Regulation of pDC fate determination by histone deacetylase 3
Curation statements for this article:-
Curated by eLife
eLife assessment
This study examines the expression of HDAC3 within DC compartment. Taking advantage of tamoxifen inducible ERT2-cre mouse model they observe the dependency of pDCs but not cDCs on HDAC3. The requirement of this histone modifier appears to occur during development around the CLP stage. Tamoxifen treated mice lack almost all pDC besides lymphoid progenitors. RNA seq studies identify multiple DC specific target genes within the remaining pDC - using Cut and Tag technology they validate some of the identified targets of HDAC3. Taken together, this study shows the requirement of HDAC3 on pDC but not cDC, congruent with the recent findings of a lymphoid origin of pDC.
This article has been Reviewed by the following groups
Discuss this preprint
Start a discussion What are Sciety discussions?Listed in
- Evaluated articles (eLife)
Abstract
Dendritic cells (DCs), the key antigen-presenting cells, are primary regulators of immune responses. Transcriptional regulation of DC development had been one of the major research interests in DC biology; however, the epigenetic regulatory mechanisms during DC development remains unclear. Here, we report that Histone deacetylase 3 ( Hdac3 ), an important epigenetic regulator, is highly expressed in pDCs, and its deficiency profoundly impaired the development of pDCs. Significant disturbance of homeostasis of hematopoietic progenitors was also observed in HDAC3-deficient mice, manifested by altered cell numbers of these progenitors and defective differentiation potentials for pDCs. Using the in vitro Flt3L supplemented DC culture system, we further demonstrated that HDAC3 was required for the differentiation of pDCs from progenitors at all developmental stages. Mechanistically, HDAC3 deficiency resulted in enhanced expression of cDC1-associated genes, owing to markedly elevated H3K27 acetylation (H3K27ac) at these gene sites in BM pDCs. In contrast, the expression of pDC-associated genes was significantly downregulated, leading to defective pDC differentiation.
Article activity feed
-

-

Author Response
Reviewer #1 (Public Review):
The work by Yijun Zhang and Zhimin He at al. analyzes the role of HDAC3 within DC subsets. Using an inducible ERT2-cre mouse model they observe the dependency of pDCs but not cDCs on HDAC3. The requirement of this histone modifier appears to be early during development around the CLP stage. Tamoxifen treated mice lack almost all pDCs besides lymphoid progenitors. Through bulk RNA seq experiment the authors identify multiple DC specific target gens within the remaining pDCs and further using Cut and Tag technology they validate some of the identified targets of HDAC3. Collectively the study is well executed and shows the requirement of HDAC3 on pDCs but not cDCs, in line with the recent findings of a lymphoid origin of pDC.
- While the authors provide extensive data on the requirement of …
Author Response
Reviewer #1 (Public Review):
The work by Yijun Zhang and Zhimin He at al. analyzes the role of HDAC3 within DC subsets. Using an inducible ERT2-cre mouse model they observe the dependency of pDCs but not cDCs on HDAC3. The requirement of this histone modifier appears to be early during development around the CLP stage. Tamoxifen treated mice lack almost all pDCs besides lymphoid progenitors. Through bulk RNA seq experiment the authors identify multiple DC specific target gens within the remaining pDCs and further using Cut and Tag technology they validate some of the identified targets of HDAC3. Collectively the study is well executed and shows the requirement of HDAC3 on pDCs but not cDCs, in line with the recent findings of a lymphoid origin of pDC.
- While the authors provide extensive data on the requirement of HDAC3 within progenitors, the high expression of HDAC3 in mature pDCs may underly a functional requirement. Have you tested INF production in CD11c cre pDCs? Are there transcriptional differences between pDCs from HDAC CD11c cre and WT mice?
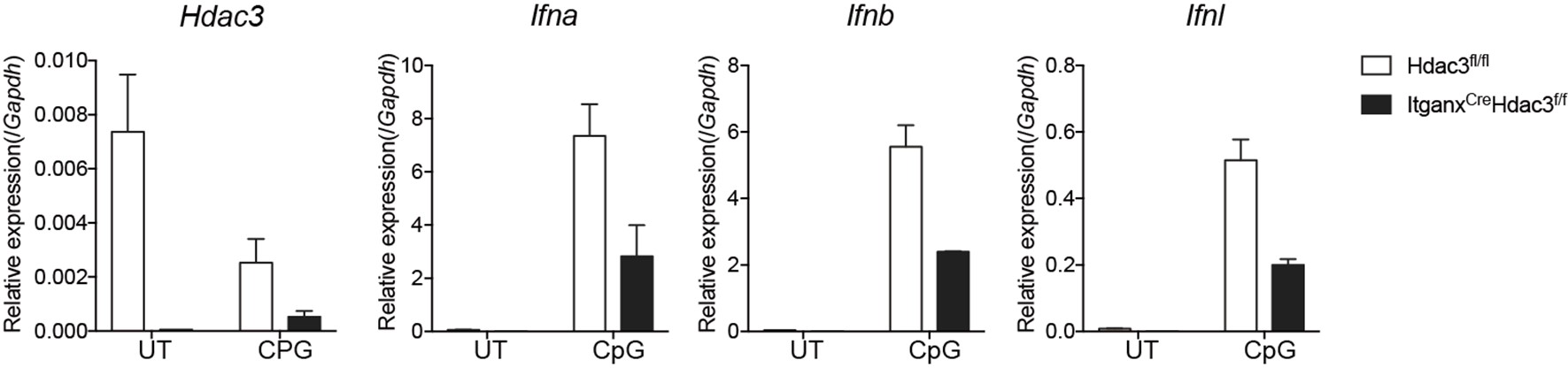
We greatly appreciate the reviewer’s point. We have confirmed that Hdac3 can be efficiently deleted in pDCs of Hdac3fl/fl-CD11c Cre mice (Figure 5-figure supplement 1 in revised manuscript). Furthermore, in those Hdac3fl/fl-CD11c Cre mice, we have observed significantly decreased expression of key cytokines (Ifna, Ifnb, and Ifnl) by pDCs upon activation by CpG ODN (shown in Author response image 1). Therefore, HDAC3 is also required for proper pDC function. However, we have yet to conduct RNA-seq analysis comparing pDCs from HDAC CD11c cre and WT mice.
Author response image 1.
Cytokine expression in Hdac3 deficient pDCs upon activation
- A more detailed characterization of the progenitor compartment that is compromised following depletion would be important, as also suggested in the specific points.
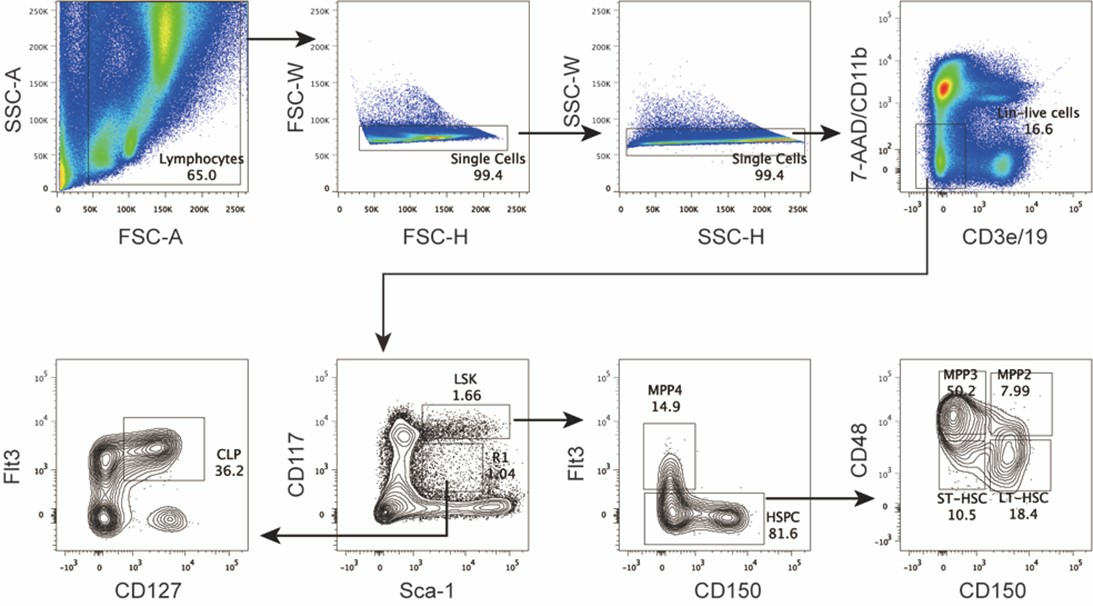
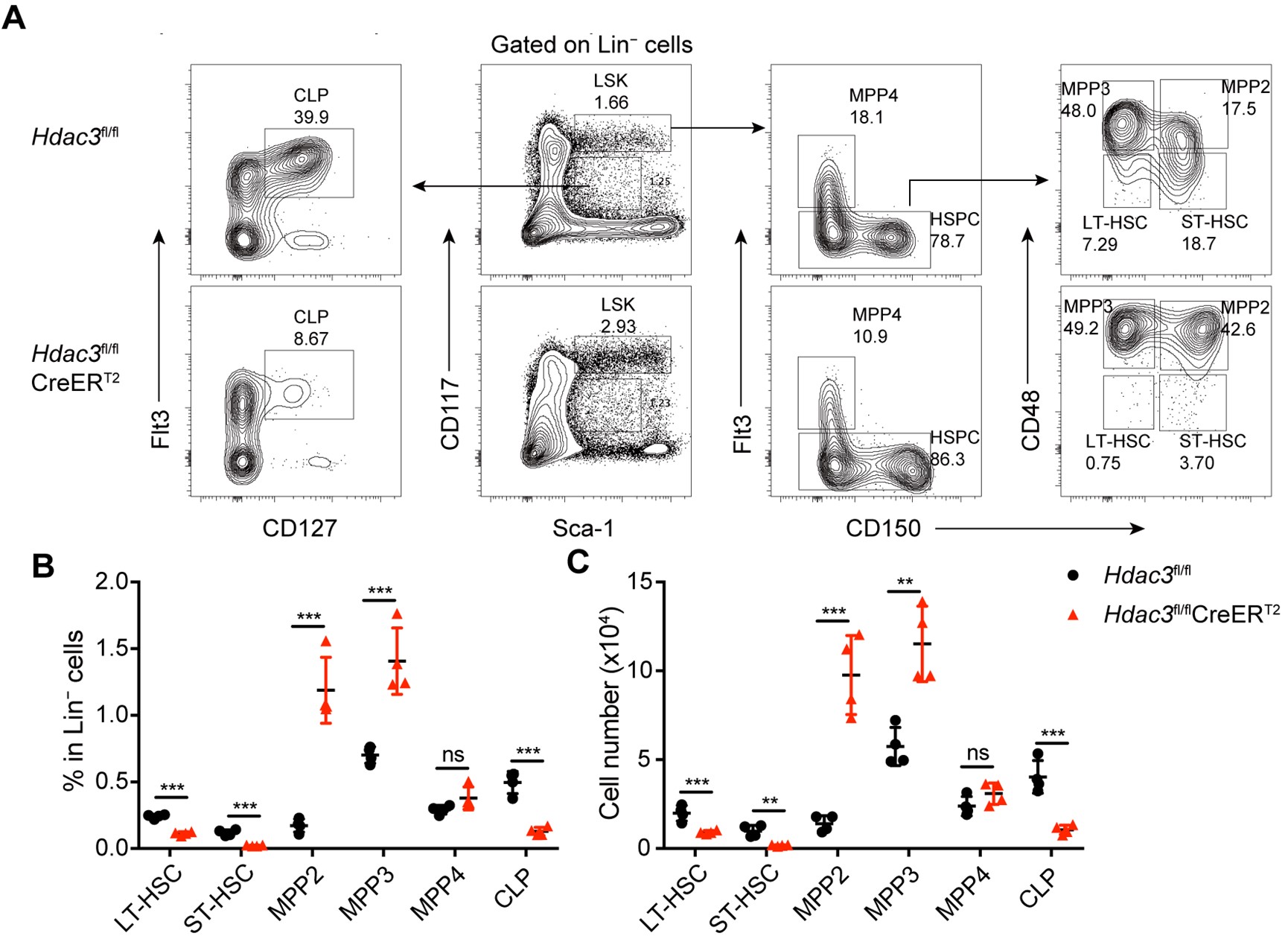
We thank the reviewer for this constructive suggestion. We have performed thorough analysis of the phenotype of hematopoietic stem cells and progenitor cells at various developmental stages in the bone marrow of Hdac3 deficient mice, based on the gating strategy from the recommended reference. Briefly, we analyzed the subpopulations of progenitors based on the description in the published report by "Pietras et al. 2015", namely MPP2, MPP3 and MPP4, using the same gating strategy for hematopoietic stem/progenitor cells. As shown in Author response image 2 and Author response image 3, we found that the number of LSK cells was increased in Hdac3 deficient mice, especially the subpopulations of MPP2 and MPP3, whereas no significant changes in MPP4. In contrast, the numbers of LT-HSC, ST-HSC and CLP were all dramatically decreased. This result has been optimized and added as Figure 3A in revised manuscript. The relevant description has been added and underlined in the revised manuscript Page 6 Line 164-168.
Author response image 2.
Gating strategy for hematopoietic stem/progenitor cells in bone marrow.
Author response image 3.
Hematopoietic stem/progenitor cells in Hdac3 deficient mice
Reviewer #2 (Public Review):
In this article Zhang et al. report that the Histone Deacetylase-3 (HDAC3) is highly expressed in mouse pDC and that pDC development is severely affected both in vivo and in vitro when using mice harbouring conditional deletion of HDAC3. However, pDC numbers are not affected in Hdac3fl/fl Itgax-Cre mice, indicating that HDCA3 is dispensable in CD11c+ late stages of pDC differentiation. Indeed, the authors provide wide experimental evidence for a role of HDAC3 in early precursors of pDC development, by combining adoptive transfer, gene expression profiling and in vitro differentiation experiments. Mechanistically, the authors have demonstrated that HDAC3 activity represses the expression of several transcription factors promoting cDC1 development, thus allowing the expression of genes involved in pDC development. In conclusion, these findings reveals HDAC3 as a key epigenetic regulator of the expression of the transcription factors required for pDC vs cDC1 developmental fate.
These results are novel and very promising. However, supplementary information and eventual further investigations are required to improve the clarity and the robustness of this article.
Major points
- The gating strategy adopted to identify pDC in the BM and in the spleen should be entirely described and shown, at least as a Supplementary Figure. For the BM the authors indicate in the M & M section that they negatively selected cells for CD8a and B220, but both markers are actually expressed by differentiated pDC. However, in the Figures 1 and 2 pDC has been shown to be gated on CD19- CD11b- CD11c+. What is the precise protocol followed for pDC gating in the different organs and experiments?
We apologize for not clearly describing the protocols used in this study. Please see the detailed gating strategy for pDC in bone marrow, and for pDC and cDC in spleen (Figure 4 and Figure 5). These information are now added to Figure1−figure supplement 3, The relevant description has been underlined in Page 5 Line 113-116, in revised manuscript.
We would like to clarify that in our study, we used two different panels of antibody cocktails, one for bone marrow Lin- cells, including mAbs to CD2/CD3/TER-119/Ly6G/B220/CD11b/CD8/CD19; the other for DC enrichment, including mAbs to CD3/CD90/TER-119/Ly6G/CD19. We included B220 in the Lineage cocktails to deplete B cells and pDCs, in order to enrich for the progenitor cells from bone marrow. However, when enriching for the pDC and cDC, B220 or CD8a were not included in the cocktail to avoid depletion of pDC and cDC1 subsets . For the flow cytometry analysis of pDCs, we gated pDCs as the CD19−CD11b−CD11c+B220+SiglecH+ population in both bone marrow and spleen. The relevant description has been underlined in the revised manuscript Page 16 Line 431-434.
- pDC identified in the BM as SiglecH+ B220+ can actually contain DC precursors, that can express these markers, too. This could explain why the impact of HDAC3 deletion appears stronger in the spleen than in the BM (Figures 1A and 2A). Along the same line, I think that it would important to show the phenotype of pDC in control vs HDAC3-deleted mice for the different pDC markers used (SiglecH, B220, Bst2) and I would suggest to include also Ly6D, taking also in account the results obtained in Figures 4 and 7. Finally, as HDCA3 deletion induces downregulation of CD8a in cDC1 and pDC express CD8a, it would important to analyse the expression of this marker on control vs HDAC3-deleted pDC.
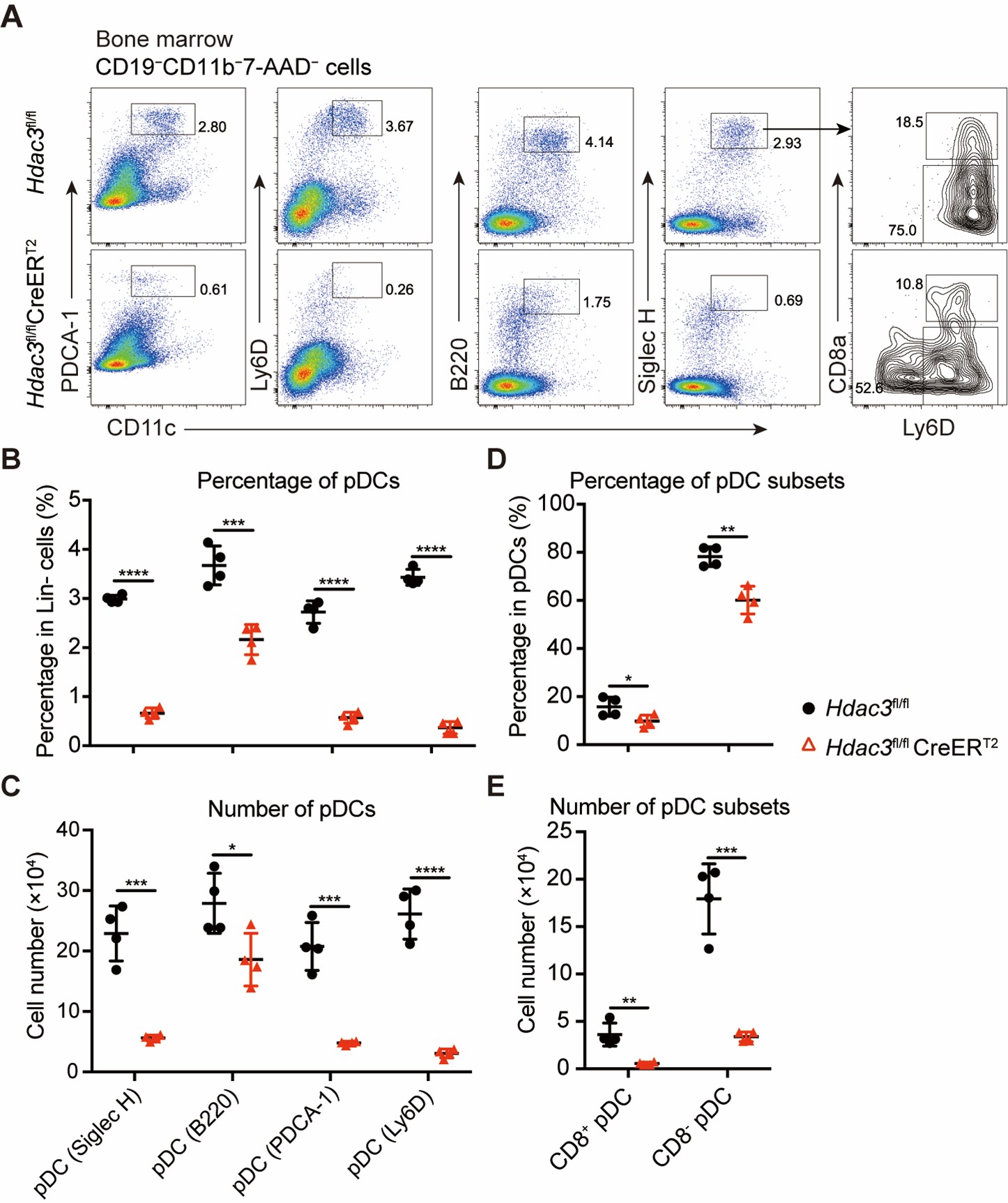
We agree with the reviewer’s points. In the revised manuscript, we incorporated major surface markers, including Siglec H, B220, Ly6D, and PDCA-1, all of which consistently demonstrated a substantial decrease in the pDC population in Hdac3 deficient mice. Moreover, we did notice that Ly6D+ pDCs showed higher degree of decrease in Hdac3 deficient mice. Additionally, percentage and number of both CD8+ pDC and CD8- pDC were decreased in Hdac3 deficient mice (Author response image 4). These results are shown in Figure1−figure supplement 4 of the revised manuscript. The relevant description has been added and underlined in the revised manuscript Page 5 Line 121-125.
Author response image 4.
Bone marrow pDCs in Hdac3 deficient mice revealed by multiple surface markers
- How do the authors explain that in the absence of HDAC3 cDC2 development increased in vivo in chimeric mice, but reduced in vitro (Figures 2B and 2E)?
As shown in the response to the Minor point 5 of Reviewer#1. Briefly, we suggested that the variabilities maybe explained by the timing of anaysis after HDAC3 deletion. In Figure 2C, we analyzed cells from the recipients one week after the final tamoxifen treatment and observed no significant change in the percentage of cDC2 when further pooled all the experiment data. In Figure 2E, where tamoxifen was administered at Day 0 in Flt3L-mediated DC differentiation in vitro, the DC subsets generated were then analyzed at different time points. We observed no significant changes in cDCs and cDC2 at Day 5, but decreases in the percentage of cDC2 were observed at Day 7 and Day 9. This suggested that the cDC subsets at Day 5 might have originated from progenitors at a later stage, while those at Day 7 and Day 9 might originate form the earlier progenitors. Therefore, based on these in vitro and in vivo experiments, we believe that the variation in the cDC2 phenotype might be attributed to the progenitors at different stages that generated these cDCs.
- More generally, as reported also by authors (line 207), the reconstitution with HDAC3-deleted cells is poorly efficient. Although cDC seem not to be impacted, are other lymphoid or myeloid cells affected? This should be expected as HDAC3 regulates T and B development, as well as macrophage function. This should be important to know, although this does not call into question the results shown, as obtained in a competitive context.
In this study, we found no significant influence on T cells, mature B cells or NK cells, but immature B cells were significantly decreased, in Hdac3-ERT2-Cre mice after tamoxifen treatment (Figure 6). However, in the bone marrow chimera experiments, the numbers of major lymphoid cells were decreased due to the impaired reconstitution capacity of Hdac3 deficient progenitors. Consistent with our finding, it has been reported that HDAC3 was required for T cell and B cell generation, in HDAC3-VavCre mice (Summers et al., 2013), and was necessary for T cell maturation (Hsu et al., 2015). Moreover, HDAC3 is also required for the expression of inflammatory genes in macrophages upon activation (Chen et al., 2012; Nguyen et al., 2020).
- What are the precise gating strategies used to identify the different hematopoietic precursors in the Figure 4 ? In particular, is there any lineage exclusion performed?
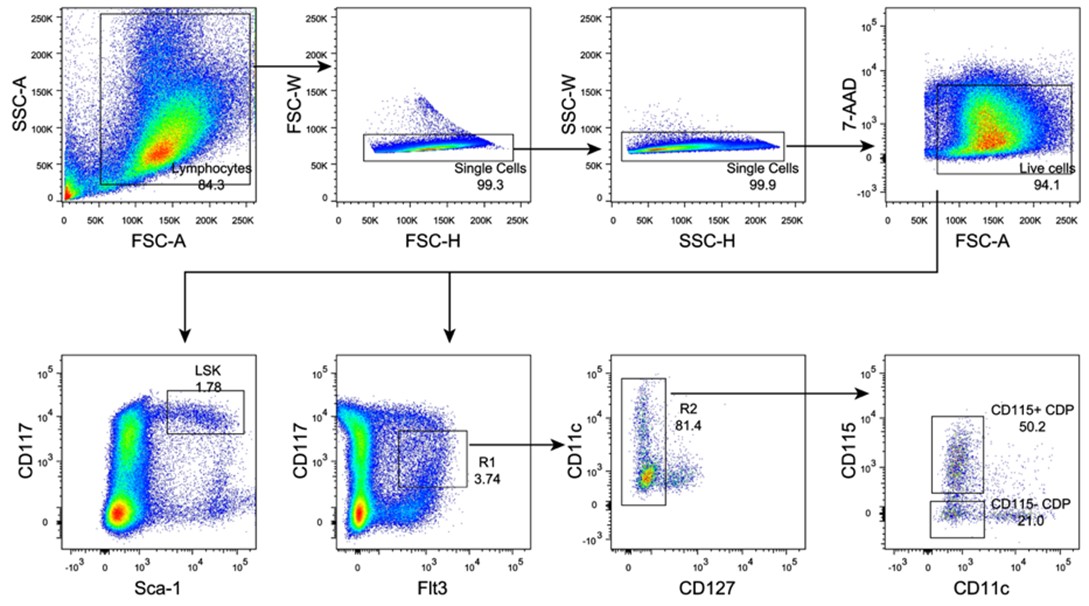
We apologize for not describing the experimental procedures clearly. In this study we enriched the lineage negative (Lin−) cells from the bone marrow using a Lineage-depleting antibody cocktail including mAbs to CD2/CD3/TER-119/Ly6G/B220/CD11b/CD8/CD19. We also provide the gating strategy implemented for sorting LSK and CDP populations from the Lin− cells in the bone marrow (Author response image 5), shown in the Figure 3A and Figure4−figure supplement 1 of revised manuscript.
Author response image 5.
Gating strategy for LSK, CD115+ CDP and CD115− CDP in bone marrow
- Moreover, what is the SiglecH+ CD11c- population appearing in the spleen of mice reconstituted with HDAC3-deleted CDP, in Fig 4D?
We also noticed the appearance of a SiglecH+CD11c− cell population in the spleen of recipient mice reconstituted with HDAC3-deficient CD115−CDPs, while the presence of this population was not as significant in the HDAC3-Ctrl group, as shown in Figure 4D. We speculate that this SiglecH+CD11c− cell population might represent some cells at a differentiation stage earlier than pre-DCs. Alternatively, the relatively increased percentage of this population derived from HDAC3-deficient CD115−CDP might be due to the substantially decreased total numbers of DCs. This could be clarified by further analysis using additional cell surface markers.
- Finally, in Fig 4H, how do the authors explain that Hdac3fl/fl express Il7r, while they are supposed to be sorted CD127- cells?
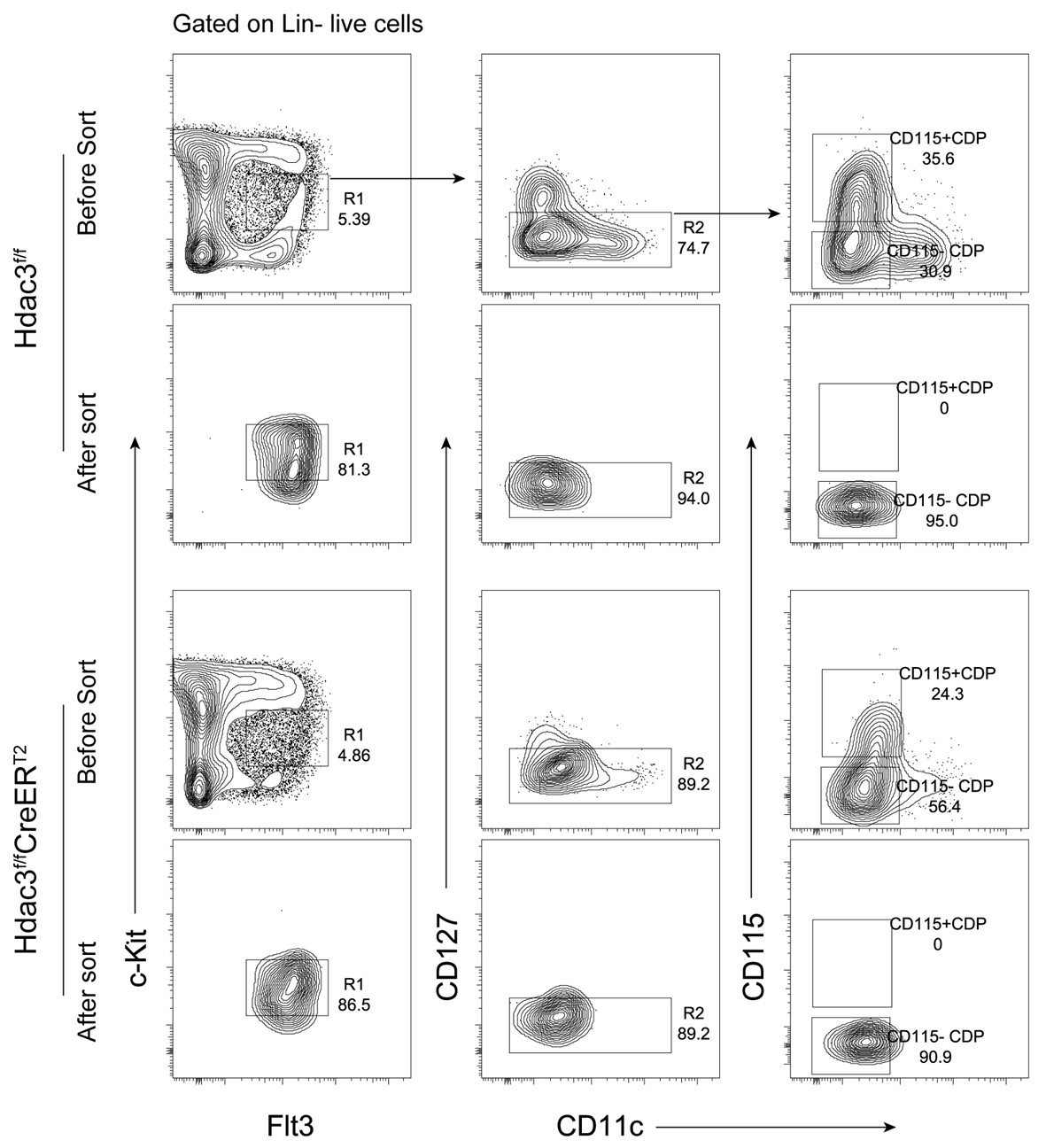
This is indeed an interesting question. In this study, we confirmed that CD115−CDPs were isolated from the surface CD127− cell population for RNA-seq analysis, and the purity of the sorted cells were checked (Author response image 6), as shown in Figure4−figure supplement 1 in revised manuscript.
The possible explanation for the expression of Il7r mRNA in some HDAC3fl/fl CD115−CDPs, as revealed in Figure 4H by RNA-seq analysis, could be due to a very low level of cell surface expression of CD127, these cells therefore could not be efficiently excluded by sorting for surface CD127- cells.
Author response image 6.
CD115−CDPs sorting from Hdac3-Ctrl and Hdac3-KO mice
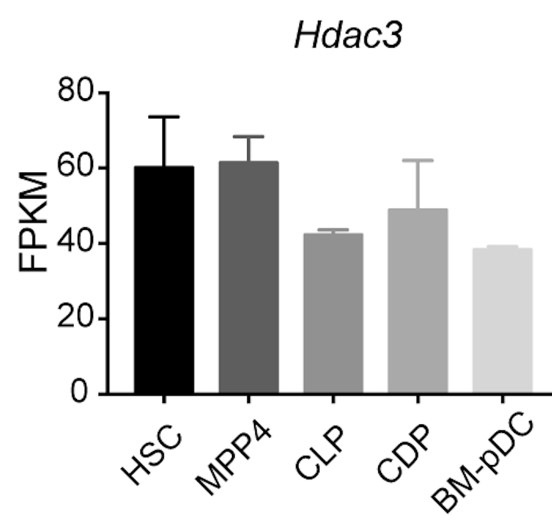
- What is known about the expression of HDAC3 in the different hematopoietic precursors analysed in this study? This information is available only for a few of them in Supplementary Figure 1. If not yet studied, they should be addressed.
We conducted additional analysis to address the expression of Hdac3 in various hematopoietic progenitor cells at different stages, based on the RNA-seq analyis. The data revealed a relatively consistent level of Hdac3 expression in progenitor populations, including HSC, MMP4, CLP, CDP and BM pDCs (Author response image 7). That suggests that HDAC3 may play an important role in the regulation of hematopoiesis at multiple stages. This information is now added in Figure1−figure supplement 1B of revised manuscript.
Author response image 7.
Hdac3 expression in hematopoietic progenitor cells
- It would be highly informative to extend CUT and Tag studies to Irf8 and Tcf4, if this is technically feasible.
We totally agree with the reviewer. We have indeed attempted using CUT and Tag study to compare the binding sites of IRF8 and TCF4 in wild-type and Hdac3-deficient pDCs. However, it proved that this is technically unfeasible to get reliable results due to the limited number of cells we could obtain from the HDAC3 deficient mice. We are committed to explore alternative approaches or technologies in future studies to address this issue.
-

eLife assessment
This study examines the expression of HDAC3 within DC compartment. Taking advantage of tamoxifen inducible ERT2-cre mouse model they observe the dependency of pDCs but not cDCs on HDAC3. The requirement of this histone modifier appears to occur during development around the CLP stage. Tamoxifen treated mice lack almost all pDC besides lymphoid progenitors. RNA seq studies identify multiple DC specific target genes within the remaining pDC - using Cut and Tag technology they validate some of the identified targets of HDAC3. Taken together, this study shows the requirement of HDAC3 on pDC but not cDC, congruent with the recent findings of a lymphoid origin of pDC.
-

Reviewer #1 (Public Review):
The work by Yijun Zhang and Zhimin He at al. analyzes the role of HDAC3 within DC subsets. Using an inducible ERT2-cre mouse model they observe the dependency of pDCs but not cDCs on HDAC3. The requirement of this histone modifier appears to be early during development around the CLP stage. Tamoxifen treated mice lack almost all pDCs besides lymphoid progenitors. Through bulk RNA seq experiment the authors identify multiple DC specific target gens within the remaining pDCs and further using Cut and Tag technology they validate some of the identified targets of HDAC3.
Collectively the study is well executed and shows the requirement of HDAC3 on pDCs but not cDCs, in line with the recent findings of a lymphoid origin of pDC.While the authors provide extensive data on the requirement of HDAC3 within …
Reviewer #1 (Public Review):
The work by Yijun Zhang and Zhimin He at al. analyzes the role of HDAC3 within DC subsets. Using an inducible ERT2-cre mouse model they observe the dependency of pDCs but not cDCs on HDAC3. The requirement of this histone modifier appears to be early during development around the CLP stage. Tamoxifen treated mice lack almost all pDCs besides lymphoid progenitors. Through bulk RNA seq experiment the authors identify multiple DC specific target gens within the remaining pDCs and further using Cut and Tag technology they validate some of the identified targets of HDAC3.
Collectively the study is well executed and shows the requirement of HDAC3 on pDCs but not cDCs, in line with the recent findings of a lymphoid origin of pDC.While the authors provide extensive data on the requirement of HDAC3 within progenitors, the high expression of HDAC3 in mature pDCs may underly a functional requirement. Have you tested INF production in CD11c cre pDCs? Are there transcriptional differences between pDCs from HDAC CD11c cre and WT mice?
A more detailed characterization of the progenitor compartment that is compromised following depletion would be important, as also suggested in the specific points.
-

Reviewer #2 (Public Review):
In this article Zhang et al. report that the Histone Deacetylase-3 (HDAC3) is highly expressed in mouse pDC and that pDC development is severely affected both in vivo and in vitro when using mice harbouring conditional deletion of HDAC3. However, pDC numbers are not affected in Hdac3fl/fl Itgax-Cre mice, indicating that HDCA3 is dispensable in CD11c+ late stages of pDC differentiation. Indeed, the authors provide wide experimental evidence for a role of HDAC3 in early precursors of pDC development, by combining adoptive transfer, gene expression profiling and in vitro differentiation experiments. Mechanistically, the authors have demonstrated that HDAC3 activity represses the expression of several transcription factors promoting cDC1 development, thus allowing the expression of genes involved in pDC …
Reviewer #2 (Public Review):
In this article Zhang et al. report that the Histone Deacetylase-3 (HDAC3) is highly expressed in mouse pDC and that pDC development is severely affected both in vivo and in vitro when using mice harbouring conditional deletion of HDAC3. However, pDC numbers are not affected in Hdac3fl/fl Itgax-Cre mice, indicating that HDCA3 is dispensable in CD11c+ late stages of pDC differentiation. Indeed, the authors provide wide experimental evidence for a role of HDAC3 in early precursors of pDC development, by combining adoptive transfer, gene expression profiling and in vitro differentiation experiments. Mechanistically, the authors have demonstrated that HDAC3 activity represses the expression of several transcription factors promoting cDC1 development, thus allowing the expression of genes involved in pDC development. In conclusion, these findings reveals HDAC3 as a key epigenetic regulator of the expression of the transcription factors required for pDC vs cDC1 developmental fate.
These results are novel and very promising. However, supplementary information and eventual further investigations are required to improve the clarity and the robustness of this article.
Major points
The gating strategy adopted to identify pDC in the BM and in the spleen should be entirely described and shown, at least as a Supplementary Figure. For the BM the authors indicate in the M & M section that they negatively selected cells for CD8a and B220, but both markers are actually expressed by differentiated pDC. However, in the Figures 1 and 2 pDC has been shown to be gated on CD19- CD11b- CD11c+. What is the precise protocol followed for pDC gating in the different organs and experiments?
pDC identified in the BM as SiglecH+ B220+ can actually contain DC precursors, that can express these markers, too. This could explain why the impact of HDAC3 deletion appears stronger in the spleen than in the BM (Figures 1A and 2A). Along the same line, I think that it would important to show the phenotype of pDC in control vs HDAC3-deleted mice for the different pDC markers used (SiglecH, B220, Bst2) and I would suggest to include also Ly6D, taking also in account the results obtained in Figures 4 and 7. Finally, as HDCA3 deletion induces downregulation of CD8a in cDC1 and pDC express CD8a, it would important to analyse the expression of this marker on control vs HDAC3-deleted pDC.
How do the authors explain that in the absence of HDAC3 cDC2 development increased in vivo in chimeric mice, but reduced in vitro (Figures 2B and 2E)? More generally, as reported also by authors (line 207), the reconstitution with HDAC3-deleted cells is poorly efficient. Although cDC seem not to be impacted, are other lymphoid or myeloid cells affected? This should be expected as HDAC3 regulates T and B development, as well as macrophage function. This should be important to know, although this does not call into question the results shown, as obtained in a competitive context.
What are the precise gating strategies used to identify the different hematopoietic precursors in the Figure 4 ? In particular, is there any lineage exclusion performed? Moreover, what is the SiglecH+ CD11c- population appearing in the spleen of mice reconstituted with HDAC3-deleted CDP? Data shown in Figure 4F should be expressed as log2 and not10. Finally, how do the authors explain that Hdac3fl/fl express Il7r, while they are supposed to be sorted CD127- cells?
What is known about the expression of HDAC3 in the different hematopoietic precursors analysed in this study? This information is available only for a few of them in Supplementary Figure 1. If not yet studied, they should be addressed.
It would be highly informative to extend CUT and Tag studies to Irf8 and Tcf4, if this is technically feasible.
-