Multifaceted Functional Complexity of SARS-CoV-2 Helicase Nsp13 Underlies Its Integrated Motor and Remodeling Activities
Curation statements for this article:-
Curated by eLife
eLife Assessment
This study characterizes several novel activities of SARS-CoV-2 helicase nsp13, providing valuable insights into potentially new functions of this essential RNA-processing enzyme in the virus life cycle. However, the experimental evidence to support the authors' claims is incomplete. In addition, the placement of the polyhistidine affinity tag on nsp13 may cause artifacts, raising concerns about the interpretation of the results.
This article has been Reviewed by the following groups
Discuss this preprint
Start a discussion What are Sciety discussions?Listed in
- Evaluated articles (eLife)
Abstract
SARS-CoV-2 nonstructural protein 13 (Nsp13) is a superfamily 1 helicase essential for viral replication. Although its canonical ATP-dependent unwinding activity is well established, the broader functional repertoire of Nsp13 remains unclear. Here, we show that Nsp13 encodes a high degree of complexity by acting as a tunable nucleic acid remodeler that integrates motor and non-motor activities within a single protein. Nsp13 operates in multiple mechanistically distinct regimes, including a canonical ATP-dependent helicase mode and a Mg²⁺-primed, ATP-independent remodeling state capable of destabilizing short duplexes, hairpins, and G-quadruplexes. Mg²⁺ binding allosterically stabilizes a compact RecA1–RecA2 configuration, priming the enzyme for ATP-independent remodeling. Unwinding polarity is substrate-dependent, with duplexes supporting bidirectional remodeling, whereas G-quadruplexes are preferentially resolved in the 5′→3′ direction. Beyond strand separation, Nsp13 also exhibits robust strand annealing and nucleic acid chaperone activities. Cofactors, substrate topology, and enzyme concentration dynamically regulate these activities. Together, our findings establish Nsp13 as a highly integrated nucleic acid remodeling system and reveal how a single viral helicase switches between motor-driven and remodeling-dominated states to meet the structural demands of replication and transcription.
Article activity feed
-

eLife Assessment
This study characterizes several novel activities of SARS-CoV-2 helicase nsp13, providing valuable insights into potentially new functions of this essential RNA-processing enzyme in the virus life cycle. However, the experimental evidence to support the authors' claims is incomplete. In addition, the placement of the polyhistidine affinity tag on nsp13 may cause artifacts, raising concerns about the interpretation of the results.
-

Reviewer #1 (Public review):
In the manuscript by Li et al., the authors perform a comprehensive study on the template and cofactor determinants of the SARS-CoV-2 nsp13 protein. They find that, alongside the classical processive unwinding ability of helicases driven by ATP consumption, other chaperone-like and ATP-independent functions exist for this enzyme. By testing DNA and RNA oligos in several conformations, the authors show that these functions are highly dependent on template identity, but also on the ratio of ATP to divalent cations. Ultimately, it is suggested that these distinct mechanisms of action are employed by nsp13 to orchestrate viral replication.
Overall, this study provides some novel insights into the functionality of a central and conserved enzyme of a relevant human pathogenic virus. While the approach is important …
Reviewer #1 (Public review):
In the manuscript by Li et al., the authors perform a comprehensive study on the template and cofactor determinants of the SARS-CoV-2 nsp13 protein. They find that, alongside the classical processive unwinding ability of helicases driven by ATP consumption, other chaperone-like and ATP-independent functions exist for this enzyme. By testing DNA and RNA oligos in several conformations, the authors show that these functions are highly dependent on template identity, but also on the ratio of ATP to divalent cations. Ultimately, it is suggested that these distinct mechanisms of action are employed by nsp13 to orchestrate viral replication.
Overall, this study provides some novel insights into the functionality of a central and conserved enzyme of a relevant human pathogenic virus. While the approach is important and adds to the field, particularly by characterizing the chaperoning activities and adding G-quadruplexes as templates, previous studies have already identified several determinants of nsp13 template binding and processing in vitro (Sommers et al., 2023, JBC; Park et al., 2025, JBC). In addition, some issues regarding experimental design need to be addressed to increase the cogency and biological relevance of the study.
(1) Generally, low concentrations of monovalent cations (20 mM), as used throughout this study, may influence helicase activity and artificially enhance protein binding/oligomerization, which could favor the observed chaperoning activity (Venus et al., 2022, Methods). In contrast, some helicases, such as HCV NS3, are inhibited by higher K+ concentrations (Gwack et al., 2004, FEBS). Thus, the influence of higher concentrations of monovalent cations should be tested in relevant assays, as intracellular K+ levels are usually >100 mM. Additionally, this could significantly affect template stability. For instance, in some G4 assays, the addition of the trap already leads to observable duplex formation (Figure 5), which may be due to low K+ conditions.
(2) As in most publications that focus strictly on helicase (or other enzymatic) functions, the activity of the isolated protein is examined. However, particularly in the case of nsp13, core functions rely on other factors, such as nsp7/8 and other components of the replication-transcription complex (RTC). The overall structure and oligomerization state of nsp13 are altered within the complex (Chen et al., 2022, NSMB). The inclusion of such factors in key experiments would greatly improve the biological relevance of the findings.
(3) In Figure 4, the authors claim that Mg2+ concentration inhibits RNA unwinding. While this is likely considering previous findings, it must be validated that duplex stabilization is not the primary cause for the observed lower dissociation rates. As the template is only 12 bp long with extensive overhangs, higher ion concentrations may significantly stabilize base pairing by reducing fraying effects. Similarly, in Figure 6, template-dependent effects of Mg2+/ATP should be ruled out.
(4) It is not entirely clear to me by which principle the templates were chosen. In my opinion, it would improve the overall comparability of the experimental results if, for instance, the blunt-ended duplex had the same sequence as the oligos with overhangs, since factors such as length, G/C content, Tm, etc., may play a significant role in binding and unwinding. Similarly, the oligos for binding and unwinding should be kept somewhat comparable, e.g., the G4 for the binding assay has 3 stacks, whereas RG1 has only 2. This discrepancy could make a significant difference. Thus, key experiments should be repeated using comparable sequence pairs.
Moreover, in the initial characterization of the binding abilities (Figure 1), the authors should include blunt-ended controls (duplex/hairpin) and, importantly, a pseudoknot (PK), as these structures are crucial for multiple steps in the viral life cycle (frameshifting, replication). Specifically, the PK in the 3'UTR (Sola et al., 2011, RNA Biology) may be an interesting target structure for unwinding assays, as it recruits the RTC, and, to my knowledge, no studies are available regarding nsp13 function at a PK. This would be particularly interesting in combination with nsp7/8 (Ohyama et al., 2024, JACS Au). -

Reviewer #2 (Public review):
Summary:
The authors are trying to broaden the understanding of SARS-CoV2 Nsp13 activity to show that a single viral protein can accomplish multiple functions. Additionally, they try to show that helicase function is not limited to ATP-driven, unidirectional unwinding.
Strengths:
The consistent application of statistics to triplicate experiments is a strength of the manuscript. The ToPif1 control in Figure S12 is a good control.
Weaknesses:
(1) All the experiments except the one in Figure S2 use N-terminally His-tagged Nsp13. Because the N-terminal tag is known to have large effects on Nsp13 activity, this calls into question virtually all of the results in this manuscript.
(2) The ATP-independent, bidirectional duplex unwinding shown for short duplex substrates is reminiscent of the trapping of thermal …
Reviewer #2 (Public review):
Summary:
The authors are trying to broaden the understanding of SARS-CoV2 Nsp13 activity to show that a single viral protein can accomplish multiple functions. Additionally, they try to show that helicase function is not limited to ATP-driven, unidirectional unwinding.
Strengths:
The consistent application of statistics to triplicate experiments is a strength of the manuscript. The ToPif1 control in Figure S12 is a good control.
Weaknesses:
(1) All the experiments except the one in Figure S2 use N-terminally His-tagged Nsp13. Because the N-terminal tag is known to have large effects on Nsp13 activity, this calls into question virtually all of the results in this manuscript.
(2) The ATP-independent, bidirectional duplex unwinding shown for short duplex substrates is reminiscent of the trapping of thermal fraying intermediates that have been reported for other helicases. Because they are only observed on short duplexes, do not require ATP, and are bidirectional, this does not suggest strand displacement as suggested in the manuscript. Instead, it suggests trapping of partially melted intermediates.
(3) Results that may be artifacts of unusual in vitro conditions are interpreted as if similar results will occur in the cell, where ATP is likely always present. Along those same lines, SARS-CoV-2 replicates in compartments of the endoplasmic reticulum, which would limit the ability of Nsp13 to access DNA substrates.
(4) There is no evidence to support the conclusion that "Duplex DNA supports bidirectional remodeling via both ATP-dependent and ATP-independent mechanisms." 3'-5' duplex melting is limited to short duplexes and is ATP-independent, suggesting it may be due to trapping of thermal fraying intermediates by the ssDNA binding Nsp13. The ATP-dependent and ATP-independent melting on the substrates with the 3'-overhang are the same, suggesting that ATP-dependent melting does not occur on this substrate, which would indicate that bidirectional ATP-dependent translocation does not occur.
(5) The description of ATP-independent unwinding as having "limited processivity," is likely not accurate. These experiments were multiturnover reactions with very high Nsp13 concentrations and no protein trap to ensure single turnover conditions. Because the reactions were multi-turnover, no information about the processivity of Nsp13 can be obtained. On the contrary, it seems likely that the product formed over the 30-minute reaction with a vast excess of Nsp13 is due to binding and dissociation of multiple Nsp13 molecules instead of processive translocation by a single enzyme.
(6) G4s are much more stable at cellular K+ concentrations than they are at 20 mM K+. As such, Nsp13's ability to unfold a G4 in the absence of ATP may be diminished or eliminated at a physiological K+ concentration.
Although the authors show that His-tagged Nsp13 can melt DNA and RNA duplexes and G-quadruplexes in an ATP-dependent and independent manner, in addition to annealing single-stranded nucleic acids into duplexes, the use of His-tagged Nsp13, which is known to cause artifacts, makes their results difficult to draw conclusions from. As such, in the opinion of this reviewer, this manuscript is likely to have little impact on the field.
-

Author response:
Public Reviews:
Reviewer #1 (Public review):
In the manuscript by Li et al., the authors perform a comprehensive study on the template and cofactor determinants of the SARS-CoV-2 nsp13 protein. They find that, alongside the classical processive unwinding ability of helicases driven by ATP consumption, other chaperone-like and ATP-independent functions exist for this enzyme. By testing DNA and RNA oligos in several conformations, the authors show that these functions are highly dependent on template identity, but also on the ratio of ATP to divalent cations. Ultimately, it is suggested that these distinct mechanisms of action are employed by nsp13 to orchestrate viral replication.
Overall, this study provides some novel insights into the functionality of a central and conserved enzyme of a relevant human pathogenic …
Author response:
Public Reviews:
Reviewer #1 (Public review):
In the manuscript by Li et al., the authors perform a comprehensive study on the template and cofactor determinants of the SARS-CoV-2 nsp13 protein. They find that, alongside the classical processive unwinding ability of helicases driven by ATP consumption, other chaperone-like and ATP-independent functions exist for this enzyme. By testing DNA and RNA oligos in several conformations, the authors show that these functions are highly dependent on template identity, but also on the ratio of ATP to divalent cations. Ultimately, it is suggested that these distinct mechanisms of action are employed by nsp13 to orchestrate viral replication.
Overall, this study provides some novel insights into the functionality of a central and conserved enzyme of a relevant human pathogenic virus. While the approach is important and adds to the field, particularly by characterizing the chaperoning activities and adding G-quadruplexes as templates, previous studies have already identified several determinants of nsp13 template binding and processing in vitro (Sommers et al., 2023, JBC; Park et al., 2025, JBC). In addition, some issues regarding experimental design need to be addressed to increase the cogency and biological relevance of the study.
We thank the reviewer for recognizing the novelty of our work, particularly the ATP-independent chaperone-like activities and G-quadruplex remodeling. We also appreciate the opportunity to clarify the conceptual distinction between our study and the prior work by Sommers et al. (2023) and Park et al. (2025). We fully agree that those studies systematically defined the canonical ATP-driven motor mechanism of Nsp13. Our results on 5′→3′ polarity, DNA preference, and tail/ATP/Mg2+ dependence align with these benchmarks, confirming the reliability of our platform.
However, the core novelty of our work lies in revealing that Nsp13 functions as a multifaceted nucleic acid remodeler, integrating motor and non-motor activities within a single protein-a functional regime absent from the JBC papers. Specifically, we uncover three novel layers: 1. Mg2+-activated, ATP-independent remodeling of short duplexes and G-quadruplexes. 2. Bidirectional remodeling on duplexes in the Mg2+-primed state. 3. Intrinsic chaperone functions including strand annealing and stem-loop restructuring.
Thus, our work fundamentally expands the biochemical model of Nsp13 from a simple ATP-driven motor to a multifunctional, mode-switchable remodeler. We will highlight these distinctions in the revised Discussion. Below, we respond point-by-point to the specific experimental design issues.
(1) Generally, low concentrations of monovalent cations (20 mM), as used throughout this study, may influence helicase activity and artificially enhance protein binding/oligomerization, which could favor the observed chaperoning activity (Venus et al., 2022, Methods). In contrast, some helicases, such as HCV NS3, are inhibited by higher K+ concentrations (Gwack et al., 2004, FEBS). Thus, the influence of higher concentrations of monovalent cations should be tested in relevant assays, as intracellular K+ levels are usually >100 mM. Additionally, this could significantly affect template stability. For instance, in some G4 assays, the addition of the trap already leads to observable duplex formation (Figure 5), which may be due to low K+ conditions.
We thank the reviewer for this critical comment regarding the ionic environment. We agree that monovalent cation concentrations are pivotal for both helicase activity and the structural stability of templates like G4s.
First, we wish to clarify that the final NaCl concentration in our reaction is not 20 mM, as this refers only to the unwinding buffer. Our protein dilution buffer contains 200 mM NaCl, and each 10 μL reaction includes 2 μL of protein, contributing ~40 mM NaCl. With 20 mM from the reaction buffer, the final concentration reaches~60 mM. We will clarify this in the Methods.
Second, our choice of ionic strength is guided by established literature. A survey of 27 published nsp13 studies (Author response table 1) shows that the majority use 20–50 mM monovalent cations, with 20 mM being most common. Mickolajczyk et al. (2021) showed that nsp13 activity is highest at low salt and declines at higher concentrations. Thus, low salt conditions are routinely used to capture nsp13’s intrinsic catalytic activity. The intracellular environment is far more complex, with crowding and interacting proteins that likely modulate helicase behavior. The low-salt conditions are therefore a deliberate simplification to isolate and define enzyme function.
Planned experiments: We fully agree that higher salt concentrations should be tested. In the revision, we will perform key assays such as ATP-independent duplex unwinding and G4 unfolding at ≥100 mM NaCl or KCl to verify that the observed activities persist under more physiological ionic conditions
(2) As in most publications that focus strictly on helicase (or other enzymatic) functions, the activity of the isolated protein is examined. However, particularly in the case of nsp13, core functions rely on other factors, such as nsp7/8 and other components of the replication-transcription complex (RTC). The overall structure and oligomerization state of nsp13 are altered within the complex (Chen et al., 2022, NSMB). The inclusion of such factors in key experiments would greatly improve the biological relevance of the findings.
We agree that examining Nsp13 within the context of the RTC is essential for establishing the biological relevance of our findings. The structural reorganization of Nsp13 upon binding to Nsp12 and Nsp7/8 (Chen et al., 2022) suggests that its enzymatic "mode" may be regulated by its protein partners.
Planned experiments: To address this, we will include the following biochemical characterizations:
(1) Nsp13/12 and Nsp13/7/8 sub-complexes will be examined to dissect the individual contributions of the polymerase and the primase-like factors to Nsp13’s multifaceted activities.
(2) The core RTC (Nsp13/12/7/8) will be used to evaluate how the full assembly modulates the functions of Nsp13 particularly on complex templates like G4 and pseudoknots.
(3) In Figure 4, the authors claim that Mg2+ concentration inhibits RNA unwinding. While this is likely considering previous findings, it must be validated that duplex stabilization is not the primary cause for the observed lower dissociation rates. As the template is only 12 bp long with extensive overhangs, higher ion concentrations may significantly stabilize base pairing by reducing fraying effects. Similarly, in Figure 6, template-dependent effects of Mg2+/ATP should be ruled out.
We thank the reviewer for this insightful suggestion. We agree that it is critical to distinguish whether the observed inhibition of RNA unwinding at higher Mg2+ concentrations is due to the physical stabilization of the RNA duplex.
Planned experiments: To address this, we will perform the following characterizations:
(1) We will measure the Tm of the RNA duplex used in Figure 4 across a range of Mg2+ concentrations (0, 0.5, and 1.0 mM). This will allow us to quantify the extent to which divalent cations stabilize the duplex RNA. These data will provide a more rigorous interpretation of the Mg2+-dependent unwinding in Figure 4.
(2) Similarly, we will perform thermal melting analyses for the various DNA and RNA templates used in Figure 6 under different Mg2+/ATP conditions to rule out the template-dependent effects of Mg2+/ATP.
(4) It is not entirely clear to me by which principle the templates were chosen. In my opinion, it would improve the overall comparability of the experimental results if, for instance, the blunt-ended duplex had the same sequence as the oligos with overhangs, since factors such as length, G/C content, Tm, etc., may play a significant role in binding and unwinding. Similarly, the oligos for binding and unwinding should be kept somewhat comparable, e.g., the G4 for the binding assay has 3 stacks, whereas RG1 has only 2. This discrepancy could make a significant difference. Thus, key experiments should be repeated using comparable sequence pairs.
We fully agree with the reviewer that maintaining sequence consistency across different assays is essential for a rigorous comparison of nsp13 activities. We apologize for the ambiguity in the initial presentation of our sequences in Table S1.
Planned revisions and experiments:
(1) We wish to clarify that several key substrates were sequence-matched. For unwinding assays, the 12-bp 3′-overhang DNA and blunt-ended DNA share the identical duplex sequence, and the 16-bp 5′-overhang and 3′-overhang DNA substrates are also sequence-matched. For annealing assays, the duplex regions for all DNA substrates (3′, 5′, blunt, and fork) are identical, and the same internal consistency was maintained for all RNA annealing substrates. To make this clear, we will reorganize Table S1 to explicitly group these sequence-paired substrates.
(2) The reviewer also notes discrepancies between binding and unwinding substrates (e.g., the difference in G4 stacks). To ensure direct comparability, we will perform additional experiments: complete binding assays for RG-1 (the 2-stack G4 used in unwinding) to match the functional data, and systematically measure binding affinities for all key unwinding substrates, including 3′-overhang, 5′-overhang, blunt-ended DNA, and the RNA fork.
(5) Moreover, in the initial characterization of the binding abilities (Figure 1), the authors should include blunt-ended controls (duplex/hairpin) and, importantly, a pseudoknot (PK), as these structures are crucial for multiple steps in the viral life cycle (frameshifting, replication). Specifically, the PK in the 3'UTR (Sola et al., 2011, RNA Biology) may be an interesting target structure for unwinding assays, as it recruits the RTC, and, to my knowledge, no studies are available regarding nsp13 function at a PK. This would be particularly interesting in combination with nsp7/8 (Ohyama et al., 2024, JACS Au).
We thank the reviewer for this insightful and inspiring suggestion. Incorporating pseudoknot (PK) structures into our analysis—particularly the well-characterized PK in the 3'UTR (Sola et al., 2011)—represents a significant opportunity to bridge our biochemical findings with the viral life cycle. To address this, we have designed a 3'UTR PK substrate based on recently reported scaffolds (Ohyama et al., 2024).
Planned experiments:
(1) We will expand our initial binding assays (Figure 1) to include blunt-ended duplexes, hairpins, and the 3'UTR PK. This will establish a baseline for how Nsp13 recognizes these structurally distinct and physiologically critical templates.
(2) We will perform unwinding assays to determine whether Nsp13, in its isolated state, possesses the mechanical capability to resolve the complex tertiary interactions within a pseudoknot.
(3) Following the reviewer's insight, we will examine whether the addition of nsp7/8 is required to facilitate the unfolding of the 3'UTR PK.
Together, these experiments will allow us to assess whether Nsp13 is capable of managing one of the most challenging structural obstacles in the SARS-CoV-2 genome.
Reviewer #2 (Public review):
Summary:
The authors are trying to broaden the understanding of SARS-CoV2 Nsp13 activity to show that a single viral protein can accomplish multiple functions. Additionally, they try to show that helicase function is not limited to ATP-driven, unidirectional unwinding.
Strengths: The consistent application of statistics to triplicate experiments is a strength of the manuscript. The ToPif1 control in Figure S12 is a good control.
We thank the reviewer for the insightful assessment and for highlighting the rigor of our experimental design, particularly our reliance on triplicate data with robust statistical validation and the inclusion of the ToPif1 control.
We are especially grateful for the detailed comments provided by the reviewer. We fully recognize that addressing these specific points is essential for strengthening the cogency of our conclusions and improving the overall rigor of the manuscript. These suggestions have provided us with a clear roadmap for further refining our experimental evidence and clarifying our mechanistic interpretations. Below, we respond point-by-point to the specific issues.
Weaknesses:
(1) All the experiments except the one in Figure S2 use N-terminally His-tagged Nsp13. Because the N-terminal tag is known to have large effects on Nsp13 activity, this calls into question virtually all of the results in this manuscript.
We thank the reviewer for raising this important concern regarding the potential influence of the N-terminal His tag on nsp13 activity. We have carefully considered this issue and provide the following lines of evidence to address it.
(1) We have generated a tag-free nsp13 variant and our preliminary characterization (Author response image 1) shows that it retains all key activities: ATP hydrolysis (comparable to His-tagged nsp13), both ATP-independent (Mg2+-activated) and ATP-dependent unwinding, as well as chaperone activity to remodel stem-loops. These results demonstrate that while the His tag may modulate enzymatic efficiency, it does not create or abolish any specific biochemical function.
(2) We conducted a systematic survey of 27 published studies on SARS-CoV/SARS-CoV-2 nsp13 (Author response table 1). The results show that 17 out of 27 studies (63%) used affinity-tagged nsp13 without tag removal, including His, MBP, GST, and Strep tags.
(3) The only study that systematically compared different affinity tags (Adedeji et al., 2012) reported that GST-tagged nsp13 exhibited ~520-fold higher ATPase activity than His-tagged nsp13, demonstrating that the choice of affinity tag can affect enzymatic efficiency. However, both tagged versions retained all core enzymatic activities, including ATP hydrolysis and duplex unwinding. Importantly, no study has compared the full functional spectrum between His-tagged and tag-free nsp13. Our preliminary data suggest that the His tag may affect efficiency but does not alter the presence or absence of any specific activity.
Planned experiments:
We fully agree with the reviewer that a more systematic comparison would strengthen the conclusions. In the revision, we will include additional characterization of tag-free nsp13: (i) quantitative nucleic acid binding affinity, (ii) G4 unfolding efficiency, (iii) strand annealing activity. These experiments are currently underway.
In summary, while we acknowledge that the His tag may influence enzymatic efficiency, our key conclusions are supported by experiments with tag-free nsp13. We will add a discussion of these points and include additional tag-free nsp13 data in the revised manuscript.
(2) The ATP-independent, bidirectional duplex unwinding shown for short duplex substrates is reminiscent of the trapping of thermal fraying intermediates that have been reported for other helicases. Because they are only observed on short duplexes, do not require ATP, and are bidirectional, this does not suggest strand displacement as suggested in the manuscript. Instead, it suggests trapping of partially melted intermediates.
We thank the reviewer for this insightful perspective. While the passive trapping of thermal fraying intermediates is a well-established model for non-catalytic protein-nucleic acid interactions, several lines of evidence suggest that nsp13 employs a more active, allosteric mechanism for ATP-independent remodeling.
(1) If nsp13 were merely a passive trap, increasing duplex stability should decrease unwinding. However, as shown in Figure S3, raising Mg2+ from 0 to 5 mM increases the DNA duplex Tm by ~10°C, yet nsp13’s remodeling activity is markedly enhanced under the same conditions (Figure 2). This positive correlation between cation-induced substrate stabilization and protein activation supports an active, protein-centered mechanism that overcomes the increased energetic barrier.
(2) The observed bidirectionality in ATP-independent remodeling does not simply imply a lack of polarity; rather, it can reflect nsp13’s intrinsic chaperone function. In the absence of ATP, nsp13 binds the ss/ds junction (Figure 2F) and, in a Mg2+-dependent manner, may use its binding energy to actively intercalate into the duplex. This mechanism is inherently symmetric for 3′ and 5′ overhangs, explaining bidirectional remodeling, while the absence of activity on blunt-ended substrates confirms the requirement for a pre-existing junction.
(3) The lack of activity on 24-bp substrates does not negate this remodeling mode but defines its energetic boundary. The binding energy released upon nsp13-nucleic acid interaction is sufficient to overcome the lower unwinding barrier of 12-16 bp duplexes, but insufficient to counteract the high stability and rapid re-annealing of a 24-bp duplex without the continuous mechanical power of ATP hydrolysis.
Planned Revision:
We thank the reviewer for prompting us to refine our mechanistic model. In the revision, we will add a dedicated discussion explicitly comparing the model of allosterically activated, binding-driven strand intrusion with the passive trapping model, incorporating the Tm data to strengthen our conclusions.
(3) Results that may be artifacts of unusual in vitro conditions are interpreted as if similar results will occur in the cell, where ATP is likely always present. Along those same lines, SARS-CoV-2 replicates in compartments of the endoplasmic reticulum, which would limit the ability of Nsp13 to access DNA substrates.
We thank the reviewer for raising this important concern regarding the physiological relevance. We fully agree that in vitro conditions do not entirely recapitulate the complex intracellular environment, and we have been careful not to over-interpret our findings. Below we address the two specific issues raised:
(1) Regarding the ATP-independent activity, we acknowledge that ATP is abundant in healthy, actively replicating cells. However, during rapid viral replication, local ATP concentrations can fluctuate due to the high energy demand of the RTC as the template contains extensive secondary structures, which may lead to transient ATP depletion. Under such energy-limited conditions, Yu et al. (2025) demonstrated that ADP-bound nsp13 exhibits chaperone activity that destabilizes nucleic acid structures without ATP hydrolysis, and Dumm et al. (2025) reported that SARS-CoV-2 nsp13 resolves RNA stem-loops in an ATP-independent manner.
Even when ATP is abundant, the ATP-independent mode may enable rapid, local structural adjustments that bypass the kinetic delay of ATP binding and hydrolysis. As shown in Figure 1D, nsp13 exhibits high binding affinity for structured nucleic acids. In this scenario, nsp13 functions not as a processive motor but through a binding-driven mechanism, using the free energy of protein-nucleic acid interaction to transiently destabilize short duplexes or resolve local secondary structures such as G4s and stem-loops in an energy-efficient manner.
(2) Regarding DNA substrates, we fully agree that RNA is the physiological substrate for nsp13. However, DNA is a validated and widely accepted surrogate for mechanistic studies because DNA is more stable and easier to manipulate than RNA to yield the mechanistic insights. A systematic survey of 27 published nsp13 studies (Author response table 1) shows that 20 out of 27 (74%) used DNA substrates for at least some of their experiments. In our study, we used DNA primarily as a mechanistic probe and a stable control, and we validated all key conclusions on physiological RNA substrates, as shown in Figures 4, 5, 6, S7, S8, S10, S11 and S12.
Planned revisions: To address the reviewer’s concerns more directly, we will revise the manuscript to include a discussion paragraph explicitly stating that the ATP-independent activity was observed under optimized in vitro conditions and may represent a latent remodeling capability that could be relevant under energy-limited conditions such as local ATP depletion during rapid replication. We will also clarify that DNA substrates were used as mechanistic probes and controls, and that all key findings were validated on physiological RNA substrates. We thank the reviewer for prompting us to strengthen the discussion of these important points.
(4) There is no evidence to support the conclusion that "Duplex DNA supports bidirectional remodeling via both ATP-dependent and ATP-independent mechanisms." 3'-5' duplex melting is limited to short duplexes and is ATP-independent, suggesting it may be due to trapping of thermal fraying intermediates by the ssDNA binding Nsp13. The ATP-dependent and ATP-independent melting on the substrates with the 3'-overhang are the same, suggesting that ATP-dependent melting does not occur on this substrate, which would indicate that bidirectional ATP-dependent translocation does not occur.
We are grateful to the reviewer for this critical evaluation of our mechanistic claims. We agree that our initial statement regarding bidirectional ATP-dependent remodeling was imprecise and not fully supported by the data. As the reviewer correctly notes, the similar unwinding efficiency on 3′-overhang substrates regardless of ATP presence indicates that ATP hydrolysis does not drive 3′→5′ translocation, which is consistent with nsp13’s known 5′→3′ motor polarity. The observed 3′→5′ activity is therefore more accurately described as an ATP-independent remodeling event, not ATP-dependent unwinding.
We will revise the Discussion and relevant Results sections to clarify the nature of this bidirectional activity. Specifically, the sentence:
"Duplex DNA supports bidirectional remodeling via both ATP-dependent and ATP-independent mechanisms..."will be corrected to: "Duplex DNA supports bidirectional remodeling via ATP-independent mechanisms."
We will also explicitly state that while nsp13 requires ATP for long-range, processive 5'→3' helicase activity, its remodeling/chaperone function is inherently bidirectional and powered by the free energy of binding to the ss/ds junction, rather than by ATP-driven mechanical work.
(5)-The description of ATP-independent unwinding as having "limited processivity," is likely not accurate. These experiments were multiturnover reactions with very high Nsp13 concentrations and no protein trap to ensure single turnover conditions. Because the reactions were multi-turnover, no information about the processivity of Nsp13 can be obtained. On the contrary, it seems likely that the product formed over the 30-minute reaction with a vast excess of Nsp13 is due to binding and dissociation of multiple Nsp13 molecules instead of processive translocation by a single enzyme.
We thank the reviewer for this important correction. We fully agree that our use of the term "processivity" was technically imprecise. Processivity strictly defines the distance a single enzyme translocates during one binding event, which our multi-turnover assays (with high nsp13 concentrations and no protein trap) were not designed to measure. Our results specifically demonstrate that the ATP-independent remodeling mode is highly sensitive to duplex length, with efficiency declining sharply as the duplex lengthens. To reflect the experimental data more faithfully, we have replaced "processivity" with more accurate descriptors throughout the manuscript.
Planned revisions:
(1) Original: "The ATP-independent unwinding mode, however, has limited processivity." Revised: "The ATP-independent unwinding mode, however, exhibits a steep decline in efficiency as the duplex length increases."
(2) Original: "...an ATP-independent, cation-activated mode with limited processivity." Revised: "...an ATP-independent, cation-activated mode specialized for localized structural remodeling"
(3) Original: "...primes Nsp13 for basal strand remodeling but supports only limited processivity." Revised: "...primes Nsp13 for basal strand remodeling but is insufficient for the sustained unwinding of extended duplexes."
(4) Original: "...primes Nsp13 for low-processivity strand displacement." Revised: "...primes Nsp13 for short-range strand displacement rather than long-range processive unwinding."
We believe these changes clarify that the ATP-independent mode acts as a molecular chaperone for local obstacles (like G4 or short stems) rather than a motor for long-range translocation. We thank the reviewer for helping us improve the precision of our description.
(6) G4s are much more stable at cellular K+ concentrations than they are at 20 mM K+. As such, Nsp13's ability to unfold a G4 in the absence of ATP may be diminished or eliminated at a physiological K+ concentration.
We thank the reviewer for this critical point regarding physiological ion concentrations. We agree that K+ significantly stabilizes G4 structures, which may raise the energy barrier for ATP-independent remodeling.
Planned experiments:
To address this, we will perform salt titration assays (up to 150 mM KCl) to evaluate the robustness of nsp13’s G4 unfolding activity under more physiological ionic conditions. We will also measure the melting temperature of our G4 substrates across this K+ range to correlate structural stability with enzymatic efficiency.
Author response image 1.
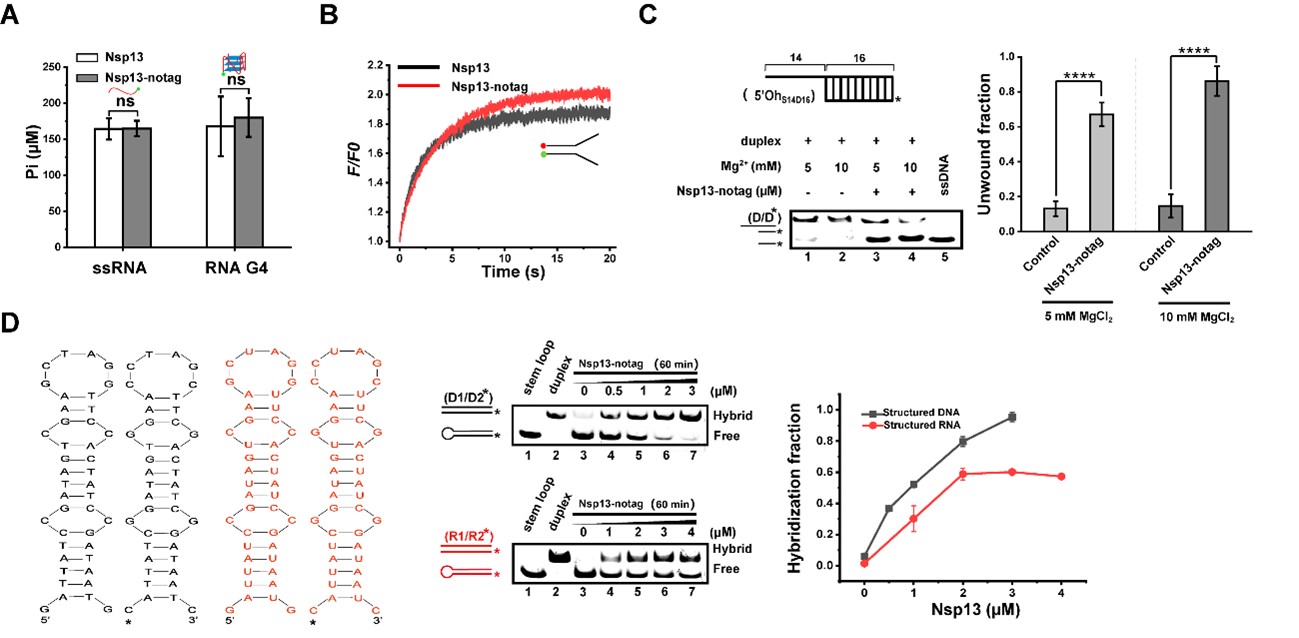
Preliminary characterization of tag-free Nsp13 enzymatic activities. (A) Comparison of ATPase activity between His-tagged and tag-free Nsp13 in the presence of ssRNA or RNA G4. (B) Raw fluorescence data from stopped-flow FRET analysis of ATP-dependent unwinding (16-bp fork DNA, 2 mM Mg2+, 2 mM ATP). F/F0 represents FAM fluorescence normalized to initial DNA intensity. (C) ATP-independent DNA duplex remodeling (data reproduced from Figure S2). (D) Chaperone activity of tag-free Nsp13 on DNA and RNA stem-loops.
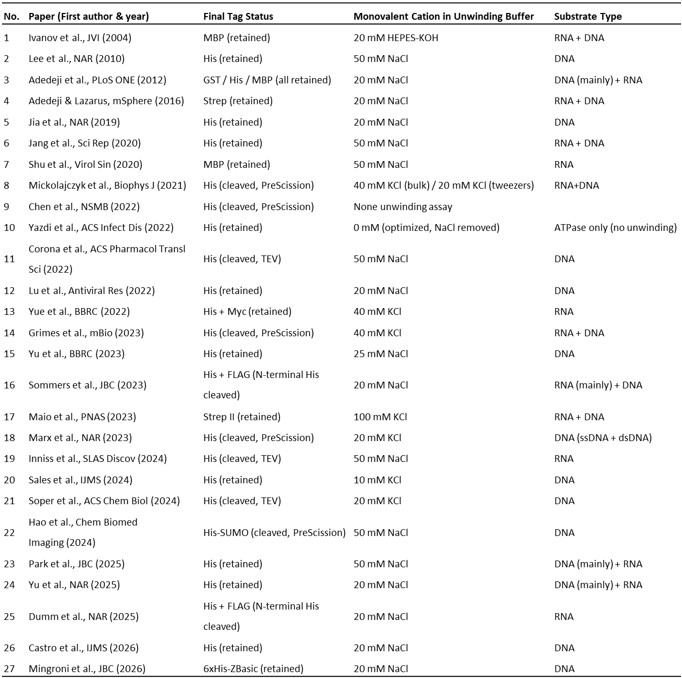
Author response table 1.
Summary of affinity tags, monovalent salt concentrations, and substrate types used in 27 published SARS-CoV/SARS-CoV-2 nsp13 studies
References:
(1) Ivanov KA, Thiel V, Dobbe JC, van der Meer Y, Snijder EJ, Ziebuhr J. Multiple enzymatic activities associated with severe acute respiratory syndrome coronavirus helicase. J Virol. 2004 Jun;78(11):5619-32.
(2) Lee NR, Kwon HM, Park K, Oh S, Jeong YJ, Kim DE. Cooperative translocation enhances the unwinding of duplex DNA by SARS coronavirus helicase nsP13. Nucleic Acids Res. 2010 Nov;38(21):7626-36.
(3) Adedeji AO, Marchand B, Te Velthuis AJ, Snijder EJ, Weiss S, Eoff RL, Singh K, Sarafianos SG. Mechanism of nucleic acid unwinding by SARS-CoV helicase. PLoS One. 2012;7(5):e36521. doi: 10.1371/journal.pone.0036521.
(4) Adedeji AO, Lazarus H. Biochemical Characterization of Middle East Respiratory Syndrome Coronavirus Helicase. mSphere. 2016 Sep 7;1(5):e00235-16.
(5) Jia Z, Yan L, Ren Z, Wu L, Wang J, Guo J, Zheng L, Ming Z, Zhang L, Lou Z, Rao Z. Delicate structural coordination of the Severe Acute Respiratory Syndrome coronavirus Nsp13 upon ATP hydrolysis. Nucleic Acids Res. 2019 Jul 9;47(12):6538-6550.
(4) Jang KJ, Jeong S, Kang DY, Sp N, Yang YM, Kim DE. A high ATP concentration enhances the cooperative translocation of the SARS coronavirus helicase nsP13 in the unwinding of duplex RNA. Sci Rep. 2020 Mar 11;10(1):4481.
(5) Shu T, Huang M, Wu D, Ren Y, Zhang X, Han Y, Mu J, Wang R, Qiu Y, Zhang DY, Zhou X. SARS-Coronavirus-2 Nsp13 Possesses NTPase and RNA Helicase Activities That Can Be Inhibited by Bismuth Salts. Virol Sin. 2020 Jun;35(3):321-329.
(6) Mickolajczyk KJ, Shelton PMM, Grasso M, Cao X, Warrington SE, Aher A, Liu S, Kapoor TM. Force-dependent stimulation of RNA unwinding by SARS-CoV-2 nsp13 helicase. Biophys J. 2021 Mar 16;120(6):1020-1030.
(7) Chen J, Wang Q, Malone B, Llewellyn E, Pechersky Y, Maruthi K, Eng ET, Perry JK, Campbell EA, Shaw DE, Darst SA. Ensemble cryo-EM reveals conformational states of the nsp13 helicase in the SARS-CoV-2 helicase replication-transcription complex. Nat Struct Mol Biol. 2022 Mar;29(3):250-260.
(8) Yazdi AK, Pakarian P, Perveen S, Hajian T, Santhakumar V, Bolotokova A, Li F, Vedadi M. Kinetic Characterization of SARS-CoV-2 nsp13 ATPase Activity and Discovery of Small-Molecule Inhibitors. ACS Infect Dis. 2022 Aug 12;8(8):1533-1542.
(9) Corona A, Wycisk K, Talarico C, Manelfi C, Milia J, Cannalire R, Esposito F, Gribbon P, Zaliani A, Iaconis D, Beccari AR, Summa V, Nowotny M, Tramontano E. Natural Compounds Inhibit SARS-CoV-2 nsp13 Unwinding and ATPase Enzyme Activities. ACS Pharmacol Transl Sci. 2022 Apr 1;5(4):226-239.
(10) Lu L, Peng Y, Yao H, Wang Y, Li J, Yang Y, Lin Z. Punicalagin as an allosteric NSP13 helicase inhibitor potently suppresses SARS-CoV-2 replication in vitro. Antiviral Res. 2022 Oct;206:105389.
(11) Yue K, Yao B, Shi Y, Yang Y, Qian Z, Ci Y, Shi L. The stalk domain of SARS-CoV-2 NSP13 is essential for its helicase activity. Biochem Biophys Res Commun. 2022 Apr 23;601:129-136.
(12) Grimes SL, Choi YJ, Banerjee A, Small G, Anderson-Daniels J, Gribble J, Pruijssers AJ, Agostini ML, Abu-Shmais A, Lu X, Darst SA, Campbell E, Denison MR. A mutation in the coronavirus nsp13-helicase impairs enzymatic activity and confers partial remdesivir resistance. mBio. 2023 Aug 31;14(4):e0106023.
(13) Yu J, Im H, Lee G. Unwinding mechanism of SARS-CoV helicase (nsp13) in the presence of Ca2+, elucidated by biochemical and single-molecular studies. Biochem Biophys Res Commun. 2023 Aug 6;668:35-41.
(14) Sommers JA, Loftus LN, Jones MP 3rd, Lee RA, Haren CE, Dumm AJ, Brosh RM Jr. Biochemical analysis of SARS-CoV-2 Nsp13 helicase implicated in COVID-19 and factors that regulate its catalytic functions. J Biol Chem. 2023 Mar;299(3):102980.
(15) Maio N, Raza MK, Li Y, Zhang DL, Bollinger JM Jr, Krebs C, Rouault TA. An iron-sulfur cluster in the zinc-binding domain of the SARS-CoV-2 helicase modulates its RNA-binding and -unwinding activities. Proc Natl Acad Sci U S A. 2023 Aug 15;120(33):e2303860120.
(16) Marx SK, Mickolajczyk KJ, Craig JM, Thomas CA, Pfeffer AM, Abell SJ, Carrasco JD, Franzi MC, Huang JR, Kim HC, Brinkerhoff H, Kapoor TM, Gundlach JH, Laszlo AH. Observing inhibition of the SARS-CoV-2 helicase at single-nucleotide resolution. Nucleic Acids Res. 2023 Sep 22;51(17):9266-9278.
(17) Inniss NL, Rzhetskaya M, Ling-Hu T, Lorenzo-Redondo R, Bachta KE, Satchell KJF, Hultquist JF. Activity and inhibition of the SARS-CoV-2 Omicron nsp13 R392C variant using RNA duplex unwinding assays. SLAS Discov. 2024 Apr;29(3):100145.
(18) Sales AH, Fu I, Durandin A, Ciervo S, Lupoli TJ, Shafirovich V, Broyde S, Geacintov NE. Variable Inhibition of DNA Unwinding Rates Catalyzed by the SARS-CoV-2 Helicase Nsp13 by Structurally Distinct Single DNA Lesions. Int J Mol Sci. 2024 Jul 19;25(14):7930.
(19) Soper N, Yardumian I, Chen E, Yang C, Ciervo S, Oom AL, Desvignes L, Mulligan MJ, Zhang Y, Lupoli TJ. A Repurposed Drug Interferes with Nucleic Acid to Inhibit the Dual Activities of Coronavirus Nsp13. ACS Chem Biol. 2024 Jul 19;19(7):1593-1603.
(20) Hao W, Hu X, Chen Q, Qin B, Tian Z, Li Z, Hou P, Zhao R, Balci H, Cui S, Diao J. Duplex Unwinding Mechanism of Coronavirus MERS-CoV nsp13 Helicase. Chem Biomed Imaging. 2024 Dec 19;3(2):111-122.
(21) Park J, Jeong YJ, Chauhan K, Koh HR, Kim DE. ATPase-dependent duplex nucleic acid unwinding by SARS-CoV-2 nsP13 relies on facile binding and translocation along single-stranded nucleic acid. J Biol Chem. 2025 Jul;301(7):110373.
(24) Yu J, Im H, Cho H, Jeon Y, Lee JB, Lee G. A novel ADP-directed chaperone function facilitates the ATP-driven motor activity of SARS-CoV helicase. Nucleic Acids Res. 2025 Jan 24;53(3):gkaf034.
(25) Dumm AJ, Zheng AY, Butler TJ, Kulikowicz T, George JC, Bombard PT, Sommers JA, Ding J, Brosh RM Jr. SARS-CoV-2 point mutations are over-represented in terminal loops of RNA stem-loop structures that can be resolved by Nsp13 helicase in a unique manner with respect to nucleotide dependence. Nucleic Acids Res. 2025 May 22;53(10):gkaf447.
(26) Castro JM, Slack RL, Ong YT, Zhang H, Gifford LB, Courouble VV, Aiken RM, Shankar V, O'Leary TR, Griffin PR, Lan S, Du Y, Fu H, Sarafianos SG. Stalling the Enemy: Targeting Nsp13 for Next-Generation SARS-CoV-2 Antivirals. Int J Mol Sci. 2026 Mar 11;27(6):2587.
(27) Mingroni MA, Enney BM, Malsick LE, Geiss BJ. Motif V is an allosteric couple between the SARS-CoV-2 nsp13 nucleotide triphosphatase and helicase active sites. J Biol Chem. 2026 Mar;302(3):111198.
-