G-protein-coupled receptor diversity and evolution in the closest living relatives of metazoa
Curation statements for this article:-
Curated by eLife
eLife Assessment
This important study fills a gap in our knowledge of the evolution of GPCRs in holozoans, as well as the phylogeny of associated signaling pathway components such as G proteins, GRKs, and RIC8 proteins. The evidence supporting the conclusions is compelling, with the analysis of extensive new genomic data from choanoflagellates and other non-animal holozoans. Overall, the study is thorough and well-executed. It will be a resource for researchers interested in both the comparative genomics of multicellularity and GPCR biology more broadly, especially given the importance of GPCRs as highly druggable targets
This article has been Reviewed by the following groups
Discuss this preprint
Start a discussion What are Sciety discussions?Listed in
- Evaluated articles (eLife)
Abstract
G-protein-coupled receptors (GPCRs) play a pivotal role in the perception of environmental cues across eukaryotic diversity. Although GPCRs have been relatively well characterized in metazoans, GPCR signaling is poorly understood in their sister group, the choanoflagellates, and in other close relatives of metazoans (CRMs). Here, we examine GPCR diversity and evolution in choanoflagellates by curating a catalog of 918 GPCRs, 141 G proteins, and 367 associated regulators from 23 choanoflagellate genomes and transcriptomes. We found that the repertoire of choanoflagellate GPCRs is larger and more diverse than previously anticipated, with 18 GPCR families found in choanoflagellates, of which 12 families are newly identified in these organisms. Comparative analyses revealed that most choanoflagellate GPCR families are conserved in metazoans and/or other eukaryotic lineages. Adhesion GPCRs and a class of GPCRs fused to kinases (the GPCR-TKL/Ks) are the most abundant GPCRs in choanoflagellates. The identification of GPCR repertoires in CRMs and other non-metazoans refines our understanding of metazoan GPCR evolution and reveals the existence of previously unreported GPCR families in metazoans and at the root of the eukaryotic tree.
Article activity feed
-

-

-

-

eLife Assessment
This important study fills a gap in our knowledge of the evolution of GPCRs in holozoans, as well as the phylogeny of associated signaling pathway components such as G proteins, GRKs, and RIC8 proteins. The evidence supporting the conclusions is compelling, with the analysis of extensive new genomic data from choanoflagellates and other non-animal holozoans. Overall, the study is thorough and well-executed. It will be a resource for researchers interested in both the comparative genomics of multicellularity and GPCR biology more broadly, especially given the importance of GPCRs as highly druggable targets
-

Reviewer #1 (Public review):
Summary:
The authors strived for an inventory of GPCRs and GPCR pathway component genes within the genomes of 23 choanoflagellates and other close relatives of metazoans.
Strengths:
The authors generated a solid phylogenetic overview of the GPCR superfamily in these species. Intriguingly, they discover novel GPCR families, novel assortments of domain combinations, novel insights into the evolution of those groups within the Opisthokonta clade. A particular focus is laid on adhesion GPCRs, for which the authors discover many hitherto unknown subfamilies based on Hidden Markov Models of the 7TM domain sequences, which were also reflected by combinations of extracellular domains of the homologs. In addition, the authors provide bioinformatic evidence that aGPCRs of choanoflagellates also contained a GAIN …
Reviewer #1 (Public review):
Summary:
The authors strived for an inventory of GPCRs and GPCR pathway component genes within the genomes of 23 choanoflagellates and other close relatives of metazoans.
Strengths:
The authors generated a solid phylogenetic overview of the GPCR superfamily in these species. Intriguingly, they discover novel GPCR families, novel assortments of domain combinations, novel insights into the evolution of those groups within the Opisthokonta clade. A particular focus is laid on adhesion GPCRs, for which the authors discover many hitherto unknown subfamilies based on Hidden Markov Models of the 7TM domain sequences, which were also reflected by combinations of extracellular domains of the homologs. In addition, the authors provide bioinformatic evidence that aGPCRs of choanoflagellates also contained a GAIN domain, which are self-cleavable thereby reflecting the most remarkable biochemical feat of aGPCRs.
Weaknesses:
The chosen classification scheme for aGPCRs may require reassessment and amendment by the authors in order to prevent confusion with previously issued classification attempts of this family.
-

Reviewer #2 (Public review):
Summary:
The authors set out to characterise the GPCR family in choanoflagellates (and other unicellular holozoans). GPCRs are the most abundant gene family in many animal genomes, playing crucial roles in a wide range of physiological processes. Although they are known to evolve rapidly, GPCRs are an ancient feature of eukaryotic biology. Identifying conserved elements across the animal-protist boundary is therefore a valuable goal, and the increasing availability of genomes from non-animal holozoans provides new opportunities to explore evolutionary patterns that were previously obscured by limited taxon sampling. This study presents a comprehensive re-examination of GPCRs in choanoflagellates, uncovering examples of differential gene retention and revealing the dynamic nature of the GPCR repertoire in …
Reviewer #2 (Public review):
Summary:
The authors set out to characterise the GPCR family in choanoflagellates (and other unicellular holozoans). GPCRs are the most abundant gene family in many animal genomes, playing crucial roles in a wide range of physiological processes. Although they are known to evolve rapidly, GPCRs are an ancient feature of eukaryotic biology. Identifying conserved elements across the animal-protist boundary is therefore a valuable goal, and the increasing availability of genomes from non-animal holozoans provides new opportunities to explore evolutionary patterns that were previously obscured by limited taxon sampling. This study presents a comprehensive re-examination of GPCRs in choanoflagellates, uncovering examples of differential gene retention and revealing the dynamic nature of the GPCR repertoire in this group. As GPCRs are typically involved in environmental sensing, understanding how these systems evolved may shed light on how our unicellular ancestors adapted their signalling networks in the transition to complex multicellularity.
Strengths:
The paper combines a broad taxonomic scope with the use of both established and recently developed tools (e.g. Foldseek, AlphaFold), enabling a deep and systematic exploration of GPCR diversity. Each family is carefully described, and the manuscript also functions as an up-to-date review of GPCR classification and evolution. Although similar attempts of understanding GPCR evolution were done over the last decade, the authors build on this foundation by identifying new families and applying improved computational methods to better predict structure and function. Notably, the presence of Rhodopsin-like GPCRs in some choanoflagellates and ichthyosporeans is intriguing, even though they do not fall within known animal subfamilies. The computational framework presented here is broadly applicable, offering a blueprint for surveying GPCR diversity in other non-model eukaryotes (and even in animal lineages), potentially revealing novel families relevant to drug discovery or helping revise our understanding of GPCR evolution beyond model systems.
Weaknesses:
While the study contributes several interesting observations, it does not radically revise the evolutionary history of the GPCR family. However, in an era increasingly concerned with the reproducibility of scientific findings, this is arguably a strength rather than a weakness. It is encouraging to see that previously established patterns largely hold, and that with expanded sampling and improved methods, new insights can be gained-especially at the level of specific GPCR subfamilies. Then, no functional follow ups are provided in the model system Salpingoeca rosetta, but I am sure functional work on GPCRs in choanoflagellates is set to reveal very interesting molecular adaptations in the future.
Comments on the latest version:
The authors have done a good job answering my questions and suggestions.
-

Author response:
The following is the authors’ response to the previous reviews
Reviewer #1:
“I am sorry to dwell on the point of naming the newly identified families of adhesion GPCRs in choanoflagellates. I commented: "Can the authors suggest another scheme (mind to avoid the subfamily I-IX or the alternative ADGRA-G,L,V subfamily schemes of metazoan aGPCRs) and adapt their numbering throughout the text and all figures/supplementary figures/supplementary files." Now the authors have changed the Roman numeral numbering (previously used by the adhesion GPCR field to denominate metazoan receptor families) to the other option that I explicitly said should be obsolete, the numbering by capital letters (which is in use since its introduction in 2015 in Hamann et al., Pharmacol Rev, 2015). The authors write: "Phylogenetic analysis of the …
Author response:
The following is the authors’ response to the previous reviews
Reviewer #1:
“I am sorry to dwell on the point of naming the newly identified families of adhesion GPCRs in choanoflagellates. I commented: "Can the authors suggest another scheme (mind to avoid the subfamily I-IX or the alternative ADGRA-G,L,V subfamily schemes of metazoan aGPCRs) and adapt their numbering throughout the text and all figures/supplementary figures/supplementary files." Now the authors have changed the Roman numeral numbering (previously used by the adhesion GPCR field to denominate metazoan receptor families) to the other option that I explicitly said should be obsolete, the numbering by capital letters (which is in use since its introduction in 2015 in Hamann et al., Pharmacol Rev, 2015). The authors write: "Phylogenetic analysis of the 7TM domains of choanoflagellates uncovered at least 19 subfamilies of aGPCRs (subfamilies A-S ...". I am thus afraid this has not addressed my point at all. For example, in the revised numbering scheme for Choanoflagellates aGPCR subfamilies of the authors the now used "A" descriptor, which are predicted to contain a HYR domain, can be mistaken for ADGRA homologs (abbreviated as "A" receptors, previously termed subfamily III aGPCRs) of metazoan aGPCRs, which contain HRM and LRR domains. Likewise, choanoflagellate "E" receptors are predicted to harbour LRR repeats, but metazoan ADGRE (abbreviated as "E" too) are characterised by their EGF domains. This clearly underlines the need to devise a numbering scheme for the newly described choanoflagellate aGPCR homologs so they cannot be confused with the receptors from other kingdoms, for which identical naming conventions exist. Please change this, e.g. by numbering/denominating the choanoflagellate subfamilies by greek letters (or your pick of any other ordering system that does not lend itself to be mistaken with the previous and existing aGPCR classifications) and change the manuscript and figures accordingly.”
We have now re-labeled the choanoflagellate aGPCR subfamilies, previously numbered from A to S, using Greek alphabetical enumeration (from α to τ). Changes have been made throughout the main text, in Figure 5, and in Supplementary Figures S6 and S7.
-

-

eLife Assessment
This important study fills a gap in our knowledge of the evolution of GPCRs in holozoans, as well as the phylogeny of associated signaling pathway components such as G proteins, GRKs, and RIC8 proteins. The evidence supporting the conclusions is compelling, with the analysis of extensive new genomic data from choanoflagellates and other non-animal holozoans. Overall, the study is thorough and well-executed. It will be a resource for researchers interested in both the comparative genomics of multicellularity and GPCR biology more broadly, especially given the importance of GPCRs as highly druggable targets.
-

Reviewer #1 (Public review):
Summary:
The authors strived for an inventory of GPCRs and GPCR pathway component genes within the genomes of 23 choanoflagellates and other close relatives of metazoans.Strengths:
The authors generated a solid phylogenetic overview of the GPCR superfamily in these species. Intriguingly, they discover novel GPCR families, novel assortments of domain combinations, novel insights into the evolution of those groups within the Opisthokonta clade. A particular focus is laid on adhesion GPCRs, for which the authors discover many hitherto unknown subfamilies based on Hidden Markov Models of the 7TM domain sequences, which were also reflected by combinations of extracellular domains of the homologs. In addition, the authors provide bioinformatic evidence that aGPCRs of choanoflagellates also contained a GAIN …Reviewer #1 (Public review):
Summary:
The authors strived for an inventory of GPCRs and GPCR pathway component genes within the genomes of 23 choanoflagellates and other close relatives of metazoans.Strengths:
The authors generated a solid phylogenetic overview of the GPCR superfamily in these species. Intriguingly, they discover novel GPCR families, novel assortments of domain combinations, novel insights into the evolution of those groups within the Opisthokonta clade. A particular focus is laid on adhesion GPCRs, for which the authors discover many hitherto unknown subfamilies based on Hidden Markov Models of the 7TM domain sequences, which were also reflected by combinations of extracellular domains of the homologs. In addition, the authors provide bioinformatic evidence that aGPCRs of choanoflagellates also contained a GAIN domain, which are self-cleavable thereby reflecting the most remarkable biochemical feat of aGPCRs.Weaknesses:
The chosen classification scheme for aGPCRs may require reassessment and amendment by the authors in order to prevent confusion with previously issued classification attempts of this family. -

Reviewer #2 (Public review):
Summary:
The authors set out to characterise the GPCR family in choanoflagellates (and other unicellular holozoans). GPCRs are the most abundant gene family in many animal genomes, playing crucial roles in a wide range of physiological processes. Although they are known to evolve rapidly, GPCRs are an ancient feature of eukaryotic biology. Identifying conserved elements across the animal-protist boundary is therefore a valuable goal, and the increasing availability of genomes from non-animal holozoans provides new opportunities to explore evolutionary patterns that were previously obscured by limited taxon sampling. This study presents a comprehensive re-examination of GPCRs in choanoflagellates, uncovering examples of differential gene retention and revealing the dynamic nature of the GPCR repertoire in …Reviewer #2 (Public review):
Summary:
The authors set out to characterise the GPCR family in choanoflagellates (and other unicellular holozoans). GPCRs are the most abundant gene family in many animal genomes, playing crucial roles in a wide range of physiological processes. Although they are known to evolve rapidly, GPCRs are an ancient feature of eukaryotic biology. Identifying conserved elements across the animal-protist boundary is therefore a valuable goal, and the increasing availability of genomes from non-animal holozoans provides new opportunities to explore evolutionary patterns that were previously obscured by limited taxon sampling. This study presents a comprehensive re-examination of GPCRs in choanoflagellates, uncovering examples of differential gene retention and revealing the dynamic nature of the GPCR repertoire in this group. As GPCRs are typically involved in environmental sensing, understanding how these systems evolved may shed light on how our unicellular ancestors adapted their signalling networks in the transition to complex multicellularity.Strengths:
The paper combines a broad taxonomic scope with the use of both established and recently developed tools (e.g. Foldseek, AlphaFold), enabling a deep and systematic exploration of GPCR diversity. Each family is carefully described, and the manuscript also functions as an up-to-date review of GPCR classification and evolution. Although similar attempts of understanding GPCR evolution were done over the last decade, the authors build on this foundation by identifying new families and applying improved computational methods to better predict structure and function. Notably, the presence of Rhodopsin-like GPCRs in some choanoflagellates and ichthyosporeans is intriguing, even though they do not fall within known animal subfamilies. The computational framework presented here is broadly applicable, offering a blueprint for surveying GPCR diversity in other non-model eukaryotes (and even in animal lineages), potentially revealing novel families relevant to drug discovery or helping revise our understanding of GPCR evolution beyond model systems.Weaknesses:
While the study contributes several interesting observations, it does not radically revise the evolutionary history of the GPCR family. However, in an era increasingly concerned with the reproducibility of scientific findings, this is arguably a strength rather than a weakness. It is encouraging to see that previously established patterns largely hold, and that with expanded sampling and improved methods, new insights can be gained-especially at the level of specific GPCR subfamilies. Then, no functional follow ups are provided in the model system Salpingoeca rosetta, but I am sure functional work on GPCRs in choanoflagellates is set to reveal very interesting molecular adaptations in the future.Comments on the latest version:
The authors have done a good job answering my questions and suggestions.
-

Author response:
The following is the authors’ response to the original reviews
Reviewer #1:
The chosen classification scheme for aGPCRs may require reassessment and amendment by the authors in order to prevent confusion with previously issued classification attempts of this family. (…) Can the authors suggest another scheme (mind to avoid the subfamily IIX or the alternative ADGRA-G,L,V subfamily schemes of metazoan aGPCRs), and adapt their numbering throughout the text and all figures/supplementary figures/supplementary files?
We appreciate the reviewer's comment and agree that a different nomenclature should be used for choanoflagellate aGPCRs to avoid possible confusion. We have now re-labeled the choanoflagellate aGPCR subfamilies, previously numbered from I to XIX, using alphabetical enumeration (from A to S). Changes have been …
Author response:
The following is the authors’ response to the original reviews
Reviewer #1:
The chosen classification scheme for aGPCRs may require reassessment and amendment by the authors in order to prevent confusion with previously issued classification attempts of this family. (…) Can the authors suggest another scheme (mind to avoid the subfamily IIX or the alternative ADGRA-G,L,V subfamily schemes of metazoan aGPCRs), and adapt their numbering throughout the text and all figures/supplementary figures/supplementary files?
We appreciate the reviewer's comment and agree that a different nomenclature should be used for choanoflagellate aGPCRs to avoid possible confusion. We have now re-labeled the choanoflagellate aGPCR subfamilies, previously numbered from I to XIX, using alphabetical enumeration (from A to S). Changes have been made throughout the main text, in Figure 5, and in Supplementary Figures S6 and S7.
line 10: The abbreviation 'GPCR-TKL/Ks' is not explained.
Thank you for pointing this out. We have now revised the text to explain the abbreviation:
“Adhesion GPCRs and a class of GPCRs fused to kinases (the GPCR-TKL/Ks) are the most abundant GPCRs in choanoflagellates.”
line 30: "7TM domain is diagnostic for GPCRs": strange wording. Use an alternative expression.
We changed the wording to:
“A conserved seven transmembrane (7TM) domain is a hallmark of GPCRs, while the wide spectrum of extracellular and intracellular domains in some GPCRs reflects the diversification of the gene family and its functions (Schiöth and Lagerström 2008).”
line 33: In the case of rhodopsins, not the GPCR (i.e., the apoprotein) responds directly to photons, but the retinal, which isomerises upon illumination.
We thank the reviewer for bringing this to our attention, and we have now removed mention of photons from the list of cues detected by GPCRs.
“For example, the extracellular N-terminus and the three extracellular loops of the 7TM domain respond to a wide range of cues, including odorant molecules, peptides, amines, lipids, nucleotides, and other molecules (Yang et al. 2021).”
line 111: What are "genome-enabled choanoflagellates"? Explain the term. As it stands, it doesn't make sense to me.
We meant only to highlight that these two species have sequenced genomes. We have deleted the phrase “genome enabled.”
“To assess the predictive power of our protein-detection pipeline, we then compared the new GPCR and cytosolic signaling component datasets from two choanoflagellates – Salpingoeca rosetta and Monosiga brevicollis – with previously published GPCR and downstream GPCR signaling component counts for these two species (Nordström et al. 2009a; Krishnan et al. 2012; De Mendoza et al. 2014; Krishnan et al. 2015; Lokits et al. 2018).”
line 145: Please give a reasoning for the naming of each of the new families (e.g., RemiSens, Hidden Gold, GPCR-TLK/K, etc.) or at least the explanations of the acronyms/names early in the manuscript, even if they are discussed later in more detail.
Thank you for identifying this as an area of confusion. While we feel that going into the rationale behind each of the names here would interrupt the flow of the manuscript, we have added a phrase encouraging readers to “hold that thought” with the hope that they can wait for the sections that specifically focus on each of these new GPCR families.
“This left twelve new GPCR families that had not, to our knowledge, been previously detected in choanoflagellates: Rhodopsin, TMEM145, GPR180, TMEM87, GPR155, GPR157, and six additional GPCR families that appear to fall outside all previously characterized GPCR families in eukaryotes. For reasons that will be discussed further below, we have named these six new GPCR families “Rémi-Sans-Famille” (RSF), “Hidden Gold” (Hi-GOLD), GPCR-TKL/K, GPRch1, GPRch2, and GPRch3. (Fig. 1B; Table 1).”
lines 297/298 and 2049: Rename tethered agonist "peptide" to "element". Synthetic peptides resembling the TA were used in experiments to test for the sufficiency of the TA for receptor activation, but because the naturally occurring TAs are part of the receptor protein, they are not peptides.
Thank you for pointing this out. We have revised the text as suggested.
line 2026: I think the letters in the acronym "CMR" are mixed up and were intended to read "CRM".
Good catch! We have corrected the text.
line 2048: "diagnostic" again. Change to "tell-tale", "hallmark", or another similar descriptor.
We have corrected the text accordingly.
2058: Strike "motif" in order to avoid confusion with the now obsolete term "GPS motif", which entailed the five most C-terminal β-strands of GAIN subdomain B (not thus neither the full GAIN domain nor the GPS).
Thank you for pointing this out. We have corrected the text.
Figure 5: Did the authors also find homologs placed in the aGPCR family based on their 7TM domain sequence but lacking a GAIN domain similar to vertebrate ADGRA/GPR123, the only aGPCR known to lack a GAIN domain (10.1016/j.tips.2013.06.002)? Irrespective of the authors' findings or non-finding on that matter, please insert a note on this in the results text.
We thank the reviewer for bringing this interesting point to our attention. We have now added a new supplementary figure A in Fig. S9 to answer the reviewer's comment. We also modified the legend of Fig. S9 to take into account this change and uploaded a new supplementary data file 20 to support Fig. S9A. Finally, we revised the main text under the section “Adhesion GPCRs” as requested:
Lines 328-331: “ While the GAIN and aGPCR 7TM domains evolved before the origin of opisthokonts (Araç et al.2012; Krishnan et al. 2012; De Mendoza et al. 2014), we detected the fusion of these two domains into a single module (GAIN/7TM) in most, but not all, holozoan aGPCRs (Fig. 5D, Fig.S7B and S9A; Supplementary file 20; Prömel et al, 2013; Krishnan et al. 2014).
Reviewer #2:
While the study contributes several interesting observations, it does not radically revise the evolutionary history of the GPCR family. However, in an era increasingly concerned with the reproducibility of scientific findings, this is arguably a strength rather than a weakness. It is encouraging to see that previously established patterns largely hold, and that with expanded sampling and improved methods, new insights can be gained, especially at the level of specific GPCR subfamilies. Then, no functional follow-ups are provided in the model system Salpingoeca rosetta, but I am sure functional work on GPCRs in choanoflagellates is set to reveal very interesting molecular adaptations in the future.
We agree with the reviewer and anticipate that this work will provide a useful resource to motivate the future functional characterization of GPCRs in choanoflagellates, other CRMs, as well as in metazoans.
The GPCR-TKL fusion is a particularly interesting finding, especially given the presence of such sequences in sponges. This could potentially represent a synapomorphy shared between sponges and choanoflagellates, later lost in other animals. The authors mention that BLASTP searches using the kinase domain recover the sponge GPCR-TKLs, suggesting the fusion may be ancestral. It would be useful to include phylogenetic trees of both the GPCR and TKL domains to assess this possibility. The authors might also consider examining sponge genomes released by the DTOL project to increase representation from this group.
We agree and thank the reviewer for this suggestion. We have now added the requested phylogenetic analyses to the new Figure S17, revised the supplementary files and Methods accordingly, and commented on these results in the main text under the section “GPCR-TKL/K and GPCR-TKs“.
Lines 579 – 589: “While no metazoan homologs were found when using the 7TM domain of choanoflagellate GPCR-TKs as queries, using the conserved tyrosine kinase domains as queries recovered GPCR-TKs in sponges but not in other metazoan lineages or other holozoans (Fig. S17E). To test whether GPCR-TKs in sponges and choanoflagellates are homologous, we performed phylogenetic analyses of their TK and 7TM domains (Fig. S17F and G). While the TK domains of GPCR-TKs from sponges and choanoflagellates formed a well-supported clade, their 7TM domains did not. These results point to a heterogeneous evolutionary history that may include domain swapping (i.e. ancestral GPCR-TKs in which the 7TM domain was replaced in either the sponge or choanoflagellate lineages) or convergent evolution, in which homologous 7TM domains fused with unrelated 7TM domains in the sponge and choanoflagellate lineages.”
Added to the Method section “Sequence alignment and phylogenetic analyses”:
Lines 913 – 933: “Phylogenetic analyses of holozoan aGPCRs, Glutamate Receptors, and Gα subunits, and the 7TM and Kinase domains from GPCR TK/TKL/Ks were performed in this study. (…) To construct the phylogenies of the Kinase domain and 7TM domain from the GPCR TK/TKL/Ks, we first built a dataset including all the GPCR TK/TKL/Ks sequences identified in choanoflagellates and in sponges, as well as the GPCR TKL/Ks previously published in oomycetes and amoebozoans (Van Den Hoogen et al. 2018). We extracted the 7TM domain and Kinase domain from each sequence by combining the transmembrane domain prediction tool TMHMM-2.0 and the protein domain prediction tool InterProScan with the alignment tool MAFFT (E-INS-I algorithm) on Geneious Prime v2024.07 (Supplementary Files 30 and 32). We then aligned the aGPCR, Glutamate and Glutamate GPCR TK/TKL/K Receptor 7TMs, the GPCR TK/TKL/Ks Kinase domain, or the full-length Gα sequences using MAFFT with the E-INS-I algorithm. The resulting alignments were then used for Maximum-likelihood and/or Bayesian inference of phylogenies (Fig. 3B, Fig. 5A, Fig. S3D, and Fig. S6A, and Fig. S17F and G; Supplementary Files 5, 9, 16,18, 31, and 33).”
Rhodopsin-like receptors are proposed in the discussion to be potential cases of lateral gene transfer (LGT) between eukaryotes. To support or refute this hypothesis, it would be valuable to place the choanoflagellate and ichthyosporean Rhodopsins within a broader phylogeny of this family, including (a few) representatives from animals and other eukaryotes. Even if deep branching relationships remain unresolved, signs such as unusually short branches could point toward recent LGT events.
Thank you for your suggestion. While we originally considered testing these alternative hypotheses in this manuscript by building a phylogeny, the rapid sequence evolution of the Rhodopsin family has stymied similar efforts in the past and instead motivated others to use clustering approaches like those used in our study (Hu et al. 2017; Thiel et al. 2023). Unfortunately, these types of analyses cannot be used to readily identify instances of LGT.
Therefore, following the suggestion of the reviewer, we bit the bullet and performed phylogenetic analyses on the sequences in question. Unfortunately, these analyses were completely inconclusive, and we feel they do not warrant inclusion in the manuscript. The topologies of the sequence trees recovered were poorly supported and sensitive to most of the variables we tested – the set of rhodopsin sequences included, the multiple alignment algorithms used, and the probabilistic methods employed to infer the phylogenies.
Instead, we have revised the manuscript to highlight the challenge of differentiating between the different hypotheses that are consistent with the phylogenetic distribution of Rhodopsins:
Lines 670 – 678: “Thus, while it is formally possible that Rhodopsins existed in stem choanoflagellates and were lost in most modern choanoflagellate lineages, either horizontal gene transfer or convergent evolution in the shared ancestor of S. macrocollata and S. punica are similarly plausible explanations for their presence in these species. Differentiating between these alternative evolutionary scenarios is challenging because of rapid rate of sequence evolution within the family and the resultant loss of phylogenetic signal. Our own preliminary investigations of Rhodopsin evolution in non-metazoans were inconclusive. Therefore, ambiguities about the provenance and function of CRM Rhodopsins currently obscure the ancestry of metazoan Rhodopsins and opsins.”
While the study surveys most available holozoan genomes, it appears that the genomes of Amoebidium spp.-which are cited in the manuscript- were not included. It may not be necessary to repeat all analyses with these two species (A. appalachense and A. parasiticum), but a preliminary search indicates the presence of four candidate 7tm_1 (Rhodopsin-like) proteins in their proteomes. These may warrant closer inspection (e.g., via BLASTP against animal databases) to confirm whether they are genuine GPCRs or false positives.
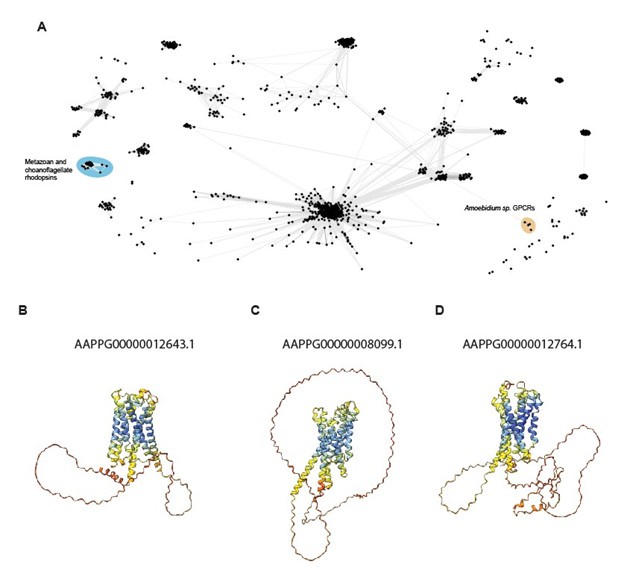
Author response image 1.
We thank the reviewer for bringing these sequences to our attention. To be clear, we did not analyze the Amoebidium spp. genome and we can find no reference to it in our manuscript. If the reviewer had the impression that the genome was analyzed, we would be grateful to know the source of the confusion so that it can be corrected. (We did not intentionally exclude the genome; it simply was not available on the Multicell Genome database from which we retrieved the ichthyosporean genomes and transcriptomes used in this study.)
Nevertheless, out of curiosity, we have now analyzed the sequences provided by the reviewer and summarize our findings here for the interest of the reviewer. Although the sequences were annotated as 7tm_1 (Rhodopsin-like) proteins in the original genome study, none of these sequences group with metazoan or choanoflagellate Rhodopsins in our clustering analysis; instead, we found that these putative GPCRs form a distinct cluster that only weakly resembles cAMP receptors, both on the basis of their sequence and predicted structures.
It is not surprising to find new GPCR clusters as new taxa are folded into the study, and these Amoebidium sequences do not add to our understanding of Rhodopsin evolution. Therefore, we have not added their analysis to the manuscript, but we hope the reviewer finds our quick analysis of interest.
Author response image 2.
In Figure 2, perhaps expanding the other holozoan clades would have been nice, as there are not too many species, but I understand if that's beyond the point of the manuscript, focused on choanoflagellates.
Thank you for this comment. However, given the focus of this study, we feel that an expansion of the other holozoan clades would reduce the clarity of the figure.
line 87 - "To this end, the 671 validated choanoflagellate GPCRs were sorted by sequence similarity, resulting in 18 clusters. "Some details in the results section would be nice, or at least clear references to where this is explained in more detail. How were the extra choanoflagellate GPCRs added if they failed to be identified with quite sensitive HMM profiles?
We apologize for the possible confusion and thank the reviewer for the suggestion; we have now added specific references to the related sections from the material and methods for interested readers.
We believe that the "extra choanoflagellate GPCRs" mentioned by the reviewer refer to the choanoflagellate GPCRs that failed to be detected when the choanoflagellate genomes and transcriptomes were searched with the predominantly metazoan-derived GPCRHMM and HMMs from the GPCR_A Pfam clan (CL0192). We were able to recover these extra choanoflagellate GPCRs by using custom choanoflagellate-specific GPCR HMMs and by blasting the choanoflagellate GPCRs previously identified as queries against the 23 choanoflagellate proteomes. We hope that the referencing of the Methods section "Recovering additional choanoflagellate GPCRs using choanoflagellate GPCR BLAST queries and custom choanoflagellate GPCR HMMs", in lines 91 and 93, will help clarify this point.
line 108 - Well, from the figure it seems that most eukaryotes have an 'animal-like' G protein signalling, so that's perhaps more of an eukaryotic signature than something that puts choanoflagellates and animals together.
Excellent point! We have revised the text.
line 132 - It is unclear what the criteria are to include these taxa as helpers for choanoflagellate classification, and not adding the other unicellular holozoans. Just some text justification could help.
Thank you for pointing this out. We have added an explanation of the rationale to the methods — section “Clustering of the 918 validated choanoflagellate GPCRs” — and referred to it in the main text.
New text added to methods:
“The non-choanoflagellate sequences added to the dataset were either top blast hits recovered after searching the entire Eukprot v3 dataset (993 species) with choanoflagellate GPCRs as queries, or previously published and well-documented GPCR sequences from metazoans.”
line 145 - These families are listed, but perhaps it would be nice to explicitly mention that they will be covered in more detail later on in the manuscript. I found myself wondering about those exotic names, until I reached the sections in the manuscript where they are explained.
Thank you for this suggestion. We have now modified our sentence to refer to the related sections.
“For reasons that will be discussed further below, we have named these six new GPCR families “Rémi-Sans-Famille” (RSF), “Hidden Gold” (Hi-GOLD), GPCR-TKL/K, GPRch1, GPRch2, and GPRch3. (Fig. 1B; Table 1).”
line 199 - perhaps would be nice to explain domain architecture of validated Dictyostelium GABA-like receptors (ANF domain?).
Thank you for your suggestion. We have now modified the sentence to mention the protein domain composition of the validated GABA-like receptor, GrlE, in Dictyostelium.
“The Glutamate Receptors from the amoebozan Dictyostelium discoideum, of which at least one, GrlE, binds both GABA and Glutamate presumably through its conserved ANF domain (Anjard and Loomis 2006; Taniura et al. 2006; Wu and Janetopoulos 2013), grouped separately from metazoan and CRM GPCRs in our analysis.”
Figure S4 - Perhaps a stacked bar chart would be easier to browse than a bunch of pie charts, notoriously difficult to quantify.
Thank you for this comment. Opinions differ on how best on whether pie charts or bar charts are more effective in this context (including between the authors of this manuscript). However, we think the point of Figure S4 a minor point, only to be appreciated by a tiny number of readers, and therefore have left the data presentation as it was in the original submission.
-

-

eLife Assessment
This important study fills an gap in our knowledge of the evolution of GPCRs in holozoans, as well as the phylogeny of associated signaling pathway components such as G proteins, GRKs, and RIC8 proteins. The evidence supporting the conclusions is compelling, with the analysis of extensive new genomic data from choanoflagellates and other non-animal holozoans. Overall, the study is thorough and well-executed. It will be a resource for researchers interested in both the comparative genomics of multicellularity and GPCR biology more broadly, especially given the importance of GPCRs as highly druggable targets.
-

Reviewer #1 (Public review):
Summary:
The authors strived for an inventory of GPCRs and GPCR pathway component genes within the genomes of 23 choanoflagellates and other close relatives of metazoans.
Strengths:
The authors generated a solid phylogenetic overview of the GPCR superfamily in these species. Intriguingly, they discover novel GPCR families, novel assortments of domain combinations, and novel insights into the evolution of those groups within the Opisthokonta clade. A particular focus is laid on adhesion GPCRs, for which the authors discover many hitherto unknown subfamilies based on Hidden Markov Models of the 7TM domain sequences, which were also reflected by combinations of extracellular domains of the homologs. In addition, the authors provide bioinformatic evidence that aGPCRs of choanoflagellates also contain a GAIN …
Reviewer #1 (Public review):
Summary:
The authors strived for an inventory of GPCRs and GPCR pathway component genes within the genomes of 23 choanoflagellates and other close relatives of metazoans.
Strengths:
The authors generated a solid phylogenetic overview of the GPCR superfamily in these species. Intriguingly, they discover novel GPCR families, novel assortments of domain combinations, and novel insights into the evolution of those groups within the Opisthokonta clade. A particular focus is laid on adhesion GPCRs, for which the authors discover many hitherto unknown subfamilies based on Hidden Markov Models of the 7TM domain sequences, which were also reflected by combinations of extracellular domains of the homologs. In addition, the authors provide bioinformatic evidence that aGPCRs of choanoflagellates also contain a GAIN domain, which is self-cleavable, thereby reflecting the most remarkable biochemical feat of aGPCRs.
Weaknesses:
The chosen classification scheme for aGPCRs may require reassessment and amendment by the authors in order to prevent confusion with previously issued classification attempts of this family.
-

Reviewer #2 (Public review):
Summary:
The authors set out to characterise the GPCR family in choanoflagellates (and other unicellular holozoans). GPCRs are the most abundant gene family in many animal genomes, playing crucial roles in a wide range of physiological processes. Although they are known to evolve rapidly, GPCRs are an ancient feature of eukaryotic biology. Identifying conserved elements across the animal-protist boundary is therefore a valuable goal, and the increasing availability of genomes from non-animal holozoans provides new opportunities to explore evolutionary patterns that were previously obscured by limited taxon sampling. This study presents a comprehensive re-examination of GPCRs in choanoflagellates, uncovering examples of differential gene retention and revealing the dynamic nature of the GPCR repertoire in …
Reviewer #2 (Public review):
Summary:
The authors set out to characterise the GPCR family in choanoflagellates (and other unicellular holozoans). GPCRs are the most abundant gene family in many animal genomes, playing crucial roles in a wide range of physiological processes. Although they are known to evolve rapidly, GPCRs are an ancient feature of eukaryotic biology. Identifying conserved elements across the animal-protist boundary is therefore a valuable goal, and the increasing availability of genomes from non-animal holozoans provides new opportunities to explore evolutionary patterns that were previously obscured by limited taxon sampling. This study presents a comprehensive re-examination of GPCRs in choanoflagellates, uncovering examples of differential gene retention and revealing the dynamic nature of the GPCR repertoire in this group. As GPCRs are typically involved in environmental sensing, understanding how these systems evolved may shed light on how our unicellular ancestors adapted their signalling networks in the transition to complex multicellularity.
Strengths:
The paper combines a broad taxonomic scope with the use of both established and recently developed tools (e.g., Foldseek, AlphaFold), enabling a deep and systematic exploration of GPCR diversity. Each family is carefully described, and the manuscript also functions as an up-to-date review of GPCR classification and evolution. Although similar attempts to understand GPCR evolution were made over the last decade, the authors build on this foundation by identifying new families and applying improved computational methods to better predict structure and function. Notably, the presence of Rhodopsin-like GPCRs in some choanoflagellates and ichthyosporeans is intriguing, even though they do not fall within known animal subfamilies. The computational framework presented here is broadly applicable, offering a blueprint for surveying GPCR diversity in other non-model eukaryotes (and even in animal lineages), potentially revealing novel families relevant to drug discovery or helping revise our understanding of GPCR evolution beyond model systems.
Weaknesses:
While the study contributes several interesting observations, it does not radically revise the evolutionary history of the GPCR family. However, in an era increasingly concerned with the reproducibility of scientific findings, this is arguably a strength rather than a weakness. It is encouraging to see that previously established patterns largely hold, and that with expanded sampling and improved methods, new insights can be gained, especially at the level of specific GPCR subfamilies. Then, no functional follow-ups are provided in the model system Salpingoeca rosetta, but I am sure functional work on GPCRs in choanoflagellates is set to reveal very interesting molecular adaptations in the future.
-