrating cells. Solid follow up experiments reveal a reciprocal relationship between mechanosensitive caveolae and RhoGTPase signalling in migrating cells, but evidence supporting a direct link between the newly identified factors with a specific caveolae subpopulation remains incomplete at this stage.
-

Reviewer #1 (Public Review):
In this study, Girardello et al. use proteomics to reveal the membrane tension sensitive caveolin-1 interactome in migrating cells. The authors use EM and surface rendering to demonstrate that caveolae formed at the rear of migrating cells are complex membrane-linked multilobed structures, and they devise a robust strategy to identify caveolin-1 associated proteins using APEX2-mediated proximity biotinylation. This important dataset is further validated using proximity ligation assays to confirm key interactions, and follows up with an interrogation of a surprising relationship between caveolae and RhoGTPase signalling, where caveolin-1 recruits ROCK1 under high membrane tension conditions, and ROCK1 activity is required to reform caveolae upon reversion to isotonic solution. However, caveolin-1 recruits the …
Reviewer #1 (Public Review):
In this study, Girardello et al. use proteomics to reveal the membrane tension sensitive caveolin-1 interactome in migrating cells. The authors use EM and surface rendering to demonstrate that caveolae formed at the rear of migrating cells are complex membrane-linked multilobed structures, and they devise a robust strategy to identify caveolin-1 associated proteins using APEX2-mediated proximity biotinylation. This important dataset is further validated using proximity ligation assays to confirm key interactions, and follows up with an interrogation of a surprising relationship between caveolae and RhoGTPase signalling, where caveolin-1 recruits ROCK1 under high membrane tension conditions, and ROCK1 activity is required to reform caveolae upon reversion to isotonic solution. However, caveolin-1 recruits the RhoA inactivator ARHGAP29 when membrane tension is low and ARHGAP29 overexpression leads to disassembly of caveolae and reduced cell motility. This study builds on previous findings linking caveolae to positive feedback regulation of RhoA signalling, and provides further evidence that caveolae serve to drive rear retraction in migration but also possess an intrinsic brake to limit RhoA activation, leading the authors to suggest that cycles of caveolae assembly and disassembly could thereby be central to establish a stable cell rear for persistent cell migration
A major strength of the manuscript is the robust proteomic dataset. The experimental set up is well defined and mostly well controlled, and there is good internal validation in that the high abundance of core caveolar proteins in low membrane tension (isotonic) conditions, and absence under high membrane tension (brief hypo-osmotic shock) conditions, correlating very well with previous finding. The data could however be better presented to show where statically robust changes occur, and supplementary information should include a table of showing abundance. It's very good to see a link to PRIDE, providing a useful resource for the community.
The authors detail several known interactions and their mechanosensitivty, but also report new interactors of caveolin-1. Several mechanosensitive interactions of caveolin-1 take place at the cell rear, but others are more diffuse across the cell looking at the PLA data (e.g FLN1, CTTN, HSPB1; Figure 4A-F and Figure 4 supplement 1). It is interesting to speculate that those at the cell rear are involved in caveolae, whilst others are linked specifically to caveolin-1 (e.g. dolines). PLA or localisation analysis with Cavin1/PTRF may be able to resolve this and further specify caveolae versus non-caveolae mechanosensitive interactions.
The Cav1/ARHGAP29 influence on YAP signalling is interesting, but appear to be quite isolated from the rest of the manuscript. Does overexpression of ARHGAP29 influence YAP signalling and/or caveolar protein expression/Cav1pY14?
ARHGAP29 and RhoA/ROCK1 related observations are very interesting and potentially really important. However, the link between ARHGAP29 and caveolae is not well established (other than in proteomic data). PLA or FRET could help establish this.
The relationship between ARHGAP29 and RhoA signalling is not well defined. Is GAP activity important in determining the effect on migration and caveolae formation? What is the effect on RhoA activity? Alternatively, the authors could investigate YAP dependent transcriptional regulation downstream of overexpression. -

Reviewer #2 (Public Review):
Girardello et al investigated the composition of the molecular machinery of caveolae governing their mechano-regulation in migrating cells. Using live cell imaging and RPE1 cells, the authors provide a spatio-temporal analysis of cavin-3 distribution during cell migration and reveal that caveolae are preferentially localized at the rear of the cell in a stable manner. They further characterize these structures using electron tomography and reveal an organization into clusters connected to the cell surface. By performing a proteomic approach, they address the interactome of caveolin-1 proteins upon mechanical stimulation by exposing RPE1 cells to hypo-osmotic shock (which aims to increase cell membrane tension) or not as a control condition. The authors identify over 300 proteins, notably proteins related to …
Reviewer #2 (Public Review):
Girardello et al investigated the composition of the molecular machinery of caveolae governing their mechano-regulation in migrating cells. Using live cell imaging and RPE1 cells, the authors provide a spatio-temporal analysis of cavin-3 distribution during cell migration and reveal that caveolae are preferentially localized at the rear of the cell in a stable manner. They further characterize these structures using electron tomography and reveal an organization into clusters connected to the cell surface. By performing a proteomic approach, they address the interactome of caveolin-1 proteins upon mechanical stimulation by exposing RPE1 cells to hypo-osmotic shock (which aims to increase cell membrane tension) or not as a control condition. The authors identify over 300 proteins, notably proteins related to actin cytoskeleton and cell adhesion. These results were further validated in cellulo by interrogating protein-protein interactions using proximity ligation assays and hypo-osmotic shock. These experiments confirmed previous data showing that high membrane tension induces caveolae disassembly in a reversible manner. Eventually, based on literature and on the results collected by the proteomic analysis, authors investigated more deeply the molecular signaling pathway controlling caveolae assembly upon mechanical stimuli. First, they confirm the targeting of ROCK1 with Caveolin-1 and the implication of the kinase activity for caveolae formation (at the rear of the cell). Then, they show that RhoGA ARHGAP29, a factor newly identified by the proteomic analysis, is also implicated in caveolae mechano-regulation likely through YAP protein and found that overexpression of RHoGA ARHGAP29 affects cell motility. Overall, this paper interrogated the role of membrane tension in caveolae located at the rear of the cell and identified a new pathway controlling cell motility.
Strengths:
Using a proximity-based proteomic assay, the authors reveal the protein network interacting with caveolae upon mechanical stimuli. This approach is elegant and allows to identify a substantial new set of factors involved in the mechano-regulation of caveolin-1, some of which have been verified directly in the cell by PLA. This study provides a compelling set of data on the interactions between caveolae and its cortical network which was so far ill-characterized.
Weaknesses:
The methodology demonstrating an impact of membrane tension is not precise enough to directly assess a direct role on caveolae at a subcellular scale, that is between the front and the rear of the cell. First, a better characterization of the "front-rear" cellular model is encouraged. Secondly, authors frequently present osmotic shock as "high membrane tension" stimuli. While osmotic shock is widely used in the field, this study is focused only on caveolae localized at the rear of cell and it remains unclear how the level of a global mechanical stimuli triggered by an osmotic shock could mimic a local stimuli. In the present case, it remains unknown the extent to which this mechanical stress is physiologically relevant to mimic mechanical forces applied at the rear of a migrating cell.
Some images are not satisfying to fully support the conclusions of the article. At this stage, the lack of an unbiased quantitative analysis of the spatio-temporal analysis of caveolae upon well-defined mechanical stimuli is also needed. Cells on images, in particular Figure 1, are difficult to see. Signal-to noise ratio in different cell area could generate a biased. Since there is inconsistency between caveolae density and localization between Figures, more solid illustrations are needed along quantitative analysis. -