Splicing factor SRSF1 is essential for homing of precursor spermatogonial stem cells in mice
Curation statements for this article:-
Curated by eLife
eLife assessment
In this valuable study, the authors characterize the role of splicing factor SRSF1 during spermatogenesis with a conditional knockout of Srsf1 in male germ cells. The phenotype and molecular role of SRSF1 in regulating alternative splicing in precursor spermatogonial stem cells in juvenile testes are convincingly supported. The paper also provides convincing evidence that the mRNA encoding Tial, a factor relevant to spermatogonial maintenance and male fertility, is alternatively spliced in testis and that this splicing is regulated by SRSF1. The work will be of interest to the fields of reproductive biology, stem cell biology, and alternative splicing.
This article has been Reviewed by the following groups
Discuss this preprint
Start a discussion What are Sciety discussions?Listed in
- Evaluated articles (eLife)
Abstract
Spermatogonial stem cells (SSCs) are essential for continuous spermatogenesis and male fertility. The underlying mechanisms of alternative splicing (AS) in mouse SSCs are still largely unclear. We demonstrated that SRSF1 is essential for gene expression and splicing in mouse SSCs. Crosslinking immunoprecipitation and sequencing data revealed that spermatogonia-related genes (e.g. Plzf , Id4 , Setdb1, Stra8 , Tial1 / Tiar , Bcas2 , Ddx5 , Srsf10 , Uhrf1 , and Bud31 ) were bound by SRSF1 in the mouse testes. Specific deletion of Srsf1 in mouse germ cells impairs homing of precursor SSCs leading to male infertility. Whole-mount staining data showed the absence of germ cells in the testes of adult conditional knockout (cKO) mice, which indicates Sertoli cell-only syndrome in cKO mice. The expression of spermatogonia-related genes (e.g. Gfra1 , Pou5f1 , Plzf , Dnd1 , Stra8 , and Taf4b ) was significantly reduced in the testes of cKO mice. Moreover, multiomics analysis suggests that SRSF1 may affect survival of spermatogonia by directly binding and regulating Tial1 / Tiar expression through AS. In addition, immunoprecipitation mass spectrometry and co-immunoprecipitation data showed that SRSF1 interacts with RNA splicing-related proteins (e.g. SART1, RBM15, and SRSF10). Collectively, our data reveal the critical role of SRSF1 in spermatogonia survival, which may provide a framework to elucidate the molecular mechanisms of the posttranscriptional network underlying homing of precursor SSCs.
Article activity feed
-

-

-

-

Author Response
The following is the authors’ response to the previous reviews.
Reviewer #3 (Recommendations For The Authors):
- Fig. 2B: In their previous comment #6, I assume that Reviewer #2 was asking about peaks that were called as statistically significant above background, not just "higher" as assessed by eye. The authors have now marked peaks that are "higher" but still do not indicate that they were called as statistically significant by any software. I agree that they need to indicate in the figure which peaks were discovered by formal analysis.
Response: Thank you for the professional suggestions. We used the Piranha (version 1.2.1) software to call peaks from CLIP-seq data, in which the P-value threshold for peaks (i.e., the -p parameter) was set as 0.05. And then any region above the IgG peak could be a binding region, …
Author Response
The following is the authors’ response to the previous reviews.
Reviewer #3 (Recommendations For The Authors):
- Fig. 2B: In their previous comment #6, I assume that Reviewer #2 was asking about peaks that were called as statistically significant above background, not just "higher" as assessed by eye. The authors have now marked peaks that are "higher" but still do not indicate that they were called as statistically significant by any software. I agree that they need to indicate in the figure which peaks were discovered by formal analysis.
Response: Thank you for the professional suggestions. We used the Piranha (version 1.2.1) software to call peaks from CLIP-seq data, in which the P-value threshold for peaks (i.e., the -p parameter) was set as 0.05. And then any region above the IgG peak could be a binding region, and of course, the higher the peak, the more pre-mRNA SRSF1 binds in that region.
- Similar to the above comment, in Fig. 7G "visual analysis" of IGV tracks is not an assay. It is fine to show the tracks as an example of the differential expression called using DESeq2, but this should be described for what it is.
Response: We thank the reviewer for the professional comments. Following this advice, we have corrected the text in this revised version (Page 11, Line 233).
- Fig 5C: TUNEL results are supported by a single image of only a few cells. It is important to include quantitation as has been done for other microscopy data.
Response: Thank you for the professional suggestions. Following this advice, we have added the quantitative data in Figure 5C. Also, we have added specific quantification methods to the text (Page 23, Line 484-485).
- Legend to Fig 6C-E: I assume n=4 refers to the number of animals. It would be best to also know many cells/tubules were counted for each animal.
Response: Thank you for the helpful comments. Following this advice, we have revised the legend for Figure 6D, E (Page 12, Line 246-249).
- There appears to be a mistake in line 285-287, which reads: "the overall analysis of aberrant AS events showed that SRSF1 effectively promotes the occurrence of SE and MXE events and inhibits the occurrence of RI events." The data in Fig 8C appears to show the opposite, with more SE and MXE, and fewer RI events, in the SRSF1 KO. This would imply that SRSF1 normally inhibits SE/MXE and promotes RI.
Response: Thank you very much for the professional comments. Following this advice, we have corrected the text in this revised version (Page 14, Line 286-288).
- In Fig. 8E, an upper band is depleted in SRSF1 KO, but in Figure 8J, a much lower band is depleted. How is this explained?
Response: Thank you for the professional suggestions. Since exon 7 of Tial1 is in the non-coding region, the lower band in Figure 8E does not correspond to the lower band in Figure 8J. For better understanding, we show the detailed information of Tial1 in the attached Figure S3.
- Line 81: As a very minor point, "AS" is defined as alternative splicing in the abstract, but should be re-defined again in the main text when first mentioned.
Response: Thank you for the helpful comments. Following this advice, we have corrected the text in this revised version (Page 3, Line 81).
-

eLife assessment
In this valuable study, the authors characterize the role of splicing factor SRSF1 during spermatogenesis with a conditional knockout of Srsf1 in male germ cells. The phenotype and molecular role of SRSF1 in regulating alternative splicing in precursor spermatogonial stem cells in juvenile testes are convincingly supported. The paper also provides convincing evidence that the mRNA encoding Tial, a factor relevant to spermatogonial maintenance and male fertility, is alternatively spliced in testis and that this splicing is regulated by SRSF1. The work will be of interest to the fields of reproductive biology, stem cell biology, and alternative splicing.
-

Reviewer #1 (Public Review):
In this study, the authors seek to characterize the role of splicing factor SRSF1 during spermatogenesis using Vasa-Cre;Srsf1Fl/del mice model. The authors first revealed that spermatogonia-related genes (e.g., Plzf, Id4, Setdb1, Stra8, Tial1/Tiar, Bcas2, Ddx5, Srsf10, Uhrf1, and Bud31) were bound by SRSF1 in the mouse testes by CLIP-seq. The authors convincingly demonstrated that specific deletion of SRSF1 in mouse gem cells with vasa-cre lead to NOA by impairing homing and failure survival of spermatogonia. To investigate the molecular mechanisms of SRSF1 in spermatogonia, further multiomics analysis including CLIP-seq, IP-MS, and RNA-seq were conducted. The results showed that SRSF1 coordinated with other RNA splicing-related proteins to directly bind and regulate the expression of nine …
Reviewer #1 (Public Review):
In this study, the authors seek to characterize the role of splicing factor SRSF1 during spermatogenesis using Vasa-Cre;Srsf1Fl/del mice model. The authors first revealed that spermatogonia-related genes (e.g., Plzf, Id4, Setdb1, Stra8, Tial1/Tiar, Bcas2, Ddx5, Srsf10, Uhrf1, and Bud31) were bound by SRSF1 in the mouse testes by CLIP-seq. The authors convincingly demonstrated that specific deletion of SRSF1 in mouse gem cells with vasa-cre lead to NOA by impairing homing and failure survival of spermatogonia. To investigate the molecular mechanisms of SRSF1 in spermatogonia, further multiomics analysis including CLIP-seq, IP-MS, and RNA-seq were conducted. The results showed that SRSF1 coordinated with other RNA splicing-related proteins to directly bind and regulate the expression of nine spermatogonia-related genes especially Tial1/Tiar via alternative splicing. The authors revealed the critical role of SRSF1-mediated AS in precursor SSCs homing and survival, which may provide a framework to elucidate the molecular mechanisms of the posttranscriptional network underlying the formation of SSC pools and the establishment of niches. This work will be of interest to stem cell and reproductive biologists. The experiments are well-designed and conducted, and the overall methods and results are convincing except for the claim that altered splicing of the Tial1 transcript mediates the effect of SRSF1 loss.
-

Reviewer #2 (Public Review):
Summary
The authors seek to characterize the role of splicing factor SRSF1 during spermatogenesis. Using a conditional deletion of Srsf1 in germ cells, they find that SRSF1 is required for male fertility. Via immunostaining and RNA-seq analysis of the Srsf1 conditional knockout (cKO) testes, combined with SRSF1 CLIP-seq and IP-MS data from the testis, they ultimately conclude that Srsf1 is required for homing of precursor spermatogonial stem cells (SCCs) due to alternative splicing.Strengths
The overall methods and results are robust. The histological analysis of the Srsf1 cKO traces the origins of the fertility defect to the postnatal testis, and the authors have generated interesting datasets characterizing SRSF1's RNA targets and interacting proteins specifically in the testis.Ultimately, the authors …
Reviewer #2 (Public Review):
Summary
The authors seek to characterize the role of splicing factor SRSF1 during spermatogenesis. Using a conditional deletion of Srsf1 in germ cells, they find that SRSF1 is required for male fertility. Via immunostaining and RNA-seq analysis of the Srsf1 conditional knockout (cKO) testes, combined with SRSF1 CLIP-seq and IP-MS data from the testis, they ultimately conclude that Srsf1 is required for homing of precursor spermatogonial stem cells (SCCs) due to alternative splicing.Strengths
The overall methods and results are robust. The histological analysis of the Srsf1 cKO traces the origins of the fertility defect to the postnatal testis, and the authors have generated interesting datasets characterizing SRSF1's RNA targets and interacting proteins specifically in the testis.Ultimately, the authors have shown that SRSF1's effects on alternative splicing are required to establish spermatogenesis. In the absence of Srsf1, the postnatal gonocytes do not properly mature into spermatogonia and consequently never initiate spermatogenesis.
-

Reviewer #3 (Public Review):
In this study, Sun et al examine the role of the splicing factor SRSF1 in spermatogenesis in mice. Alternative splicing is important for spermatogenic development, but its regulation and major developmental roles during spermatogenesis are not well understood. The authors set out to better define both SRSF1 function in testes and the contribution of alternative splicing. They generate several large 'omics datasets to define SRSF1 targets in testis, including RNA interactions by CLIP-seq in whole testis, protein interactions by IP-mass spec in whole testis, and RNA sequencing to detect expression levels and splice variants. They also examine the phenotype of germline conditional knockouts (cKO) for Srsf1, using the early-acting Vasa-Cre, and find a severe depletion of germ cells starting at 7 days post partum …
Reviewer #3 (Public Review):
In this study, Sun et al examine the role of the splicing factor SRSF1 in spermatogenesis in mice. Alternative splicing is important for spermatogenic development, but its regulation and major developmental roles during spermatogenesis are not well understood. The authors set out to better define both SRSF1 function in testes and the contribution of alternative splicing. They generate several large 'omics datasets to define SRSF1 targets in testis, including RNA interactions by CLIP-seq in whole testis, protein interactions by IP-mass spec in whole testis, and RNA sequencing to detect expression levels and splice variants. They also examine the phenotype of germline conditional knockouts (cKO) for Srsf1, using the early-acting Vasa-Cre, and find a severe depletion of germ cells starting at 7 days post partum (dpp) and culminating with a lack of germ cells (Sertoli Cell Only Syndrome) by adulthood. They detect differences in gene expression as well as differences in splicing between control and knockout, including 9 genes that are downregulated, experience alternative splicing, and whose transcripts are also bound by SRSF1, and identify the Tial1/Tiar transcript as one of these targets. They conclude that SRSF1 is required for homing and self-renewal of precursor spermatogonial stem cells, and suggest that this role may be mediated in part though its regulation of Tial1/Tiar splicing.
Strengths of the paper include detailed phenotyping of the Srsf1 cKO, which convincingly supports the Sertoli Cell Only phenotype, establishes the timing of the first appearance of the spermatogonial defect, and provides new insight into the role of splicing factors and SRSF1 specifically in spermatogenesis. Another strength is the generation of CLIP-seq, IP-MS, and RNA-seq datasets which will be a useful resource for the field of germ cell development. Overall, the results support the claims made. While the study does not provide a full mechanistic understanding of how alternative splicing mediated by SRSF1 affects SSC precursors, the contributions are novel and useful, and will be of interest to the fields of alternative splicing and male reproductive biology.
-

-

Author Response
The following is the authors’ response to the original reviews.
Reviewer #1 (Recommendations For The Authors):
The conclusions of this paper are mostly well supported by data, but some aspects need to be corrected.
- Line 99. The title is not suitable for summarizing this part of the results. In this paragraph, the results mainly describe SRSF1 expression pattern and binding of spermatogonia-associated gene's transcripts in testes. There is no functional assay to conclude SRSF1 has an essential role in mouse testes. The data only indicate that SRSF1 may have a vital role in posttranscriptional regulation in the testes.
Thank you for the professional suggestions. Following this advice, we have corrected the text in this revised version (Page 4, Line 98 and 112).
- Line 141. In the mating scheme, Vasa-Cre Srsf1Fl/del …
Author Response
The following is the authors’ response to the original reviews.
Reviewer #1 (Recommendations For The Authors):
The conclusions of this paper are mostly well supported by data, but some aspects need to be corrected.
- Line 99. The title is not suitable for summarizing this part of the results. In this paragraph, the results mainly describe SRSF1 expression pattern and binding of spermatogonia-associated gene's transcripts in testes. There is no functional assay to conclude SRSF1 has an essential role in mouse testes. The data only indicate that SRSF1 may have a vital role in posttranscriptional regulation in the testes.
Thank you for the professional suggestions. Following this advice, we have corrected the text in this revised version (Page 4, Line 98 and 112).
- Line 141. In the mating scheme, Vasa-Cre Srsf1Fl/del mice should be obtained instead of Vasa-Cre Srsf1Fl/Fl mice.
Thank you for the professional suggestions. Following this advice, we have corrected the text in this revised version (Page 4, Line 118).
- Fig 2 C, "PZLF" should be corrected to "PLZF".
Thank you very much for the helpful comments. We have corrected this in Figure 2C.
- Fig 5 B, "VASA" and "Merge" should be interchanged.
Thank you very much for the helpful comments. We have interchanged "VASA" and "Merge" in Figure 5B.
- Fig 5 D, "Ctrl" should be added in the up panel.
Thank you very much for the helpful suggestions. We have added "Ctrl" in Figure 5C.
- The legend for Figure 6 D should be revised.
Thank you very much for the helpful suggestions. We have revised the legend for Figure 7D
- The legend for Figure 7 G should be revised.
Thank you very much for the helpful suggestions. We have revised the legend for Figure 8D
- Immunoprecipitation mass spectrometry (IP-MS) data showed that t SRSF1 interacts with other RNA splicing-related proteins (e.g., SRSF10, SART1, RBM15, SRRM2, SF3B6, and SF3A2). The authors should verify the interactions in testis or cells.
We thank the reviewer for the professional comments and suggestions. Following this advice, we performed co-transfection and co-IP to verify the protein-protein interactions in 293T cells, the results showed that the RRM1 domain of SRSF1 interacted with SART1, RBM15 and SRSF10 in 293T cells. In addition, the fluorescence results showed complete co-localization of mCherry-SRSF1 with eGFP-SART1, eGFP-RBM15 and eGFP-SRSF10 in 293T cells. Therefore, we have incorporated the data into the Figure 9G-J. Meanwhile, these have been incorporated into the text, given descriptions, and highlighted (Page 17, Lines 338-347).
- To avoid overstatement, the authors should pay attention to the use of adjectives and adverbs in the article, especially when drawing conclusions about the role of Tail1.
We thank the reviewer for the professional comments and suggestions. To avoid overstatement, we have revised the entire text (Page 4, Lines 98, and 112; Page 16, Lines 308; Page 17, Lines 346-347; Page 20, Lines 413-414; Page 21, Lines 432-433).
Reviewer #2 (Recommendations For The Authors):
Major
- I find the use of "SSC homing" misleading/confusing because this "homing" or relocation of postnatal gonocytes/nascent spermatogonia to the basement membrane precedes the maturation of the nascent spermatogonia into SSCs. In addition, "SSC homing" is commonly used in the SSC transplantation field to describe a transplanted SSC's ability to find and colonize its niche within the seminiferous tubules. I appreciate that "postnatal gonocytes/nascent spermatogonia homing" is not easily grasped by a broader audience. Perhaps "homing of precursor SSCs" is more appropriate.
Thank you very much for the helpful comments and suggestions. Following this advice, we have corrected the text in this revised version (Line 1-2, 39, 44, 49, 54-55, 68, 70, 72-73, 77, 84, 93-95, 191, 201, 240, 384-387, 397, 417-422, and 433)
- If I am misunderstanding the description of the Srsf1 cKO phenotype, and the authors truly believe SSCs have formed in the Srsf1 cKO testis, I strongly recommend immunostaining to show that the cKO germ cells robustly express SSC markers, not just markers of undifferentiated spermatogonia.
We thank the reviewer for the professional suggestions. We fully agree with the reviewer. Immunohistochemical staining for FOXO1 and statistical results indicated a reduced number of prospermatogonia (Figure 6C-E). So, we have corrected the text in this revised version (Line 1-2, 39, 44, 49, 54-55, 68, 70, 72-73, 77, 84, 93-95, 191, 201, 240, 384-387, 397, 417-422, and 433).
- If the authors have the available resources, the significance of this report would be enhanced by additional characterization of the cKO phenotype at the transition from gonocyte to nascent spermatogonia. Do any cKO germ cells exhibit defects in maturing from gonocytes to nascent spermatogonia at the molecular level? I.e., by P5-7, do all cKO germ cells express PLZF and localize FOXO1 to cytoplasm, as expected of nascent spermatogonia? If the cKO germ cells are actually a heterogenous population of gonocytes and nascent spermatogonia, what is the distribution of each subpopulation in the lumen vs basement membrane?
Thank you for the professional suggestions. Following this advice, immunohistochemical staining for FOXO1 was performed on 5 dpp mouse testis sections (Figure 6C). Further, germ cell statistics of FOXO1 expression in the nucleus showed a reduced number of prospermatogonia in cKO mice (Figure 6D). And germ cells in which FOXO1 is expressed in the nucleus similarly undergo abnormal homing (Figure 6E). Thus, all the above data indicated that SRSF1 has an essential role in the homing of precursor SSCs. we have incorporated the data into the Figure 6C-E. Meanwhile, these have been incorporated into the text, given descriptions, and highlighted (Page 9, Lines 191-201; Page 20, Lines 389-391).
Minor
- Could the authors clarify why Tial1 exon exclusion in the cKO results in reduced protein expression? Is it creating a transcript isoform that undergoes nonsense-mediated decay?
Thank you for the professional suggestions. Following this advice, we analyzed Tial1 transcripts again, and we found that Tial1 exon exclusion resulted in reduced expression of protein isoform X2 (Figure 8J). Since this region is not in the CDS, no clear evidence of nonsense-mediated decay was found in the analysis.
- Could the authors confirm that the TIAL1 antibody is not detecting the portion of the protein encoded by the alternatively spliced exon?
Thank you for the helpful comments. The TIAL1 monoclonal antibody is produced by Proteintech Group under the product number 66907-1-Ig. Immunogen is TIAL1 fusion protein Ag11981. The sequence is as follows. MDARVVKDMATGKSKGYGFVSFYNKLDAENAIVHMGGQWLGGRQIRTNWATRKPPAPKSTQENNTKQLRFEDVVNQSSPKNCTVYCGGIASGLTDQLMRQTFSPFGQIMEIRVFPEKGYSFVRFSTHESAAHAIVSVNGTTIEGHVVKCYWGKESPDMTKNFQQVDYSQWGQWSQVYGNPQQYGQYMANGWQVPPYGVYGQPWNQQGFGVDQSPSAAWMGGFGAQPPQGQAPPPVIPPPNQAGYGMASYQTQ The homology was 99% in mice and all TIAL1 isoforms were detected. So, TIAL1 antibody is detecting the portion of the protein encoded by the alternatively spliced exon.
- Lines 143: should "cKO" actually be "control"?
Thank you for the helpful suggestions. There is a real problem in the text description. we have corrected the text in this revised version (Page 6, Line 138-139).
- Lines 272-3 "visual analysis using IGV showed the peak of Tial1/Tiar was stabilized in 5 dpp cKO mouse testes (Figure 7H)": "peak stabilization" is not evident to me from the figure nor do I see Tial1 listed as differentially expressed in the supplemental. I would refrain from using IGV visualization as the basis for the differential abundance of a transcript.
Thank you very much for the helpful comments and suggestions. Tial1/Tiar is one of 39 stabilizing genes that are bound by SRSF1 and undergo abnormal AS. Following this advice, we have substituted Tial1/Tiar's FPKM for his peaks (Figure 8H). Meanwhile, we have corrected the text in this revised version (Page 15, Line 296-300; Page 16, Line 303-304).
- Lines 468-473: please clarify the background list used for GO enrichment analyses. By default, the genes expressed in the testis are enriched for spermatogenesis-related genes. To control for this and test whether a gene list is enriched for spermatogenesis-related genes beyond what is already seen in the testis, I recommend using a list of all expressed genes (for example, defined by TPM>=1) as the background list.
We thank the reviewer for the professional comments and suggestions. Following this advice, all expressed genes (TPM sum of all samples >=1) are listed background for GO enrichment analyses. The results of GO enrichment analysis of the AS gene turned out to be the same. The results of GO enrichment analysis of the SRSF1 peak-containing genes, differential genes, and IP proteins-associated genes have corrected in the figure (Figure 2A, 7E, and 9E)
- Figure 2B: Could the authors mark where the statistically significant peaks appear on the tracks? There are many small peaks and it's unclear if they are significant or not.
Thank you for the helpful suggestions. Following this advice, we have marked the areas of higher peaks in the figure (Figure 2B). We generally believe that any region above the peaks of IgG is likely to be a binding region, and of course, the higher the peak value, the more pre-mRNA is bound by SRSF1 in that region.
- Figure 7A: I assume the SRSF1 CLIP-seq genes are all the genes from the adult testis experiments. I would suggest limiting the CLIP-seq gene set to only those expressed in the P5 RNA-seq data, as if the target is not expressed at P5, there's no way it will be differentially expressed or differentially spliced in at P5.
Thank you very much for the helpful comments and suggestions. Following this advice, we found that 3543 of the 4824 genes bound by SRSF1 were expressed in testes at 5 dpp. we have corrected in the figure (Figure 8A). these have been incorporated into the text, given descriptions, and highlighted (Page 14, Lines 274-277).
- Figure 7F: Could the authors clarify where the alternatively spliced exon is relative to the total transcript, shown in 7H?
Thank you for the helpful suggestions. Following this advice, we have labeled the number of exons where variable splicing occurs. (Figure 8F).
- Please include where the sequencing and mass spec data will be publicly available.
Thank you very much for the helpful comments and suggestions. Following this advice, these have been incorporated into the text, given descriptions, and highlighted (Page 25, Lines 560-565).
Reviewer #3 (Recommendations For The Authors):
Suggestions for improving the data and analysis
- The claim that TIAL1 mediates SRSF1 effects is not well supported; this claim should be adjusted or additional supporting data should be provided. To support a claim that alternative splicing of Tial1 mediates the effects of SRSF1, at least two additional pieces of data are needed: first, a demonstration that the two alternative protein isoforms have different molecular functions, either in vitro or in vivo; and second, a better quantitation of the levels and ratios of expression of the two different isoforms in vivo.
Thank you for the helpful comments and suggestions. Following this advice, we quantified the expression levels and ratios of two different isoforms in vivo, and we found that Tial1 exon exclusion resulted in reduced expression of protein isoform X2 (Figure 8J). However, it is not possible to prove that the two alternative protein isoforms have different molecular functions. So, this claim has been adjusted in the text. these have been incorporated into the text, given descriptions, and highlighted (Lines 1-2, 43-45, 95, 306, 323-325, 408, 413-414).
- Likewise, the claim that "SRSF1 is required for "homing and self-renewal" of SSCs should be adjusted or better supported. As of now, the data supports a claim that SRSF1 is required for the establishment of the SSC population in the testis after birth. This could be due to defects in homing, self-renewal, or survival. To support claims about homing and self-renewal, these phenotypes should be tested more directly, for example by quantitating numbers of spermatogonia at the basal membrane in juvenile testes (homing) and expression of SSC markers in addition to the pan-germ cell marker VASA across early postnatal time points.
Thank you very much for the helpful comments and suggestions. Immunohistochemical staining for FOXO1 was performed on 5 dpp mouse testis sections (Figure 6C). Further, germ cell statistics of FOXO1 expression in the nucleus showed a reduced number of prospermatogonia in cKO mice (Figure 6D). And germ cells in which FOXO1 is expressed in the nucleus similarly undergo abnormal homing (Figure 6E). Thus, all the above data indicated that SRSF1 has an essential role in the homing of precursor SSCs. we have incorporated the data into the Figure 6C-E. These have been incorporated into the text, given descriptions, and highlighted (Page 9, Lines 191-201; Page 20, Lines 387-389). Meanwhile, "homing and self-renewal" of SSCs have corrected the text in this revised version (Line 1-2, 39, 44, 49, 54-55, 68, 70, 72-73, 77, 84, 93-95, 191, 201, 240, 384-387, 397, 417-422, and 433).
- Additional, more detailed analyses of CLIP-seq and RNA-seq data at least showing that the libraries are of good quality should be provided.
Thank you very much for suggestions. Following this advice, detailed analyses of RNA-seq data have been incorporated the data into the figures (Figure S2). But detailed analyses of CLIP-seq have already been used in another paper (Sun et al., 2023), and we have not provided it in order to avoid multiple uses of one figure. Meanwhile, we made a citation in the article (Page 4, Lines 105; Page 25, Lines 564-565).
- Gene Ontology analyses should be redone with a more appropriate background gene set.
Thank you for the helpful suggestions. All expressed genes (TPM sum of all samples >=1) are listed background for GO enrichment analyses. The results of GO enrichment analysis of the AS gene turned out to be the same. The results of GO enrichment analysis of the SRSF1 peak-containing genes, differential genes, and IP proteins-associated genes have been corrected in the figure (Figure 2A, 7E, and 9E)
Minor points about the text and figures
- The species (mouse) should be stated earlier in the Introduction.
Thank you for the professional suggestions. Following this advice, the mouse has been stated earlier in the Introduction (Page 3, Line 65).
- In Fig. 1C (Western blot), the results would be more convincing if quantitation of band intensities normalized to the loading control was added.
Thank you very much for comments and suggestions. Following this advice, ACTB served as a loading control. The value in 16.5 dpc testes were set as 1.0, and the relative values of testes in other developmental periods are indicated. Therefore, we have incorporated the data into the figures (Figure 1C).
- In Fig 5D, TUNEL signal in the single-channel image is difficult to see; please adjust the contrast.
Thank you for the professional suggestions. Following this advice, the images of the channels have been replaced by enlarged images for better visibility (Figure 5C).
Major comments
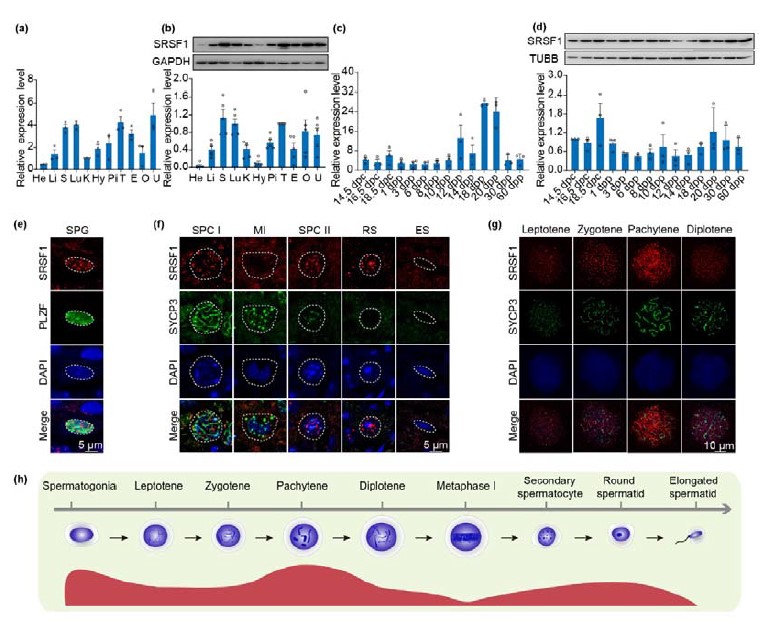
- In Fig 1D, it appears that SRSF1 is expressed most strongly in spermatogonia by immunofluorescence, but this is inconsistent with the sharp rise in expression detected by RT-qPCR at 20 days post partum (dpp) (Fig. 1B), which is when round spermatids are first added; this discrepancy should be explained or addressed.
We appreciate the important comments from the reviewer. In another of our studies, we showed that SRSF1 expression is higher in pachytene spermatocytes and round spermatids (Sun et al., 2023). So, it is normal for the sharp rise in expression detected by RT-qPCR at 20 days post partum (dpp).
Author response image 1.
Dynamic localization of SRSF1 in male mouse germ cells. (Sun et al., 2023)
- It is important to provide a more comprehensive basic description of the CLIP-seq datasets beyond what is shown in the tracks shown in Fig. 2B. This would allow a better assessment of the data quality and would also provide information about the transcriptome-wide patterns of SRSF1 binding. No information or quality metrics are provided about the libraries, and it is not stated how replicates are handled to maximize the robustness of the analysis. The distribution of peaks across exons, introns, and other genomic elements should also be shown.
Thank you very much for the helpful comments and suggestions. In fact, detailed analyses of CLIP-seq have already been presented in another paper (Sun et al., 2023), and we have not provided it in order to avoid multiple uses of one figure. Meanwhile, we made a citation in the article (Page 4, Lines 105; Page 25, Lines 564-565). In addition, the distribution of peaks in exons, introns, and other genomic elements is shown in Figure 2B.
- The claim that SRSF1 is required for "homing and self-renewal" of SSCs is made in multiple places in the manuscript. However, neither homing nor self-renewal is ever directly tested. A single image is shown in Fig. 5E of a spermatogonium at 5dpp that does not appropriately sit on the basal membrane, potentially indicating a homing defect, but this is not quantified or followed up. There is good evidence for depletion of spermatogonia starting at 7 dpp, but no further explanation of how homing and/or self-renewal fit into the phenotype.
Thank you very much for the helpful comments and suggestions. Following this advice, immunohistochemical staining for FOXO1 was performed on 5 dpp mouse testis sections (Figure 6C). Further, germ cell statistics of FOXO1 expression in the nucleus showed a reduced number of prospermatogonia in cKO mice (Figure 6D). And germ cells in which FOXO1 is expressed in the nucleus similarly undergo abnormal homing (Figure 6E). Thus, all the above data indicated that SRSF1 has an essential role in the homing of precursor SSCs. we have incorporated the data into the Figure 6C-E. These have been incorporated into the text, given descriptions, and highlighted (Page 9, Lines 191-201; Page 20, Lines 387-389). Meanwhile, "homing and self-renewal" of SSCs have corrected the text in this revised version (Line 1-2, 39, 44, 49, 54-55, 68, 70, 72-73, 77, 84, 93-95, 191, 201, 240, 384-387, 397, 417-422, and 433).
- In Fig. 6A (lines 258-260) very few genes downregulated in the cKO are bound by SRSF1 and undergo abnormal splicing. The small handful that falls into this overlap could simply be noise. A much larger fraction of differentially spliced genes are CLIP-seq targets (~33%), which is potentially interesting, but this set of genes is not explored.
Thank you for the helpful comments. Following this advice, this was specifically indicated by the fact that 39 stabilizing genes were bound by SRSF1 and underwent abnormal AS. In our study, Tial1/Tiar is one of 39 stabilizing genes that are bound by SRSF1 and undergo abnormal AS. Therefore, we fully agree with the reviewers' comments. These have been added in this revised version (Page 14, Lines 279-280; Page 15, Lines 296-300).
- The background gene set for Gene Ontology analyses is not specified. If these were done with the whole transcriptome as background, one would expect enrichment of spermatogenesis genes simply because they are expressed in testes. The more appropriate set of genes to use as background in these analyses is the total set of genes that are expressed in testis.
We thank the reviewer for the professional comments and suggestions. All expressed genes (TPM sum of all samples >=1) are listed background for GO enrichment analyses. The results of GO enrichment analysis of the AS gene turned out to be the same. The results of GO enrichment analysis of the SRSF1 peak-containing genes, differential genes, and IP proteins-associated genes have been corrected in the figure (Figure 2A, 7E, and 9E)
- In general, the model is over-claimed: aside from interactions by IP-MS, little is demonstrated in this study about how SRSF1 affects alternative splicing in spermatogenesis, or how alternative splicing of TIAL1 specifically would result in the phenotype shown. It is not clear why Tial1/Tiar is selected as a candidate mediator of SRSF1 function from among the nine genes that are downregulated in the cKO, are bound by SRSF1, and undergo abnormal splicing. Although TIAL1 levels are reduced in cKO testes by Western blot (Fig. 7J), this could be due just be due to a depletion of germ cells from whole testis. The reported splicing difference for Tial1 seems very subtle and the ratio of isoforms does not look different in the Western blot image.
Thank you very much for the helpful comments and suggestions. In our study, Tial1/Tiar is one of 39 stabilizing genes that are bound by SRSF1 and undergo abnormal AS. However, Western blotting showed that expression levels of TIAL1/TIAR isoform X2 were significantly suppressed (Figure 8J). So, the data indicate that SRSF1 is required for TIAL1/TIAR expression and splicing.
Sun, L., Chen, J., Ye, R., Lv, Z., Chen, X., Xie, X., Li, Y., Wang, C., Lv, P., Yan, L., et al. (2023). SRSF1 is crucial for male meiosis through alternative splicing during homologous pairing and synapsis in mice. Sci Bull 68, 1100-1104. 10.1016/j.scib.2023.04.030.
-

eLife assessment
In this valuable study, the authors characterize the role of splicing factor SRSF1 during spermatogenesis with a conditional knockout for Srsf1 in male mouse germ cells. The requirement of SRSF1 for maturation of postnatal gonocytes into spermatogonia, and the molecular role of SRSF1 in regulating alternative splicing in juvenile testes are convincingly supported. The paper also provides strong evidence that the mRNA encoding Tial, a factor relevant for spermatogonial maintenance and male fertility, is alternatively spliced in testis and that this splicing is regulated by SRSF1. The work will be of interest to reproductive biologists and stem cell biologists.
-

Joint Public Review:
Summary:
In this study, the authors seek to characterize the role of splicing factor SRSF1. Using a conditional deletion of Srsf1 in germ cells, they find that SRSF1 is required for male fertility. Via immunostaining and RNA-seq analysis of the Srsf1 conditional knockout (cKO) testes, combined with SRSF1 CLIP-seq and IP-MS data from the testis, they conclude that Srsf1 is required for homing of precursor spermatogonial stem cells (SCCs) due to alternative splicing of Tial1. They further show that spermatogonia-related genes (Plzf, Id4, Setdb1, Stra8, Tial1/Tiar, Bcas2, Ddx5, Srsf10, Uhrf1, and Bud31) were bound by SRSF1 in the mouse testes by CLIP-seq. They show that SRSF1 coordinates with other RNA splicing-related proteins to directly bind and regulate the expression of several spermatogonia-related genes, …Joint Public Review:
Summary:
In this study, the authors seek to characterize the role of splicing factor SRSF1. Using a conditional deletion of Srsf1 in germ cells, they find that SRSF1 is required for male fertility. Via immunostaining and RNA-seq analysis of the Srsf1 conditional knockout (cKO) testes, combined with SRSF1 CLIP-seq and IP-MS data from the testis, they conclude that Srsf1 is required for homing of precursor spermatogonial stem cells (SCCs) due to alternative splicing of Tial1. They further show that spermatogonia-related genes (Plzf, Id4, Setdb1, Stra8, Tial1/Tiar, Bcas2, Ddx5, Srsf10, Uhrf1, and Bud31) were bound by SRSF1 in the mouse testes by CLIP-seq. They show that SRSF1 coordinates with other RNA splicing-related proteins to directly bind and regulate the expression of several spermatogonia-related genes, including Tial1/Tiar, via alternative splicing Ultimately, the study shows that SRSF1's effects on alternative splicing are required to establish spermatogenesis. In the absence of Srsf1, the postnatal gonocytes do not properly mature into spermatogonia and consequently never initiate spermatogenesis.Strengths:
This study shows a role of SRSF1-mediated alternative splicing in establishment and survival of precursor SSCs, which may provide a framework to elucidate the molecular mechanisms of the posttranscriptional network underlying the formation of SSC pools. The histological analysis of the Srsf1 cKO traces the origins of the fertility defect to the postnatal testis, and the authors have generated interesting CLIP-seq, IP-MS, and RNA-seq datasets characterizing SRSF1's RNA targets and interacting proteins specifically in the testis. Together, this study provides detailed phenotyping of the Srsf1 cKO, which convincingly supports the Sertoli Cell Only phenotype, establishes the timing of the first appearance of the spermatogonial defect, and provides new insight into the role of splicing factors and SRSF1 specifically in spermatogenesis. The experiments are well-designed and conducted, the overall methods and results are robust and convincing.Weaknesses:
This study does not provide a full mechanistic explanation connecting altered splicing with defects in SSC precursors. The claim that altered splicing of the Tial1 transcript mediates the effect of SRSF1 loss is not convincingly supported. In addition, some regions of the text suggest that misregulated splicing of Tial1 disrupts spermatogonial survival; while Tial1 is required for primordial germ cell survival in embryonic gonads (E11.5-13.5; Beck et al 1998), it is unclear if Tial1 is required for germline development beyond this embryonic stage. -

-

eLife assessment
In this valuable study, the authors seek to characterize the role of splicing factor SRSF1 during spermatogenesis with a conditional knockout for Srsf1 in male germ cells. The spermatogenesis phenotype is convincingly supported, but two central claims of the study, that SRSF1 is required for spermatogonial homing and self-renewal and that this function is mediated by regulation of splicing of the gene Tial1, are inadequately supported. Support is inadequate because homing and self-renewal phenotypes were not explicitly tested, and functional data were not provided to support a role for alternative splicing of Tial1 in the fertility phenotype. The work will be of interest to reproduction and stem cell biologists.
-

Reviewer #1 (Public Review):
The authors revealed that spermatogonia-related genes (e.g., Plzf, Id4, Setdb1, Stra8, Tial1/Tiar, Bcas2, Ddx5, Srsf10, Uhrf1, and Bud31) were bound by SRSF1 in the mouse testes by Crosslinking immunoprecipitation and sequencing (CLIP-seq). Using Vasa-cre mouse line, the authors successfully evidenced that SRSF1 in the testis is essential for homing and self-renewal in mouse spermatogonial stem cells. Further evidence showed that SRSF1 directly binds and regulates the expression of Tial1/Tiar via AS to implement SSC homing and self-renewal. Immunoprecipitation mass spectrometry (IP-MS) data showed that the AS of SSC is regulated by SRSF1 coordinated with other RNA splicing-related proteins (e.g., SRSF10, SART1, RBM15, SRRM2, SF3B6, and SF3A2). The authors revealed the critical role of SRSF1-mediated AS in …
Reviewer #1 (Public Review):
The authors revealed that spermatogonia-related genes (e.g., Plzf, Id4, Setdb1, Stra8, Tial1/Tiar, Bcas2, Ddx5, Srsf10, Uhrf1, and Bud31) were bound by SRSF1 in the mouse testes by Crosslinking immunoprecipitation and sequencing (CLIP-seq). Using Vasa-cre mouse line, the authors successfully evidenced that SRSF1 in the testis is essential for homing and self-renewal in mouse spermatogonial stem cells. Further evidence showed that SRSF1 directly binds and regulates the expression of Tial1/Tiar via AS to implement SSC homing and self-renewal. Immunoprecipitation mass spectrometry (IP-MS) data showed that the AS of SSC is regulated by SRSF1 coordinated with other RNA splicing-related proteins (e.g., SRSF10, SART1, RBM15, SRRM2, SF3B6, and SF3A2). The authors revealed the critical role of SRSF1-mediated AS in SSC homing and self-renewal, which may provide a framework to elucidate the molecular mechanisms of the posttranscriptional network underlying the formation of SSC pools and the establishment of niches. The experiments are well-designed and conducted, the overall conclusions are convincing. This work will be of interest to stem cell and reproductive biologists.
-

Reviewer #2 (Public Review):
Summary
The authors seek to characterize the role of splicing factor SRSF1 during spermatogenesis. Using a conditional deletion of Srsf1 in germ cells, they find that SRSF1 is required for male fertility. Via immunostaining and RNA-seq analysis of the Srsf1 conditional knockout (cKO) testes, combined with SRSF1 CLIP-seq and IP-MS data from the testis, they ultimately conclude that Srsf1 is required for spermatogonial stem cell (SSC) homing and self-renewal due to alternative splicing of Tial1.Strengths
The overall methods and results are robust. The histological analysis of the Srsf1 cKO traces the origins of the fertility defect to the postnatal testis, and the authors have generated interesting datasets characterizing SRSF1's RNA targets and interacting proteins specifically in the testis.Ultimately, the …
Reviewer #2 (Public Review):
Summary
The authors seek to characterize the role of splicing factor SRSF1 during spermatogenesis. Using a conditional deletion of Srsf1 in germ cells, they find that SRSF1 is required for male fertility. Via immunostaining and RNA-seq analysis of the Srsf1 conditional knockout (cKO) testes, combined with SRSF1 CLIP-seq and IP-MS data from the testis, they ultimately conclude that Srsf1 is required for spermatogonial stem cell (SSC) homing and self-renewal due to alternative splicing of Tial1.Strengths
The overall methods and results are robust. The histological analysis of the Srsf1 cKO traces the origins of the fertility defect to the postnatal testis, and the authors have generated interesting datasets characterizing SRSF1's RNA targets and interacting proteins specifically in the testis.Ultimately, the authors have shown that SRSF1's effects on alternative splicing are required to establish spermatogenesis. In the absence of Srsf1, the postnatal gonocytes/nascent spermatogonia do not properly relocate from the seminiferous tubules' lumen to basement membrane, and consequently, never initiate spermatogenesis. I believe this relocation event is what the authors are referring to as "SSC homing".
Weaknesses
I do not think there is enough evidence to support two major conclusions. First, the authors conclude that SRSF1 is required for "SSC self-renewal." Given the defect in nascent spermatogonial development, it is not evident to me that SSCs actually form in the Srsf1 cKO. Second, the authors conclude that SRSF1 controls alternative splicing of Tial1 to "implement SSC homing and self-renewal." I'm unsure as to the basis for TIAL1's role in "SSC homing and self-renewal," particularly as the only reference provided about Tial1's effects on the germ line shows that germ cells are completely lost during embryonic gestation. As a result, it's ultimately unclear to me how SRSF1 mechanistically regulates the relocation of gonocytes/nascent spermatogonia to the basement membrane.Alternative splicing is quite pronounced in the testis relative to other tissues. The authors have presented interesting work that shows that alternative splicing is required in the testicular germ line for the establishment of spermatogenesis.
-

Reviewer #3 (Public Review):
In this study, Sun et al examine the role of the splicing factor SRSF1 in spermatogenesis in mice. Alternative splicing is important for spermatogenic development, but its regulation and major developmental roles during spermatogenesis are not well understood. The authors set out to better define both SRSF1 function in testes and the contribution of alternative splicing. They collect several large 'omics datasets to define SRSF1 targets in testis, including RNA interactions by CLIP-seq in whole testis, protein interactions by IP-mass spec in whole testis, and RNA sequencing to detect expression levels and splice variants. They also examine the phenotype of germline conditional knockouts (cKO) for Srsf1, using the early-acting Vasa-Cre, and find a severe depletion of germ cells starting at 7 days post partum …
Reviewer #3 (Public Review):
In this study, Sun et al examine the role of the splicing factor SRSF1 in spermatogenesis in mice. Alternative splicing is important for spermatogenic development, but its regulation and major developmental roles during spermatogenesis are not well understood. The authors set out to better define both SRSF1 function in testes and the contribution of alternative splicing. They collect several large 'omics datasets to define SRSF1 targets in testis, including RNA interactions by CLIP-seq in whole testis, protein interactions by IP-mass spec in whole testis, and RNA sequencing to detect expression levels and splice variants. They also examine the phenotype of germline conditional knockouts (cKO) for Srsf1, using the early-acting Vasa-Cre, and find a severe depletion of germ cells starting at 7 days post partum (dpp) and culminating with a lack of germ cells (Sertoli Cell Only Syndrome) by adulthood. They detect differences in gene expression as well as differences in splicing between control and knockout, including 9 genes that are downregulated, experience alternative splicing, and whose transcripts are also bound by SRSF1, and identify the Tial1/Tiar transcript as one of these targets. They conclude that SRSF1 is required for homing and self-renewal of spermatogonial stem cells, at least in part through its regulation of Tial1/Tiar splicing.
Strengths of the paper include detailed phenotyping of the Srsf1 cKO, which convincingly supports the Sertoli Cell Only phenotype, establishes the timing of the first appearance of the spermatogonial defect, and provides new insight into the role of splicing factors and SRSF1 specifically in spermatogenesis. Another strength is the generation of CLIP-seq, IP-MS, and RNA-seq datasets which will be a useful resource for the field of germ cell development. Major weaknesses include a lack of robust support for two major claims: first, there is inadequate support for the claim of defects in either "homing" or "self-renewal" of spermatogonia in the cKO, and second, there is inadequate support for the claim that altered splicing of the Tial1 transcript mediates the effect of SRSF1 loss. A moderate weakness is the superficial discussion of the CLIP, RNA-seq, and IP-MS datasets, limiting their otherwise high utility for other researchers. Overall, the paper as it stands will have a moderate impact on the field of male reproductive biology. Specific points that should be addressed to improve support for the claims are below.
Major comments
In Fig 1D, it appears that SRSF1 is expressed most strongly in spermatogonia by immunofluorescence, but this is inconsistent with the sharp rise in expression detected by RT-qPCR at 20 days post partum (dpp) (Fig. 1B), which is when round spermatids are first added; this discrepancy should be explained or addressed.
It is important to provide a more comprehensive basic description of the CLIP-seq datasets beyond what is shown in the tracks shown in Fig. 2B. This would allow a better assessment of the data quality and would also provide information about the transcriptome-wide patterns of SRSF1 binding. No information or quality metrics are provided about the libraries, and it is not stated how replicates are handled to maximize the robustness of the analysis. The distribution of peaks across exons, introns, and other genomic elements should also be shown.
The claim that SRSF1 is required for "homing and self-renewal" of SSCs is made in multiple places in the manuscript. However, neither homing nor self-renewal is ever directly tested. A single image is shown in Fig. 5E of a spermatogonium at 5dpp that does not appropriately sit on the basal membrane, potentially indicating a homing defect, but this is not quantified or followed up. There is good evidence for depletion of spermatogonia starting at 7 dpp, but no further explanation of how homing and/or self-renewal fit into the phenotype.
In Fig. 6A (lines 258-260) very few genes downregulated in the cKO are bound by SRSF1 and undergo abnormal splicing. The small handful that falls into this overlap could simply be noise. A much larger fraction of differentially spliced genes are CLIP-seq targets (~33%), which is potentially interesting, but this set of genes is not explored.
The background gene set for Gene Ontology analyses is not specified. If these were done with the whole transcriptome as background, one would expect enrichment of spermatogenesis genes simply because they are expressed in testes. The more appropriate set of genes to use as background in these analyses is the total set of genes that are expressed in testis.
In general, the model is over-claimed: aside from interactions by IP-MS, little is demonstrated in this study about how SRSF1 affects alternative splicing in spermatogenesis, or how alternative splicing of TIAL1 specifically would result in the phenotype shown. It is not clear why Tial1/Tiar is selected as a candidate mediator of SRSF1 function from among the nine genes that are downregulated in the cKO, are bound by SRSF1, and undergo abnormal splicing. Although TIAL1 levels are reduced in cKO testes by Western blot (Fig. 7J), this could be due just be due to a depletion of germ cells from whole testis. The reported splicing difference for Tial1 seems very subtle and the ratio of isoforms does not look different in the Western blot image.
-