A pH-sensitive switch activates virulence in Salmonella
Curation statements for this article:-
Curated by eLife
eLife assessment
Salmonella invades and survives in host cells via SPI-1 and SPI-2 type III secretion system mechanisms, with the SPI-2 system allowing for intracellular survival in Salmonella-containing vacuoles, which have a low-pH environment. Transcription of SPI-2 genes at low pH is activated by the DNA-binding SsrB protein, which sits at the top of the SPI-2 regulatory hierarchy. This study provides important insights as to how SsrB is allosterically affected by pH resulting in acid-dependent DNA binding. However, there are concerns about some experiments, and the evidence presented is not fully conclusive.
This article has been Reviewed by the following groups
Discuss this preprint
Start a discussion What are Sciety discussions?Listed in
- Evaluated articles (eLife)
Abstract
The transcriptional regulator SsrB acts as a switch between virulent and biofilm lifestyles of non-typhoidal Salmonella enterica serovar Typhimurium. During infection, phosphorylated SsrB activates genes on Salmonella Pathogenicity Island-2 (SPI-2) essential for survival and replication within the macrophage. Low pH inside the vacuole is a key inducer of expression and SsrB activation. Previous studies demonstrated an increase in SsrB protein levels and DNA-binding affinity at low pH; the molecular basis was unknown (Liew et al., 2019). This study elucidates its underlying mechanism and in vivo significance. Employing single-molecule and transcriptional assays, we report that the SsrB DNA-binding domain alone (SsrBc) is insufficient to induce acid pH-sensitivity. Instead, His12, a conserved residue in the receiver domain confers pH sensitivity to SsrB allosterically. Acid-dependent DNA binding was highly cooperative, suggesting a new configuration of SsrB oligomers at SPI-2-dependent promoters. His12 also plays a role in SsrB phosphorylation; substituting His12 reduced phosphorylation at neutral pH and abolished pH-dependent differences. Failure to flip the switch in SsrB renders Salmonella avirulent and represents a potential means of controlling virulence.
Article activity feed
-

-

Author Response
Reviewer #2 (Public Review):
Reviewer #2 was critical of every aspect of our manuscript and we were disappointed that they failed to appreciate the significance of our findings. However, we have responded to each point as described below:
- The experiment displayed in Figure 5 is deeply flawed for multiple reasons and should be removed from the manuscript entirely. A Michaelis-Menton plot compares the initial rate of a reaction versus substrate concentration. Instead, the authors plotted the fraction of SsrB that is phosphorylated after 10 minutes at various substrate concentrations. Such a plot must reach saturation because the enzyme is limiting, whereas it is not always possible to achieve saturation in a genuine Michaelis-Menton plot. Because no reaction rates were measured, it is not possible to derive kcat values …
Author Response
Reviewer #2 (Public Review):
Reviewer #2 was critical of every aspect of our manuscript and we were disappointed that they failed to appreciate the significance of our findings. However, we have responded to each point as described below:
- The experiment displayed in Figure 5 is deeply flawed for multiple reasons and should be removed from the manuscript entirely. A Michaelis-Menton plot compares the initial rate of a reaction versus substrate concentration. Instead, the authors plotted the fraction of SsrB that is phosphorylated after 10 minutes at various substrate concentrations. Such a plot must reach saturation because the enzyme is limiting, whereas it is not always possible to achieve saturation in a genuine Michaelis-Menton plot. Because no reaction rates were measured, it is not possible to derive kcat values from the data.
Mea culpa. We now plot our phosphorylation data and describe the mid-point as a k0.5 and have removed Fig. 1g. When we directly compare the H12 mutant to wt at neutral pH, its phosphorylation level is less compared to the wt (see new Fig. 4a). The wt phosphorylation is reduced at acid pH, (Fig 4b), but with His12Q, there was no difference in phosphorylation between neutral and acid pH (Fig 4c). It is important to include this data, because in RcsB, a close homolog of SsrB, an H12A mutant was not phosphorylated by acetyl phosphate and it was incapable of binding to DNA, unlike what we show here with SsrB.
(i) Increasing the concentration of the phosphoramidite substrate increased ionic strength. Response regulator active sites contain many charged moieties and autophosphorylation of at least one response regulator (CheY) is inhibited by increasing ionic strength (PMID 10471801).
The reviewer raises some interesting points and they are based on CheY phosphorylation by small molecules. We have a long history of studying OmpR and SsrB as well as other RRs and we know that they can all behave very differently from “canonical signaling”. We examined the effect of ionic strength on SsrB phosphorylation and it was relatively insensitive to changes in ionic strength (our original buffer was 267-430 mOsm and in each case, we have 90% phosphorylation). However, we repeated all of the phosphorylation experiments and kept ionic strength constant. These data are now presented in the revised manuscript.
(ii) Autophosphorylation with phosphoramidite is pH dependent because the nitrogen on the donor must be protonated to form a good leaving group (PMID 9398221). The pKa of phosphoramidite is ~8. Therefore, the fraction of phosphoramidite that is reactive (i.e., protonated) will be very different at pH 6.1 and 7.4.
We are aware of those findings, but we are comparing the H12 mutant with the wt protein in each case. There is no reason to believe that the presence of the mutant should alter the phosphoramidate substrate, so we are comparing how the wt phosphorylation compares with the mutant (Fig 4b, c).
(iii) Response regulator autophosphorylation absolutely depends on the presence of a divalent metal ion (usually Mg2+) in the active site (PMID 2201404). There is no guarantee that the 20 mM Mg2+ included in the reaction is sufficient to saturate SsrB. Furthermore, as the authors themselves note, the amino acid at SsrB position 12 is likely to affect the affinity of Mg2+ binding. Therefore, the fraction of SsrB that is reactive (i.e. has Mg2+ bound) may differ between wildtype and the H12Q mutant, and/or between wildtype at different pHs (because the protonation state of His12 changes).
This is exactly the point that we are making. And why we varied the magnesium concentration (increasing to 50-100 mM). There was a slight increase in phosphorylation at 50 mM MgCl2 compared to 20 mM, and only a slight increase between 50 and 100 mM at pH 6.1. The revised phosphorylation experiments all contain 100 mM MgCl2.
- The data in Figures 1abcd and 3de are clearly sigmoidal rather than hyperbolic, indicating cooperativity. However, there are insufficient data points between the upper and lower bounds to accurately calculate the Hill coefficient or KD values. This limitation of the data means that comparisons of apparent Hill coefficient or KD values under different conditions cannot be the basis of credible conclusions.
We respectfully disagree. In every curve that we provide, there is at least one data point in the transition between low and high binding. With the mutant H12Q, we did manage to get two data points in the transition and the KD was the same as the wildtype (Fig. 2). We provide an analysis of the binding curve which nicely demonstrates the range of KD values based on the lowest and highest error in the point (132-168 nM) and it doesn’t significantly change the value (this is now shown in Fig.1– figure supplement 1). The very high affinity we observed at pH 6.1 (KD ~5 nM) makes the range of possibilities between 4-8 nM (i.e. still VERY high affinity). These range in affinities at neutral and acid pH are very reminiscent of affinities we measured for OmpR and OmpR~P at the porin promoters, suggesting that acid pH puts SsrB in an activated state even in the absence of phosphorylation. A similar argument holds for the Hill coefficient (see Figure).
- There are hundreds of receiver domain structures in PDB. There is some variation, but to a first approximation receiver domain structures, all exhibit an (alpha/beta)5 fold. The structure of SsrB predicted by i-TASSER breaks the standard beta-2 strand into two parts, which throws off the numbering for subsequent beta strands. Given the highly conserved receiver domain fold, I am skeptical that the predicted i-TASSER structure is correct or adds any value to the manuscript. If the authors wish to retain the structure of the manuscript, then they should point out the unusual feature and the consequence of strand numbering.
We now include a new model based on the RcsB/DNA crystal structure that eliminates this problem (see new Fig.2– figure supplement 2). We have replaced this model with an Alphafold prediction that was energy minimized to align with the RcsB dimer crystal structure (Fig.5– figure supplement 2). This model retains the original (beta/alpha)5 fold, so the classical numbering is retained.
- The detailed predictions of active site structure in Supplementary Figure 5 are not physiologically relevant because Mg2+ was not included in the simulation. The presence of a divalent cation binding to Asp10 and Asp11 is likely to substantially alter interactions between Asp 10, Asp11, His12, and Lys109.
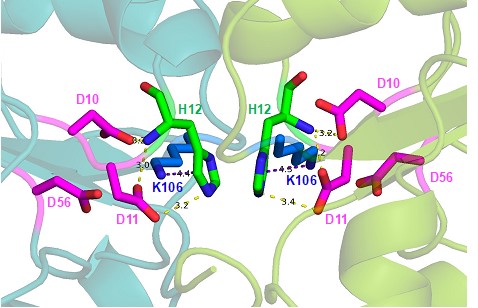
See response to 1iii, above and new Fig.5– figure supplement 2. Author response image 1 is a zoomed-in snapshot of supplementary Figure 8c that has been modelled using the RcsB dimer bound to BeF3 and Mg2+(6ZIX). Both the i-TASSER and Alphafold model receiver domains align well with this structure, and the polar contacts and pi-cation interactions made by His12 are maintained.
Author response image 1.
- The authors present an AlphaFold model of an SsrB dimer, and note that His12 is at the dimer interface. However, the authors also believe that a higher-order oligomer of SsrB binds to DNA in a pH-dependent manner. Do the authors have any suggestions or informed speculation about how His12 might affect higher-order oligomerization than dimerization?
As mentioned to point 3, above, we now include a new model of an SsrB dimer bound to DNA based on our NMR structure of the CTD and the RcsB/DNA structure. In the RcsB paper, they also have evidence for a higher-order oligomer in the crystal structure of unphosphorylated (and BeF3-) RcsB, which showed an asymmetric unit containing 6 molecules of RcsB, which form 3 dimers arranged in a hexameric structure that resembles a cylinder. This configuration involves a crossed conformation with the REC of one molecule interacting with the DBD of another and interestingly, His12 is interacting with the DBD of another molecule. We modelled an SsrB oligomer structure using the RcsB hexamer as a template and have included it as a new figure (see Fig.5– figure supplement 3) and in the revised discussion (lines 432-448).
-

eLife assessment
Salmonella invades and survives in host cells via SPI-1 and SPI-2 type III secretion system mechanisms, with the SPI-2 system allowing for intracellular survival in Salmonella-containing vacuoles, which have a low-pH environment. Transcription of SPI-2 genes at low pH is activated by the DNA-binding SsrB protein, which sits at the top of the SPI-2 regulatory hierarchy. This study provides important insights as to how SsrB is allosterically affected by pH resulting in acid-dependent DNA binding. However, there are concerns about some experiments, and the evidence presented is not fully conclusive.
-

Reviewer #1 (Public Review):
The strength of this work is the quality and quantity of data, which identify a critical histidine residue, His12 of SsrB, that is responsible for the allosteric, pH-dependent conformational change in SsrB and for phosphorylation of SsrB. That is the fundamental question to the field: the low pH response when Salmonella invades host cells and utilizes acidification within a Salmonella-containing vacuole as a signal to initiate the expression of virulence genes from the Salmonella pathogenicity island 2 (SPI-2) suite of virulence genes, which encode specific effector proteins and a unique secretion injectisome that has, to date, eluded purification. The SsrB protein will activate the transcription of non-SPI-2 genes at neutral pH in the regulation of biofilm formation. The low pH, phosphorylated SsrB …
Reviewer #1 (Public Review):
The strength of this work is the quality and quantity of data, which identify a critical histidine residue, His12 of SsrB, that is responsible for the allosteric, pH-dependent conformational change in SsrB and for phosphorylation of SsrB. That is the fundamental question to the field: the low pH response when Salmonella invades host cells and utilizes acidification within a Salmonella-containing vacuole as a signal to initiate the expression of virulence genes from the Salmonella pathogenicity island 2 (SPI-2) suite of virulence genes, which encode specific effector proteins and a unique secretion injectisome that has, to date, eluded purification. The SsrB protein will activate the transcription of non-SPI-2 genes at neutral pH in the regulation of biofilm formation. The low pH, phosphorylated SsrB structure allows for cooperative binding to DNA that is necessary for SPI-2 gene activation. Remarkably, the substitution of the single His12 residue of SsrB is enough to eliminate its activity at acidic pH, but not at normal pH. The authors employ a clever and exceptional single-molecule DNA unzipping assay for their DNA affinity measurements. Another major strength of this work is the logical flow of the results section and the lucidity of the written presentation. This work will guide the field in allowing for the expression of SPI-2 in the lab for mechanistic studies that would be otherwise impossible to do within a vacuole.
The first chapter of the results section includes the demonstration that acid pH increases SsrB affinity for SPI-2 promoter DNA. The authors employed a sophisticated single molecule DNA unzipping to measure the effects of pH on SsrB affinity to the DNA target. The DAN affinity was ~32-fold higher at acid pH (6.1) than at neutral pH (7.4). At both acidic and neutral pH conditions DNA binding was highly cooperative.
In the second results chapter, the authors investigated whether the DAN binding domain of SsrB was responsible for low pH-stimulated DNA binding. SsrB is a classic two-component regulatory protein with an N-terminal receiver domain that gets phosphorylated during activation and a C-terminal DNA binding domain to affect the regulation of gene expression in response to phosphorylation. Again, the single molecule DNA unzipping assay was employed to characterize pH effects on just the C-terminal binding domain (SsrB-C). The isolated C-terminal domain bound DNA with a 4-fold lower affinity as compared to the full-length protein. Cooperativity was also reduced. SsrB-C was shown to be unable to support acid-stimulation of SPI-2 transcription using both in vivo and in vitro transcriptional assays. The data is quite solid.
The third results chapter is a comparison of SsrB to other members of the NarL/FixJ subfamily of response regulators. SsrB is the only member to have known pH dependence on its activity. The authors found SsrB to have the highest pI of the subfamily and the second-greatest number of histidine residues. Of four histidine residues in the receiver domain His12 was conserved in the subfamily, while His28, His34, and His 72 were unique to SsrB and thus initially investigated. Since histidine residues are known to play a role in pH sensing, the three histidine residues in the receiver domain were extensively characterized for a potential role in pH-dependent transcriptional activation. The experiments ruled out the role of the three unique histidine residues in the SsrB receiver domain in pH sensing.
The fourth research chapter demonstrated that it is the conserved His14 of SsrB that is responsible for pH sensing. A striking result was the finding that the H12Q substitution retained full DNA binding activity at neutral pH, but at acidic pH, the H12Q allele was unable to activate SPI-2 transcription. Further analysis showed the mutant allele was defective in subunit cooperativity.
The fifth research chapter characterized other amino acid substitutions at His12 of SsrB. Positively charged substitutions were employed to mimic the protonated state of His12 and aromatic substitutions were chosen to mimic the aromatic nature of the imidazole ring of histidine. H12Y and H12F substitutions had substantially reduced activity but retained pH sensing. Charged substitutions were defective for both binding and pH sensing. These results support the conclusion that the aromatic nature of the histidine imidazole role was important for pH sensing.
In the final research chapter, the authors characterized His 12 substitutions for effects on SsrB phosphorylation at Asp56. The results of these assays showed that substitution at His12 reduced both SsrB phosphorylation at neutral pH and abolished pH-dependent changes in SsrB phosphorylation consistent with conformational changes in SsrB as a result of substitution at His12.
Overall, a solid study that defines the essential role of His12 in SsrB activation at low pH. His12 is critical for pH sensing, SsrB phosphorylation, SsrB oligomerization, and in vivo Salmonella virulence.
-

Reviewer #2 (Public Review):
The authors seek to explore the mechanistic basis for enhancement binding to DNA by SsrB at lower pH. Their evidence supports the conclusions listed in the Evaluation Summary. Multiple additional conclusions are not supported by the data as described below:
1. The experiment displayed in Figure 5 is deeply flawed for multiple reasons and should be removed from the manuscript entirely. A Michaelis-Menton plot compares the initial rate of a reaction versus substrate concentration. Instead, the authors plotted the fraction of SsrB that is phosphorylated after 10 minutes at various substrate concentrations. Such a plot must reach saturation because the enzyme is limiting, whereas it is not always possible to achieve saturation in a genuine Michaelis-Menton plot. Because no reaction rates were measured, it is not …
Reviewer #2 (Public Review):
The authors seek to explore the mechanistic basis for enhancement binding to DNA by SsrB at lower pH. Their evidence supports the conclusions listed in the Evaluation Summary. Multiple additional conclusions are not supported by the data as described below:
1. The experiment displayed in Figure 5 is deeply flawed for multiple reasons and should be removed from the manuscript entirely. A Michaelis-Menton plot compares the initial rate of a reaction versus substrate concentration. Instead, the authors plotted the fraction of SsrB that is phosphorylated after 10 minutes at various substrate concentrations. Such a plot must reach saturation because the enzyme is limiting, whereas it is not always possible to achieve saturation in a genuine Michaelis-Menton plot. Because no reaction rates were measured, it is not possible to derive kcat values from the data. There are also at least three potential problems with the reaction conditions themselves: (i) Increasing the concentration of the phosphoramidite substrate increased ionic strength. Response regulator active sites contain many charged moieties and autophosphorylation of at least one response regulator (CheY) is inhibited by increasing ionic strength (PMID 10471801). (ii) Autophosphorylation with phosphoramidite is pH dependent because the nitrogen on the donor must be protonated to form a good leaving group (PMID 9398221). The pKa of phosphoramidite is ~8. Therefore, the fraction of phosphoramidite that is reactive (i.e., protonated) will be very different at pH 6.1 and 7.4. (iii) Response regulator autophosphorylation absolutely depends on the presence of a divalent metal ion (usually Mg2+) in the active site (PMID 2201404). There is no guarantee that the 20 mM Mg2+ included in the reaction is sufficient to saturate SsrB. Furthermore, as the authors themselves note, the amino acid at SsrB position 12 is likely to affect the affinity of Mg2+ binding. Therefore, the fraction of SsrB that is reactive (i.e. has Mg2+ bound) may differ between wildtype and the H12Q mutant, and/or between wildtype at different pHs (because the protonation state of His12 changes).
2. The data in Figures 1abcd and 3de are clearly sigmoidal rather than hyperbolic, indicating cooperativity. However, there are insufficient data points between the upper and lower bounds to accurately calculate the Hill coefficient or KD values. This limitation of the data means that comparisons of apparent Hill coefficient or KD values under different conditions cannot be the basis of credible conclusions.
3. There are hundreds of receiver domain structures in PDB. There is some variation, but to a first approximation receiver domain structures, all exhibit an (alpha/beta)5 fold. The structure of SsrB predicted by i-TASSER breaks the standard beta-2 strand into two parts, which throws off the numbering for subsequent beta strands. Given the highly conserved receiver domain fold, I am skeptical that the predicted i-TASSER structure is correct or adds any value to the manuscript. If the authors wish to retain the structure of the manuscript, then they should point out the unusual feature and the consequence of strand numbering.
4. The detailed predictions of active site structure in Supplementary Figure 5 are not physiologically relevant because Mg2+ was not included in the simulation. The presence of a divalent cation binding to Asp10 and Asp11 is likely to substantially alter interactions between Asp 10, Asp11, His12, and Lys109.
5. The authors present an AlphaFold model of an SsrB dimer, and note that His12 is at the dimer interface. However, the authors also believe that a higher-order oligomer of SsrB binds to DNA in a pH-dependent manner. Do the authors have any suggestions or informed speculation about how His12 might affect higher-order oligomerization than dimerization?
-

Reviewer #3 (Public Review):
Once inside a cellular vacuole, Salmonella senses the low pH and activates the transcriptional regulator SsrB to induce expression of the Salmonella pathogenicity island 2 genes that are essential for intracellular survival and replication inside the host. This study investigates the mechanisms by which SsrB senses the pH changes, and with a series of elegant experiments identify a conserved residue in the receiver domain, His12, as essential for pH sensing and Salmonella virulence.
Overall, this study identifies an important mechanism of pathogen virulence, which could be targeted to control intracellular replication of the pathogen. The experiments are well conducted, the manuscript is clearly written, and the data are convincing and well presented. The authors perform a logical and detailed analysis of …
Reviewer #3 (Public Review):
Once inside a cellular vacuole, Salmonella senses the low pH and activates the transcriptional regulator SsrB to induce expression of the Salmonella pathogenicity island 2 genes that are essential for intracellular survival and replication inside the host. This study investigates the mechanisms by which SsrB senses the pH changes, and with a series of elegant experiments identify a conserved residue in the receiver domain, His12, as essential for pH sensing and Salmonella virulence.
Overall, this study identifies an important mechanism of pathogen virulence, which could be targeted to control intracellular replication of the pathogen. The experiments are well conducted, the manuscript is clearly written, and the data are convincing and well presented. The authors perform a logical and detailed analysis of several portions of SsrB to finally identify His12 as a key residue for pH sensing. This was not an easy task. Moreover, the fact that a single amino acid appears to be so important for SsrB pH sensing and SsrB phosphorylation is an important finding for potentially targeting SsrB and inhibiting Salmonella virulence.
-