Heat Shock Factor 1 forms nuclear condensates and restructures the yeast genome before activating target genes
Curation statements for this article:-
Curated by eLife
eLife assessment
This is a valuable contribution to our understanding of how different cell stressors (ethanol or heat-shock) elicit unique responses at the genomic and topographical level under the regulation of yeast transcription factor Hsf1, providing solid evidence documenting the temporal coupling (or lack thereof) between Hsf1 aggregation and long-range communication among co-regulated heat-shock loci versus chromatin remodeling and transcriptional activation. A particular strength is the combination of genomic and imaging-based experimental approaches applied to genetically engineered in vivo systems.
This article has been Reviewed by the following groups
Discuss this preprint
Start a discussion What are Sciety discussions?Listed in
- Evaluated articles (eLife)
Abstract
In insects and mammals, 3D genome topology has been linked to transcriptional states yet whether this link holds for other eukaryotes is unclear. Using both ligation proximity and fluorescence microscopy assays, we show that in Saccharomyces cerevisiae , Heat Shock Response ( HSR ) genes dispersed across multiple chromosomes and under the control of Heat Shock Factor (Hsf1) rapidly reposition in cells exposed to acute ethanol stress and engage in concerted, Hsf1-dependent intergenic interactions. Accompanying 3D genome reconfiguration is equally rapid formation of Hsf1-containing condensates. However, in contrast to the transience of Hsf1-driven intergenic interactions that peak within 10–20 min and dissipate within 1 hr in the presence of 8.5% (v/v) ethanol, transcriptional condensates are stably maintained for hours. Moreover, under the same conditions, Pol II occupancy of HSR genes, chromatin remodeling, and RNA expression are detectable only later in the response and peak much later (>1 hr). This contrasts with the coordinate response of HSR genes to thermal stress (39°C) where Pol II occupancy, transcription, histone eviction, intergenic interactions, and formation of Hsf1 condensates are all rapid yet transient (peak within 2.5–10 min and dissipate within 1 hr). Therefore, Hsf1 forms condensates, restructures the genome and transcriptionally activates HSR genes in response to both forms of proteotoxic stress but does so with strikingly different kinetics. In cells subjected to ethanol stress, Hsf1 forms condensates and repositions target genes before transcriptionally activating them.
Article activity feed
-

-

-

eLife assessment
This is a valuable contribution to our understanding of how different cell stressors (ethanol or heat-shock) elicit unique responses at the genomic and topographical level under the regulation of yeast transcription factor Hsf1, providing solid evidence documenting the temporal coupling (or lack thereof) between Hsf1 aggregation and long-range communication among co-regulated heat-shock loci versus chromatin remodeling and transcriptional activation. A particular strength is the combination of genomic and imaging-based experimental approaches applied to genetically engineered in vivo systems.
-

Reviewer #2 (Public Review):
Rubio et al. study the behavior of the transcription factor Hsf1 under ethanol stress, examining its distribution within the nucleus and the coalescence of heat shock response genes in budding yeast. In comparison to the heat shock response, the response to ethanol stress shows similar gene coalescence and Hsf1 binding. However, there is a notable delay in the transcriptional response to ethanol, and a disconnect between it and the appearance of irreversible Hsf1 condensates/puncta, highlighting important differences in how Hsf1 responds to these two related but distinct environmental stresses.
The authors have addressed the majority of my previous comments effectively. The Sis1 experiment provides a clear illustration of a distinctive response to ethanol and heat. This work offers a comprehensive …
Reviewer #2 (Public Review):
Rubio et al. study the behavior of the transcription factor Hsf1 under ethanol stress, examining its distribution within the nucleus and the coalescence of heat shock response genes in budding yeast. In comparison to the heat shock response, the response to ethanol stress shows similar gene coalescence and Hsf1 binding. However, there is a notable delay in the transcriptional response to ethanol, and a disconnect between it and the appearance of irreversible Hsf1 condensates/puncta, highlighting important differences in how Hsf1 responds to these two related but distinct environmental stresses.
The authors have addressed the majority of my previous comments effectively. The Sis1 experiment provides a clear illustration of a distinctive response to ethanol and heat. This work offers a comprehensive perspective on Hsf1 in stress response from multiple angles.
-

Reviewer #3 (Public Review):
This is an interesting manuscript that builds off of this group's previous work focused on the interface between Hsf1, heat shock protein (HSP) mRNA production, and 3D genome topology. Here the group subjects the yeast Saccharomyces cerevisiae to either heat stress (HS) or ethanol stress (ES) and examines Hsf1 and Pol II chromatin binding, Histone occupancy, Hsf1 condensates, HSP gene coalescence (by 3C and live cell imaging), and HSP mRNA expression (by RT-qPCR and live cell imaging). The manuscript is well written, and the experiments seem well done, and generally rigorous, with orthogonal approaches performed to support conclusions. The main findings are that both HS and ES result in Hsf1/Pol II-dependent intergenic interactions, along with formation of Hsf1 condensates. Yet, while HS results in rapid and …
Reviewer #3 (Public Review):
This is an interesting manuscript that builds off of this group's previous work focused on the interface between Hsf1, heat shock protein (HSP) mRNA production, and 3D genome topology. Here the group subjects the yeast Saccharomyces cerevisiae to either heat stress (HS) or ethanol stress (ES) and examines Hsf1 and Pol II chromatin binding, Histone occupancy, Hsf1 condensates, HSP gene coalescence (by 3C and live cell imaging), and HSP mRNA expression (by RT-qPCR and live cell imaging). The manuscript is well written, and the experiments seem well done, and generally rigorous, with orthogonal approaches performed to support conclusions. The main findings are that both HS and ES result in Hsf1/Pol II-dependent intergenic interactions, along with formation of Hsf1 condensates. Yet, while HS results in rapid and strong induction of HSP gene expression and Hsf1 condensate resolution, ES result in slow and weak induction of HSP gene expression without Hsf1 condensate resolution. Thus, the conclusion is somewhat phenomenological - that the same transcription factor can drive distinct transcription, topologic, and phase-separation behavior in response to different types of stress.
-

Author response:
The following is the authors’ response to the previous reviews.
Reviewer #2 (Public Review):
The authors have addressed the majority of my comments effectively. The new Sis1 experiment provides a clear illustration of a distinctive response to ethanol and heat. This work offers a comprehensive perspective on Hsf1 in stress response from multiple angles. I have two additional comments to improve the paper without re-review:
(Original point #3) Could the authors clarify the differences between DPY1561 and the original strain used? There appears to be missing statistical analysis for Figure 1E at the bottom.
DPY1561 is a haploid version of the original heterozygous diploid strain (LRY033). We opted for this strain in the analysis depicted in Figures 1D and 1E since 100% of Hsp104 is BFP-tagged; thus, the signal above …
Author response:
The following is the authors’ response to the previous reviews.
Reviewer #2 (Public Review):
The authors have addressed the majority of my comments effectively. The new Sis1 experiment provides a clear illustration of a distinctive response to ethanol and heat. This work offers a comprehensive perspective on Hsf1 in stress response from multiple angles. I have two additional comments to improve the paper without re-review:
(Original point #3) Could the authors clarify the differences between DPY1561 and the original strain used? There appears to be missing statistical analysis for Figure 1E at the bottom.
DPY1561 is a haploid version of the original heterozygous diploid strain (LRY033). We opted for this strain in the analysis depicted in Figures 1D and 1E since 100% of Hsp104 is BFP-tagged; thus, the signal above background is stronger and the scoring of Hsp104 foci cleaner. The statistical analysis (Mann Whitney test) for the lower graphs in Fig. 1E has been added. We thank the reviewer for pointing this out.
(Original point #4) In the new Figure 7F, '% transcription' and '% coalescence' are presented. My understanding is that Figures 7D and 7E aim to demonstrate the correlation between HSP104 transcription (a continuous variable) and HSP104-HSP12 coalescence (a binary variable) at the single-cell level. However, averaging the data across cells masks individual variations and potential anti-correlations. The authors could explore statistical methods that handle correlations between a continuous variable and a binary variable. Alternatively, consider converting 'HSP104 transcription' to a binary variable and then performing a chi-square test to assess the association.
We thank the reviewer for this suggestion. In response, we have made the following changes:
(1) Clarified that the data used in this analysis were derived from Fig. 7 – figure supplement 1 in which ‘HSP104 transcription’ was converted to a binary variable.
(2) Indicated that the theoretical ceiling for coalescence of these tagged alleles is 25% given their heterozygous state (Figure 7–figure supplement 1D legend). In the other 75% of cells scored, HSP104-HSP12 coalescence might also be taking place but is not detectable using this strategy. Therefore, it is not possible to elucidate any anti-correlation between HSR transcription and HSR coalescence in this experiment.
In addition, we attempted to buttress the argument suggested by the Pearson correlation coefficient analysis (Fig. 7F) that a stronger association exists between transcription and gene coalescence in heat-shocked (HS) vs. ethanol stressed (ES) cells. To do so, we used the chi-square test as suggested by the reviewer. However, the results of this test were ambiguous, and we therefore did not include it in the manuscript.
-

-

eLife assessment
This is a valuable contribution to our understanding of how different cell stressors (ethanol or heat-shock) elicit unique responses at the genomic and topographical level under the regulation of yeast transcription factor Hsf1, providing solid evidence documenting the temporal coupling (or lack thereof) between Hsf1 aggregation and long-range communication among co-regulated heat-shock loci versus chromatin remodeling and transcriptional activation. A particular strength is the combination of genomic and imaging-based experimental approaches applied to genetically engineered in vivo systems.
-

Reviewer #2 (Public Review):
Significance:
Rubio et al. study the behavior of the transcription factor Hsf1 under ethanol stress, examining its distribution within the nucleus and the coalescence of heat shock response genes in budding yeast. In comparison to the heat shock response, the response to ethanol stress shows similar gene coalescence and Hsf1 binding. However, there is a notable delay in the transcriptional response to ethanol, and a disconnect between it and the appearance of irreversible Hsf1 condensates/puncta, highlighting important differences in how Hsf1 responds to these two related but distinct environmental stresses.
Overview and general concerns (from the original review):
The authors studied how yeast responds to ethanol stress (8.5%) and compared it to the heat shock response (from 25{degree sign}C to 39{degree …
Reviewer #2 (Public Review):
Significance:
Rubio et al. study the behavior of the transcription factor Hsf1 under ethanol stress, examining its distribution within the nucleus and the coalescence of heat shock response genes in budding yeast. In comparison to the heat shock response, the response to ethanol stress shows similar gene coalescence and Hsf1 binding. However, there is a notable delay in the transcriptional response to ethanol, and a disconnect between it and the appearance of irreversible Hsf1 condensates/puncta, highlighting important differences in how Hsf1 responds to these two related but distinct environmental stresses.
Overview and general concerns (from the original review):
The authors studied how yeast responds to ethanol stress (8.5%) and compared it to the heat shock response (from 25{degree sign}C to 39{degree sign}C). They observed a more gradual increase in the expression of heat shock response (HSR) genes during ethanol stress compared to heat shock. Additionally, the recruitment of Hsf1 and Pol II to HSR genes, and the inter- and intrachromosomal interactions among these genes, showed slower kinetics under ethanol stress. They attribute the delay in transcriptional response to chromatin compaction induced by ethanol. Despite this delay, these interactions persisted longer. Hsf1 clusters, previously documented during the heat shock response, were also observed during ethanol stress and persisted for an extended period. The conditional degradation of Hsf1 and Rpb1 eliminated most inter- and intrachromosomal interactions for heat shock responsive genes in both ethanol stress and heat shock conditions, indicating the importance of these factors for long distance interactions between HSR genes. Overall, this manuscript provides novel insights into the differential behavior of HSR genes under different stress conditions. This contributes to the broader understanding of how different stressors might elicit unique responses at the genomic and topographical level under the regulation of transcription factor Hsf1.
The central finding of the study highlights the different dynamics of Hsf1, Pol II, and gene organization in response to heat shock versus ethanol stress. However, one important limitation to consider is that the two chosen conditions may not be directly comparable. For a balanced assessment, the authors should ideally expose yeast to various ethanol concentrations and different heat shock temperatures, ensuring the observed differences stem from the nature of the stressor rather than suboptimal stress intensity. At the very least, an additional single ethanol concentration point on each side of 8.5% should be investigated to ensure that 8.5% is near the optimum. In fact, comparing the number of Hsp104 foci in the two conditions in Fig. 1E and F suggests that the yeast is likely experiencing different intensities of stress for the chosen heat shock condition and ethanol concentration used in this study.
A second significant concern is the use of the term "Hsf1 condensate". Chowdhary et al.'s 2022 Molecular Cell study highlighted an inhomogeneous distribution and rapid dynamics of Hsf1 clustering upon heat shock, with sensitivity to 1,6-hexandiol, which is interpreted as evidence for condensation by LLPS. But this interpretation has been criticized severely by McSwiggen at al. Genes Dev 2019 and Mussacchio EMBO J 2022. It is important to mention that 1,6-hexandiol is known to affect chromatin organization (Itoh et al. Life Science Alliance 2021). Describing such clusters as 'condensates' without further experimental evidence is premature. I encourage authors to settle on their neutral term 'puncta' which they use interchangeably with 'condensate' so as not to confuse the reader. The dynamic binding and unbinding of the low-abundance Hsf1 at coalescent chromatin target sites might explain the liquid-like properties of these clusters without the need for invoking the phase-separation hypothesis. While Hsf1 clusters exhibit features consistent with phase-separated condensates, other equally plausible alternative mechanisms, such as dynamic site-specific interactions (Musacchio, EMBO J, 2022), should also be considered. This is best left for the discussion where the underlying mechanism for puncta formation can be addressed.
Comments on revised version:
The authors have addressed the majority of my comments effectively. The new Sis1 experiment provides a clear illustration of a distinctive response to ethanol and heat. This work offers a comprehensive perspective on Hsf1 in stress response from multiple angles. I have two additional comments to improve the paper without re-review:
(Original point #3) Could the authors clarify the differences between DPY1561 and the original strain used? There appears to be missing statistical analysis for Figure 1E at the bottom.
(Original point #4) In the new Figure 7F, '% transcription' and '% coalescence' are presented. My understanding is that Figures 7D and 7E aim to demonstrate the correlation between HSP104 transcription (a continuous variable) and HSP104-HSP12 coalescence (a binary variable) at the single-cell level. However, averaging the data across cells masks individual variations and potential anti-correlations. The authors could explore statistical methods that handle correlations between a continuous variable and a binary variable. Alternatively, consider converting 'HSP104 transcription' to a binary variable and then performing a chi-square test to assess the association.
-

Reviewer #3 (Public Review):
This is an interesting manuscript that builds off of this group's previous work focused on the interface between Hsf1, heat shock protein (HSP) mRNA production, and 3D genome topology. Here the group subjects the yeast Saccharomyces cerevisiae to either heat stress (HS) or ethanol stress (ES) and examines Hsf1 and Pol II chromatin binding, Histone occupancy, Hsf1 condensates, HSP gene coalescence (by 3C and live cell imaging), and HSP mRNA expression (by RT-qPCR and live cell imaging). The manuscript is well written, and the experiments seem well done, and generally rigorous, with orthogonal approaches performed to support conclusions. The main findings are that both HS and ES result in Hsf1/Pol II-dependent intergenic interactions, along with formation of Hsf1 condensates. Yet, while HS results in rapid and …
Reviewer #3 (Public Review):
This is an interesting manuscript that builds off of this group's previous work focused on the interface between Hsf1, heat shock protein (HSP) mRNA production, and 3D genome topology. Here the group subjects the yeast Saccharomyces cerevisiae to either heat stress (HS) or ethanol stress (ES) and examines Hsf1 and Pol II chromatin binding, Histone occupancy, Hsf1 condensates, HSP gene coalescence (by 3C and live cell imaging), and HSP mRNA expression (by RT-qPCR and live cell imaging). The manuscript is well written, and the experiments seem well done, and generally rigorous, with orthogonal approaches performed to support conclusions. The main findings are that both HS and ES result in Hsf1/Pol II-dependent intergenic interactions, along with formation of Hsf1 condensates. Yet, while HS results in rapid and strong induction of HSP gene expression and Hsf1 condensate resolution, ES result in slow and weak induction of HSP gene expression without Hsf1 condensate resolution. Thus, the conclusion is somewhat phenomenological - that the same transcription factor can drive distinct transcription, topologic, and phase-separation behavior in response to different types of stress. While identifying a mechanistic basis for these results would be a tough task perhaps beyond the scope of this study, it would nevertheless be helpful to place these results in context with a series of other studies demonstrating across various organisms showing Hsf1 driving distinct activities dependent on the context of activation. Perhaps even more importantly, this work left out PMID: 32015439 which is particularly relevant considering that it shows that it is human HSF1 condensate resolution rather than simple condensate formation that is associated with HSF1 transcriptional activity - which are similar to the findings here with this particular dose of HS resulting in resolution and high transcriptional activity versus ES resulting in resolution failure and lower activity. It is also worth noting that the stresses themselves are quite different - ethanol can be used as a carbon source and so beyond inducing proteotoxic stress, the yeast are presumably adapting to this distinct metabolic state. Basically, it is not clear whether these differences are due to the dose of stress, versus we are looking at an early timepoint as ES initiates a genome-wide chromatin restructuring and gene expression reprogramming that goes beyond a response to proteotoxic stress. This reviewer is not suggesting a barrage of new experiments, but perhaps discussion points to contextualize results.
Comments on latest version:
The authors have addressed my concerns.
-

Author response:
The following is the authors’ response to the original reviews.
Public Reviews:
Reviewer #1:
Overall, the conclusions appear appropriately supported by the data, and the data appear of high quality.
Strengths:
The particular strengths of the paper include an impressive combination of genomic and imaging-based approaches and insightful genetically engineered cell systems. The manuscript reports interesting and potentially important findings. The text is generally very well written, the ideas are clearly explained, and the reasoning is easy to follow.
Weaknesses:
The main weakness seems to be that the heat and ethanol shock approaches likely elicit pleiotropic effects, and therefore it is a challenge to test the causal relationship between various observations. Nevertheless, even as indirect effects might contribute to …
Author response:
The following is the authors’ response to the original reviews.
Public Reviews:
Reviewer #1:
Overall, the conclusions appear appropriately supported by the data, and the data appear of high quality.
Strengths:
The particular strengths of the paper include an impressive combination of genomic and imaging-based approaches and insightful genetically engineered cell systems. The manuscript reports interesting and potentially important findings. The text is generally very well written, the ideas are clearly explained, and the reasoning is easy to follow.
Weaknesses:
The main weakness seems to be that the heat and ethanol shock approaches likely elicit pleiotropic effects, and therefore it is a challenge to test the causal relationship between various observations. Nevertheless, even as indirect effects might contribute to some of the authors' observations, the results are definitively worth reporting.
We agree that these two proteotoxic stresses can impact cell physiology in multiple ways and discuss this on lines 132-143 and 500-519. Moreover, in this revision we have more rigorously quantified the extent of proteotoxic stress elicited by the 39°C heat shock and 8.5% ethanol stress (Figure 1E; see response 1 to Reviewer 2). We have additionally added new Figure 2 that reveals an important difference in the way Hsf1 and its negative regulator, the Hsp70 co-chaperone Sis1, respond to HS and ES. This difference is evident at two different intensities for each stress as described in more detail below (see response 1 to Reviewer 2).
Presentation of some of the data could be improved.
We agree and have made improvements/data additions to multiple figures: Figure 1E; Figures 3A, B; Figures 4A, B; Figure 7 (data drawn from original Fig. 6 and Fig. 6 – fig. suppl. 1 and reorganized); Fig. 8B; Figure 9; Figure 10. Corresponding enhancements to the supplemental figures have been made as well.
Reviewer #2:
(1) The central finding of the study highlights the different dynamics of Hsf1, Pol II, and gene organization in response to heat shock versus ethanol stress. However, one important limitation to consider is that the two chosen conditions may not be directly comparable. For a balanced assessment, the authors should ideally expose yeast to various ethanol concentrations and different heat shock temperatures, ensuring the observed differences stem from the nature of the stressor rather than suboptimal stress intensity. At the very least, an additional single ethanol concentration point on each side of 8.5% should be investigated to ensure that 8.5% is near the optimum. In fact, comparing the number of Hsp104 foci in the two conditions in Fig. 1E and F suggests that the yeast is likely experiencing different intensities of stress for the chosen heat shock condition and ethanol concentration used in this study.
We thank the reviewer for this important suggestion. In this revision, we have included an enhanced analysis of the yeast cellular response to each of these stresses. As illustrated in revised Figure 1, the two stresses used throughout this study – 39°C heat shock and 8.5% ethanol stress – both elicit a proteotoxic response, as assayed by the de novo formation of Hsp104 clusters. While 10 min exposure to 8.5% ethanol results in the formation of multiple discrete (spherical) foci, a 10 min exposure to the elevated temperature leads the appearance of multiple, largely diffuse Hsp104 clusters, some of which are spherical (new Fig. 1D). The difference in morphology notwithstanding, we have attempted to quantify these clusters using Imaris v. 10.0.1 image analysis software; the results are depicted in Fig. 1E. Such quantification suggests that 8.5% ethanol elicits a more intense stress than exposure to 39°C. A caveat is that it is unclear whether diffuse Hsp104 clusters are comparable to compact Hsp104 foci (see response 3 below).
Beyond the apparent difference in intensity, a new analysis presented in new Figure 2 reveals that heat shock, elicited by temperature upshift to either 39°C or 42°C, induces relocalization of the J-protein Sis1 – a key negative regulator of Hsf1 – from the nucleoplasm to the nucleolar periphery. Sis1’s perinucleolar ring localization agrees with previous findings of 39°C heat-shocked cells (Feder et al., 2021). Ethanol stress, whether 5% or 8.5%, initially causes Sis1 to relocalize diffusely throughout the nucleus and cytosol. At 10 min, Sis1 localizes to the periphery of the nucleus, thereby providing a marked contrast to what is observed in response to heat shock. These new results are described on lines 174-191.
Taking these two observations together, we asked whether a less severe ethanol stress (5%) would induce Hsf1 puncta. It does, and as rapidly as 8.5% ethanol (data are presented in revised Figure 8-figure supplement 1). Interestingly, in the presence of 5% ethanol, Hsf1 puncta begin to dissolve at 30 min. This strongly contrasts with the case when cells are exposed to 8.5% ethanol (Figure 8; Figure 8-figure supplement 1). As we state in this revision (lines 414-424), the sustained presence of condensates that we originally observed is likely the consequence of the intensity of the proteotoxic stress elicited by exposure to 8.5% ethanol; analogous responses to these two stress conditions have been observed before (lines 495-501).
(2) A second significant concern is the use of the term "Hsf1 condensate". Chowdhary et al.'s 2022 Molecular Cell study highlighted an inhomogeneous distribution and rapid dynamics of Hsf1 clustering upon heat shock, with sensitivity to 1,6-hexandiol, which is interpreted as evidence for condensation by LLPS. However this interpretation has been criticized severely by McSwiggen et al. Genes Dev 2019 and Mussacchio EMBO J 2022. It is important to mention that 1,6-hexandiol is known to affect chromatin organization (Itoh et al. Life Science Alliance 2021). Describing such clusters as 'condensates' without further experimental evidence is premature.
While we appreciate and largely agree with the point made by this reviewer, we prefer to maintain the term “condensate”. Banani et al (2017) originally defined “biomolecular condensate” to mean selforganized membrane-free compartments that concentrate specific biomolecules. It was never meant to imply LLPS although its widespread use in the literature has led to that implication. We clarify our use of this term on lines 99-104.
(3) Figure 1: Why does ethanol stress at 0 min display a larger number of Hsp104 foci per cell than heat shock at the same time? How are foci defined by the authors? In Fig. 1D, there are many smaller puncta. A comparative assessment of the number and size of foci for heat shock and ethanol stress would be beneficial.
We thank the reviewer for raising this point and have addressed it as follows. First, we repeated the assay with a different strain (DPY1561) and increased the number of cells assayed from 40 to 200. This larger sample size created the same T=0 baseline for both stresses (Figure 1E). Second, we define Hsp104 foci as diffraction-limited structures with a diameter of ~0.4 µm (lines 747-749). Third, employing Imaris v. 10.0.1, we quantified foci size (= volume) and a summary graph has been added to Figure 1E that also displays the number of foci per cell. In the legend to this figure, we point out that to conduct this analysis we assumed that the diffuse Hsp104 clusters seen in HS cells are comparable to the compact Hsp104 foci in ES cells (lines 1169-1171).
(4) Figure 2: Selecting a housekeeping gene with consistent expression levels is crucial for meaningful qPCR analysis. Do SCR1 mRNA levels fluctuate during heat shock or ethanol stress?
We thank the reviewer for this question. In revised Figure 3 – figure supplement 1C we provide a new graph (reproduced here) revealing that the levels of SCR1 do not significantly change under either heat shock or ethanol stress relative to the non-stressed control (0 min). One-way ANOVA analysis was performed for both HS and ES and p values were 0.094 and 0.083, respectively (calculated using GraphPad Prism 8).
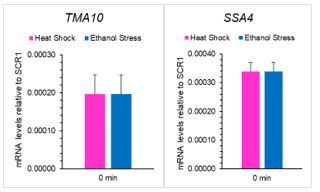
(5) Additionally, certain genes, such as TMA10 and SSA4, lack visible bars at time 0. Are these levels undetectable? The varying y-axis scales are confusing; presenting data as relative fold changes could offer a clearer perspective.
Transcript levels for all genes evaluated here are detectable, even in the basal unstressed state. They are not visible on the histogram for certain genes at T= 0 due to the prodigious fold-increase in RNA elicited by heat shock. However, to address this concern, we have added a bar graph inset displaying basal transcript levels for each gene in revised Figure 3. We reproduce data for SSA4 and TMA10 in the graphs below. In addition, we present transcript levels in new Figure 3 - figure supplement 1 for cells subjected to ethanol stress to allow a better appreciation of their increase over time.
Author response image 1.
(6) Line 239: The evidence for chromatin compaction is unconvincing. An increase in H3 occupancy by ChIP might indicate a reduction in histone exchange dynamics but may not relate to overall chromatin compaction. The authors use H2A-mCherry to suggest a decrease in chromatin volume, but this data is not persuasive. Did the authors observe any changes in nuclear size? Perhaps quantifying chromatin compaction more directly, using signal intensity per volume, would be informative.
To address this concern, we attempted to quantify integrated density for H2A-mCherry using Image J software. While the volume decreased for both stresses, the integrated density only increased for ethanol stress. We speculate that this may be due to photobleaching which has been reported for heat shock. The combination of heat and acidic pH contribute to loss of fluorescence signal (Alkaabi et al., 2005). While the integrated density supports the idea of global chromatin compaction in the ethanol stress condition, given the above concerns with the HS sample we elected to not present these data.
(7) Line 340: The claim of a "strong spatiotemporal correlation" isn't evident from the data. Could correlation coefficients be provided? There is potential anti-correlation in Fig. 6 - Figure Supplement 1C.
We thank the reviewer for this excellent suggestion. We now present an analysis of the correlation between HSP104 – HSP12 coalescence and HSP104 transcription for both HS and ES time courses, using single cell data of Figures 7D, 7E and Figure 7- suppl. 1D. This analysis is presented in new Figure 7F.
(8) Figure 8: The WT data in Fig 8 seem inconsistent with Fig. 4 (e.g. the interaction frequency for HSP104 and SSA2). Are these fluctuations between experiments, or are they side effects of IAA treatment? The use of ethanol as an IAA solvent vehicle raises concerns. It would be beneficial if the authors could demonstrate that 1.7% ethanol in the control does not induce ethanol stress.
We acknowledge that there existed an inconsistency in the magnitude of intergenic interaction frequencies reported in the two experiments for HSP104 and SSA2. Some of this might be attributed to the fact that different strains were used, W303-1B in Figure 4 and LRY016 (W303-1B; LEU2::pGPD1osTIR1) in Figure 8. Nonetheless, in each experiment there was a prodigious fold-increase in interaction frequency over the no stress (T= 0 min) control for both HS and ES conditions and moreover, in each experiment the magnitude of this interaction was greater for the 2.5 min HS sample vs. the 10 min ES sample. However, to obviate this concern, we have removed the HSP104-SSA2 analysis from Figure 9 (corresponds to original Fig. 8).
Regarding the second point, we cannot entirely rule out the concern that the 1.7% ethanol vehicle might impact 3C interaction frequencies. It is unlikely to be significant, however, given that most other pairwise tests evaluated in the two experiments (Figs. 5 and 9) resulted in similar 3C values. In particular, there was no consistent trend towards higher (or lower) interaction frequencies in the IAA experiment of Fig. 9.
Reviewer #3:
This is an interesting manuscript that builds off of this group's previous work focused on the interface between Hsf1, heat shock protein (HSP) mRNA production, and 3D genome topology. Here the group subjects the yeast Saccharomyces cerevisiae to either heat stress (HS) or ethanol stress (ES) and examines Hsf1 and Pol II chromatin binding, Histone occupancy, Hsf1 condensates, HSP gene coalescence (by 3C and live cell imaging), and HSP mRNA expression (by RT-qPCR and live cell imaging). The manuscript is well written, and the experiments seem well done, and generally rigorous, with orthogonal approaches performed to support conclusions…While identifying a mechanistic basis for the results [presented here] would be a tough task perhaps beyond the scope of this study, it would nevertheless be helpful to place these results in context with a series of other studies…importantly, this work left out PMID: 32015439 (HSF1 phase transition mediates stress adaptation and cell fate decisions) which is particularly relevant considering that it shows that it is human HSF1 condensate resolution rather than simple condensate formation that is associated with HSF1 transcriptional activity - which is similar to the findings here with this particular dose of HS resulting in resolution and high transcriptional activity versus ES resulting in resolution failure and lower activity.
We thank the Reviewer for pointing out this oversight. In this revision, we cite Gaglia et al., 2020 and several others reporting HSF1 foci formation in human cells exposed to heat shock. The single cell analysis of Gaglia et al argued that dissolution of large HSF1 foci (aka “nuclear stress bodies”), typically several µm in diameter and localized over satellite III DNA repeats (Jolly et al., 1997, 2002), correlates with HSP gene activation. Importantly, these condensates are postulated to act as reservoirs of HSF1, sequestered away from HSP genes (Gaglia et al., 2020). In contrast, Zhang et al., 2022 has shown that human HSF1 inducibly forms small condensates (~300 nm) that localize over HSP genes and whose formation directly correlates with HSP gene activation (we discuss the Jolly, Gaglia and Zhang findings on lines 382-394). Likewise, our work shows that in yeast, Hsf1 inducibly forms small, dynamic clusters that colocalize with HSR genes within 2.5 min of exposure to elevated temperature; these dissolve ~20-60 min later (Figure 8 and Figure 8-supp. 1). In concert with Hsf1 condensate formation, HSR gene repositioning and transcription/ Pol II recruitment are likewise evident within 2.5 min. Therefore, in HS cells there exists coordinate induction of condensate formation, Pol II recruitment, transcription and intergenic interactions (for a detailed kinetic analysis of HSR gene interactions, see Figures 5 and 6 of Chowdhary et al, 2017). This tight temporal relationship is absent in ethanol stressed cells (Figures 3, 4, 5, 6, 7, 8; summarized in Figure 10 and Table 1).
It is also worth noting that the stresses themselves are quite different - ethanol can be used as a carbon source and so beyond inducing proteotoxic stress, the yeast are presumably adapting to this distinct metabolic state. Basically, it is not clear whether these differences are due to the dose of stress, versus we are looking at an early timepoint as ES initiates a genome-wide chromatin restructuring and gene expression reprogramming that goes beyond a response to proteotoxic stress. This reviewer is not suggesting a barrage of new experiments, but perhaps discussion points to contextualize results.
We thank the reviewer for this suggestion and in our revised manuscript discuss these issues (lines 414424 and 486-498 [5% vs. 8.5% ethanol]; lines 500-519 [ethanol as a metabolite]).
Recommendations for the authors:
Reviewer #1:
(1) In Figure 1E, the number of foci in control (0 min) cells is very different for the two conditions. Could the authors clarify/check this? Based on the mean numbers at time point 0, the control cells for the ethanol treatment already contain about 10-20 Hsp104 foci, compared to around 5 foci per cell in the control for heat shock.
We thank the reviewer for raising this point and have repeated the assay with a different strain (DPY1561). And as shown in Figure 1E, have confirmed that the control samples have similar number of foci.
(2) In the same Figure 1E, is the P-value relative to the control or the same time point in the other treatment? A comparison across treatments would be necessary to support the claim in lines 168-171 of the text.
The statistical analysis (Mann Whitney test) was performed by comparing each stress timepoint to the no stress control. We clarify this in the figure legend.
(3) In Figure 1D, the heat-shock condition shows the same cells that are used in the control, but the cells in the ethanol-shock condition are different. This is a bit visually misleading compared to the experimental setup shown in panel 1C. The authors could show the control cells for the ethanol condition as well.
We thank the reviewer for this excellent suggestion and have added the 0 min image for the ethanol stress conditions.
(4) In Figure 7B adding images at 60min would help underscore the point that the condensates are stable in ethanol shocked cells.
We appreciate this suggestion as well and have included a 60 min timepoint for both stresses (Figure 8B).
Reviewer #2:
(1) Line 113: Has it not been established that yeast Hsf1 is constitutively trimeric?
In yeast, only a fraction of Hsf1 is thought to be constitutively trimeric and it is this species that binds high-affinity HSEs even under non-stressful conditions (Giardina & Lis, 1995; Pincus et al., 2018). We have added this clarification to the text (lines 121-123).
(2) Ethanol can precipitate proteins, especially in rich media like YPD. Did the authors notice any protein precipitation? If yes, how do they account for effects due to nutrient loss by precipitation?
This is an interesting point, but we did not notice any precipitates in either rich or synthetic liquid media containing 8.5% (v/v) ethanol for any of the time points used in the experiments.
(3) Figure 3: The figure appears incomplete. Can enhancer, promoter, coding region, and 3'UTR be shown consistently for all genes examined?
In response to this point, we have simplified this figure (new Fig. 4) by uniform presentation of factor occupancy at enhancer, promoter, and coding region loci for all but one of the genes evaluated. For HSP12 (330 bp), we were unable to distinguish promoter from coding region since the average sonicated chromatin fragment obtained using a Bioruptor is ~300 bp. Therefore, we evaluated only the HSP12 coding region for Pol II and histone H3 occupancy.
(4) Figure 4: The comparison between heat shock at 2.5 min and ethanol stress at later points is puzzling. Why not use consistent time points as in Fig. 3?
Time points for the two stresses examined in this figure (new Fig. 5) were selected to represent times of peak intergenic interaction between HSR genes. These times were derived from our earlier analysis of 3C interactions during a heat shock time course (Figs. 5, 6 of Chowdhary et al., 2017) and ES data presented in this study, including Fig. 4 (Pol II ChIP time course) and Fig. 6 (3C time course). Data presented in Figs. 5 and 6 are consistent with the notion that intergenic interactions in cells subjected to ethanol stress are delayed relative to those observed in heat shocked cells, peaking in most cases at ~10 min (vs. ~2.5 min for heat stress (Chowdhary et al., 2017)).
(5) Figure 5: Fig. 5B top panel seems to show color inconsistencies for bars at 0 and 120 min. Also, the xaxis on the top left panel seems to have a typo; should it read "10," not "0?"
We thank the reviewer for the observation. We changed the graphs in new Figure 6 to display the same color for all time points. We also fixed the typo.
(6) Line 302: The evidence presented supports maximal mRNA levels, but the claim of "maximal transcription" requires support from nascent RNA analysis.
We agree that RT-qPCR measures mRNA abundance, not nascent transcription. We have changed the text to refer to “transcript levels” where pertinent (lines 301-302; 1331-1332).
(7) How long do loci remain coalescent during heat shock versus ethanol stress? Both 3C and imaging analyses do not differentiate between frequency and duration, which seems essential for understanding interaction dynamics.
We thank the reviewer for this excellent question. In new Fig. 7D,E (data drawn from Fig. 6 – fig. suppl. 1), HSR gene coalescence detected in single cells over a HS or ES time course is charted. Interpretable data exist for a small number of cells. Moreover, for both HS and ES states, in certain cells coalescence between the representative Hsf1 target genes HSP104 and HSP12 dissolves and then reappears. With this caveat in mind, the data suggest that HSP104-HSP12 coalescence can last at least 15 min in HS cells and up to 30 min in ES cells. We have not emphasized this point in the manuscript since a far more comprehensive analysis – beyond the scope of this study – is required.
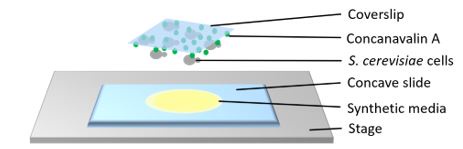
(8) For longer analyses, how do the authors accommodate potential ethanol concentration changes due to evaporation?
For liquid cultures, we relied on maintaining minimal changes in the vapor pressure within the experimental vessel; to facilitate that, flasks were tightly covered to minimize evaporation and temperature was kept at 25°C. For most molecular analyses (RT-qPCR, ChIP, 3C), we confined our analysis to the first 60 min. For microscopy, the samples were encased within a concave slide, covered by a coverslip, as illustrated below. In addition, to tightly seal the coverslip on the slide we used petrolatum. This arrangement minimized evaporation.
Author response image 2.
(9) Figure 9: This legend seems to have an incomplete sentence: "(represented using ...)."
We have substituted an entirely new model in this revised manuscript (new Figure 10) that omits the use of an ellipsis. (We had used it to symbolize a delay in the appearance of HSR gene transcription in ES cells.)
References
Alkaabi, K. M., Yafea, A., & Ashraf, S. S. (2005). Effect of pH on thermal- and chemical-induced denaturation of GFP. Applied Biochemistry and Biotechnology, 126(2), 149–156. https://doi.org/10.1385/ABAB:126:2:149
Chowdhary, S., Kainth, A. S., & Gross, D. S. (2017). Heat Shock Protein Genes Undergo Dynamic Alteration in Their Three-Dimensional Structure and Genome Organization in Response to Thermal Stress. Molecular and Cellular Biology, 37(24), 1–23. https://doi.org/10.1128/mcb.00292-17
Feder, Z. A., Ali, A., Singh, A., Krakowiak, J., Zheng, X., Bindokas, V. P., Wolfgeher, D., Kron, S. J., & Pincus, D. (2021). Subcellular localization of the J-protein Sis1 regulates the heat shock response. Journal of Cell Biology, 220(1), e202005165. https://doi.org/10.1083/JCB.202005165
Gaglia, G., Rashid, R., Yapp, C., Joshi, G. N., Li, C. G., Lindquist, S. L., Sarosiek, K. A., Whitesell, L., Sorger, P. K., & Santagata, S. (2020). HSF1 phase transition mediates stress adaptation and cell fate decisions. Nature Cell Biology, 22(2), 151–158. https://doi.org/10.1038/s41556-019-0458-3
Giardina, C., & Lis, J. T. (1995). Dynamic protein-DNA architecture of a yeast heat shock promoter. Molecular and Cellular Biology, 15(5), 2737–2744. https://doi.org/10.1128/mcb.15.5.2737
Jolly, C., Konecny, L., Grady, D. L., Kutskova, Y. A., Cotto, J. J., Morimoto, R. I., & Vourc’h, C. (2002). In vivo binding of active heat shock transcription factor 1 to human chromosome 9 heterochromatin during stress. Journal of Cell Biology, 156(5), 775–781. https://doi.org/10.1083/jcb.200109018
Jolly, C., Morimoto, R. I., Robert-Nicoud, M., & Vourc’h, C. (1997). HSF1 transcription factor concentrates in nuclear foci during heat shock: Relationship with transcription sites. Journal of Cell Science, 110(23), 2935–2941. https://doi.org/10.1242/jcs.110.23.2935
Pincus, D., Anandhakumar, J., Thiru, P., Guertin, M. J., Erkine, A. M., & Gross, D. S. (2018). Genetic and epigenetic determinants establish a continuum of Hsf1 occupancy and activity across the yeast genome. Molecular Biology of the Cell, 29(26), 3168–3182. https://doi.org/10.1091/mbc.E18-060353
Zhang, H., Shao, S., Zeng, Y., Wang, X., Qin, Y., Ren, Q., Xiang, S., Wang, Y., Xiao, J., & Sun, Y. (2022). Reversible phase separation of HSF1 is required for an acute transcriptional response during heat shock. Nature Cell Biology, 24(3), 340–352. https://doi.org/10.1038/s41556-022-00846-7
-

-

-

eLife assessment
This is a useful contribution to our understanding of how different cell stressors (ethanol or heat-shock) might elicit unique responses at the genomic and topographical level under the regulation of yeast transcription factor Hsf1, and of the temporal coupling (or lack thereof) between Hsf1 aggregation and long-range communication among co-regulated heat-shock loci versus chromatin remodeling and transcriptional activation. A particular strength is the combination of genomic and imaging-based experimental approaches applied to genetically engineered in vivo systems. While much of the data is convincing, the work is incomplete in not providing strong evidence supporting (i) a similar rate and extent of proteotoxic stress under the two chosen stress conditions, (ii) relatively greater bulk chromatin compaction elicited …
eLife assessment
This is a useful contribution to our understanding of how different cell stressors (ethanol or heat-shock) might elicit unique responses at the genomic and topographical level under the regulation of yeast transcription factor Hsf1, and of the temporal coupling (or lack thereof) between Hsf1 aggregation and long-range communication among co-regulated heat-shock loci versus chromatin remodeling and transcriptional activation. A particular strength is the combination of genomic and imaging-based experimental approaches applied to genetically engineered in vivo systems. While much of the data is convincing, the work is incomplete in not providing strong evidence supporting (i) a similar rate and extent of proteotoxic stress under the two chosen stress conditions, (ii) relatively greater bulk chromatin compaction elicited by ethanol, (iii) reproducible levels of interactions between chromosomal loci, and (iv) phase-separated condensates versus other types of Hsf1 clusters.
-

Reviewer #1 (Public Review):
Summary:
The study compares the transcriptional and epigenetic response of baker's yeast cells to heat shock and ethanol shock. The authors made several interesting observations. In response to heat shock, the transcription factor HSF1 rapidly forms foci, binds upstream elements of heat-shock-response genes, facilitates long-distance genomic contacts between heat-shock-response genes, and the genes are rapidly transcribed. In response to ethanol shock, the transcription factor HSF1 rapidly forms foci, binds upstream elements of heat-shock-response genes, facilitates long-distance genomic contacts between heat-shock-response genes, and yet transcription of the genes is substantially delayed. These insights are potentially important, as current models of eukaryotic gene control predict that physical contact …Reviewer #1 (Public Review):
Summary:
The study compares the transcriptional and epigenetic response of baker's yeast cells to heat shock and ethanol shock. The authors made several interesting observations. In response to heat shock, the transcription factor HSF1 rapidly forms foci, binds upstream elements of heat-shock-response genes, facilitates long-distance genomic contacts between heat-shock-response genes, and the genes are rapidly transcribed. In response to ethanol shock, the transcription factor HSF1 rapidly forms foci, binds upstream elements of heat-shock-response genes, facilitates long-distance genomic contacts between heat-shock-response genes, and yet transcription of the genes is substantially delayed. These insights are potentially important, as current models of eukaryotic gene control predict that physical contact between genes and regulatory elements is necessary, and in some cases sufficient to transcribe a gene. The current study indicates that the two effects are virtually decoupled in response to ethanol shock in yeast cells.Overall, the conclusions appear appropriately supported by the data, and the data appear of high quality.
Strengths:
The particular strengths of the paper include an impressive combination of genomic and imaging-based approaches and insightful genetically engineered cell systems. The manuscript reports interesting and potentially important findings. The text is generally very well written, the ideas are clearly explained, and the reasoning is easy to follow.Weaknesses:
The main weakness seems to be that the heat and ethanol shock approaches likely elicit pleiotropic effects, and therefore it is a challenge to test the causal relationship between various observations. Nevertheless, even as indirect effects might contribute to some of the authors' observations, the results are definitively worth reporting. Also, the presentation of some of the data could be improved. -

Reviewer #2 (Public Review):
Significance
Rubio et al. study the behavior of the transcription factor Hsf1 under ethanol stress, examining its distribution within the nucleus and the coalescence of heat shock response genes in budding yeast. In comparison to the heat shock response, the response to ethanol stress shows similar gene coalescence and Hsf1 binding. However, there is a notable delay in the transcriptional response to ethanol, and a disconnect between it and the appearance of irreversible Hsf1 condensates/puncta, highlighting important differences in how Hsf1 responds to these two related but distinct environmental stresses.Overview and general concerns
The authors studied how yeast responds to ethanol stress (8.5%) and compared it to the heat shock response (from 25{degree sign}C to 39{degree sign}C). They observed a more …Reviewer #2 (Public Review):
Significance
Rubio et al. study the behavior of the transcription factor Hsf1 under ethanol stress, examining its distribution within the nucleus and the coalescence of heat shock response genes in budding yeast. In comparison to the heat shock response, the response to ethanol stress shows similar gene coalescence and Hsf1 binding. However, there is a notable delay in the transcriptional response to ethanol, and a disconnect between it and the appearance of irreversible Hsf1 condensates/puncta, highlighting important differences in how Hsf1 responds to these two related but distinct environmental stresses.Overview and general concerns
The authors studied how yeast responds to ethanol stress (8.5%) and compared it to the heat shock response (from 25{degree sign}C to 39{degree sign}C). They observed a more gradual increase in the expression of heat shock response (HSR) genes during ethanol stress compared to heat shock. Additionally, the recruitment of Hsf1 and Pol II to HSR genes, and the inter- and intrachromosomal interactions among these genes, showed slower kinetics under ethanol stress. They attribute the delay in transcriptional response to chromatin compaction induced by ethanol. Despite this delay, these interactions persisted longer. Hsf1 clusters, previously documented during the heat shock response, were also observed during ethanol stress and persisted for an extended period. The conditional degradation of Hsf1 and Rpb1 eliminated most inter- and intrachromosomal interactions for heat shock responsive genes in both ethanol stress and heat shock conditions, indicating the importance of these factors for long-distance interactions between HSR genes. Overall, this manuscript provides novel insights into the differential behavior of HSR genes under different stress conditions. This contributes to the broader understanding of how different stressors might elicit unique responses at the genomic and topographical level under the regulation of transcription factor Hsf1.The central finding of the study highlights the different dynamics of Hsf1, Pol II, and gene organization in response to heat shock versus ethanol stress. However, one important limitation to consider is that the two chosen conditions may not be directly comparable. For a balanced assessment, the authors should ideally expose yeast to various ethanol concentrations and different heat shock temperatures, ensuring the observed differences stem from the nature of the stressor rather than suboptimal stress intensity. At the very least, an additional single ethanol concentration point on each side of 8.5% should be investigated to ensure that 8.5% is near the optimum. In fact, comparing the number of Hsp104 foci in the two conditions in Fig. 1E and F suggests that the yeast is likely experiencing different intensities of stress for the chosen heat shock condition and ethanol concentration used in this study.
A second significant concern is the use of the term "Hsf1 condensate". Chowdhary et al.'s 2022 Molecular Cell study highlighted an inhomogeneous distribution and rapid dynamics of Hsf1 clustering upon heat shock, with sensitivity to 1,6-hexandiol, which is interpreted as evidence for condensation by LLPS. However this interpretation has been criticized severely by McSwiggen et al. Genes Dev 2019 and Mussacchio EMBO J 2022. It is important to mention that 1,6-hexandiol is known to affect chromatin organization (Itoh et al. Life Science Alliance 2021). Describing such clusters as 'condensates' without further experimental evidence is premature. I encourage authors to settle on their neutral term 'puncta' which they use interchangeably with 'condensate' so as not to confuse the reader. The dynamic binding and unbinding of the low-abundance Hsf1 at coalescent chromatin target sites might explain the liquid-like properties of these clusters without the need for invoking the phase-separation hypothesis. While Hsf1 clusters exhibit features consistent with phase-separated condensates, other equally plausible alternative mechanisms, such as dynamic site-specific interactions (Musacchio, EMBO J, 2022), should also be considered. This is best left for the discussion where the underlying mechanism for puncta formation can be addressed.
Specific comments:
- Figure 1: Why does ethanol stress at 0 min display a larger number of Hsp104 foci per cell than heat shock at the same time? How are foci defined by the authors? In Fig. 1D, there are many smaller puncta. A comparative assessment of the number and size of foci for heat shock and ethanol stress would be beneficial.
- Figure 2: Selecting a housekeeping gene with consistent expression levels is crucial for meaningful qPCR analysis. Do SCR1 mRNA levels fluctuate during heat shock or ethanol stress? Additionally, certain genes, such as TMA10 and SSA4, lack visible bars at time 0. Are these levels undetectable? The varying y-axis scales are confusing; presenting data as relative fold changes could offer a clearer perspective.
- Line 239: The evidence for chromatin compaction is unconvincing. An increase in H3 occupancy by ChIP might indicate a reduction in histone exchange dynamics but may not relate to overall chromatin compaction. The authors use H2A-mCherry to suggest a decrease in chromatin volume, but this data is not persuasive. Did the authors observe any changes in nuclear size? Perhaps quantifying chromatin compaction more directly, using signal intensity per volume, would be informative.
- Line 340: The claim of a "strong spatiotemporal correlation" isn't evident from the data. Could correlation coefficients be provided? There is potential anti-correlation in Fig. 6 - Figure Supplement 1C.
- Figure 8: The WT data in Fig 8 seem inconsistent with Fig. 4 (e.g. the interaction frequency for HSP104 and SSA2). Are these fluctuations between experiments, or are they side effects of IAA treatment? The use of ethanol as an IAA solvent vehicle raises concerns. It would be beneficial if the authors could demonstrate that 1.7% ethanol in the control does not induce ethanol stress.
-

Reviewer #3 (Public Review):
This is an interesting manuscript that builds off of this group's previous work focused on the interface between Hsf1, heat shock protein (HSP) mRNA production, and 3D genome topology. Here the group subjects the yeast Saccharomyces cerevisiae to either heat stress (HS) or ethanol stress (ES) and examines Hsf1 and Pol II chromatin binding, Histone occupancy, Hsf1 condensates, HSP gene coalescence (by 3C and live cell imaging), and HSP mRNA expression (by RT-qPCR and live cell imaging). The manuscript is well written, and the experiments seem well done, and generally rigorous, with orthogonal approaches performed to support conclusions. The main findings are that both HS and ES result in Hsf1/Pol II-dependent intergenic interactions, along with the formation of Hsf1 condensates. Yet, while HS results in rapid …
Reviewer #3 (Public Review):
This is an interesting manuscript that builds off of this group's previous work focused on the interface between Hsf1, heat shock protein (HSP) mRNA production, and 3D genome topology. Here the group subjects the yeast Saccharomyces cerevisiae to either heat stress (HS) or ethanol stress (ES) and examines Hsf1 and Pol II chromatin binding, Histone occupancy, Hsf1 condensates, HSP gene coalescence (by 3C and live cell imaging), and HSP mRNA expression (by RT-qPCR and live cell imaging). The manuscript is well written, and the experiments seem well done, and generally rigorous, with orthogonal approaches performed to support conclusions. The main findings are that both HS and ES result in Hsf1/Pol II-dependent intergenic interactions, along with the formation of Hsf1 condensates. Yet, while HS results in rapid and strong induction of HSP gene expression and Hsf1 condensate resolution, ES results in slow and weak induction of HSP gene expression without Hsf1 condensate resolution. Thus, the conclusion is somewhat phenomenological - that the same transcription factor can drive distinct transcription, topologic, and phase-separation behavior in response to different types of stress. While identifying a mechanistic basis for these results would be a tough task perhaps beyond the scope of this study, it would nevertheless be helpful to place these results in context with a series of other studies demonstrating across various organisms showing Hsf1 driving distinct activities dependent on the context of activation. Perhaps even more importantly, this work left out PMID: 32015439 which is particularly relevant considering that it shows that it is human HSF1 condensate resolution rather than simple condensate formation that is associated with HSF1 transcriptional activity - which is similar to the findings here with this particular dose of HS resulting in resolution and high transcriptional activity versus ES resulting in resolution failure and lower activity. It is also worth noting that the stresses themselves are quite different - ethanol can be used as a carbon source and so beyond inducing proteotoxic stress, the yeast are presumably adapting to this distinct metabolic state. Basically, it is not clear whether these differences are due to the dose of stress, versus we are looking at an early timepoint as ES initiates a genome-wide chromatin restructuring and gene expression reprogramming that goes beyond a response to proteotoxic stress. This reviewer is not suggesting a barrage of new experiments, but perhaps discussion points to contextualize results.
-