Metabolic consequences of various fruit-based diets in a generalist insect species
Curation statements for this article:-
Curated by eLife
eLife assessment
This useful study uses untargeted metabolomics to help us understand how some herbivores are able to be generalists, rather than specializing in the metabolism of specific plant species. This is an important area, since little is known about how generalist insect species metabolize their food. In its current form, the study lacks ecological relevance due to the exclusive use of refined sampling procedures, and the metabolomic analysis is incomplete.
This article has been Reviewed by the following groups
Discuss this preprint
Start a discussion What are Sciety discussions?Listed in
- Evaluated articles (eLife)
Abstract
Most phytophagous insect species exhibit a limited diet breadth and specialize on a few or a single host plant. In contrast, some species display a remarkably large diet breadth, with host plants spanning several families and many species. It is unclear, however, whether this phylogenetic generalism is supported by a generic metabolic use of common host chemical compounds (‘metabolic generalism’) or alternatively by distinct uses of diet-specific compounds (‘multi-host metabolic specialism’)? Here, we simultaneously investigated the metabolomes of fruit diets and of individuals of a generalist phytophagous species, Drosophila suzukii , that developed on them. The direct comparison of metabolomes of diets and consumers enabled us to disentangle the metabolic fate of common and rarer dietary compounds. We showed that the consumption of biochemically dissimilar diets resulted in a canalized, generic response from generalist individuals, consistent with the metabolic generalism hypothesis. We also showed that many diet-specific metabolites, such as those related to the particular color, odor, or taste of diets, were not metabolized, and rather accumulated in consumer individuals, even when probably detrimental to fitness. As a result, while individuals were mostly similar across diets, the detection of their particular diet was straightforward. Our study thus supports the view that dietary generalism may emerge from a passive, opportunistic use of various resources, contrary to more widespread views of an active role of adaptation in this process. Such a passive stance towards dietary chemicals, probably costly in the short term, might favor the later evolution of new diet specializations.
Article activity feed
-

-

Author Response
Reviewer #1 (Public Review):
While the mechanism about arm-races between plant and specialist herbivores has been studied, such as detoxification of specific secondary metabolites, the mechanism of the wider diet breadth, so-called generalist herbivores have been less studied. Since the heterogeneity of host plant species, the experimental validation of phylogenetic generalism of herbivores seemed as hard to be conducted. The authors declared the two major hypotheses about the large diet breadth ("metabolic generalism" and "multi-host metabolic specialism"), and carefully designed the experiment using Drosophila suzukii as a model herbivore species.
By an untargeted metabolomics approach using UHPLC-MS, authors attempted to falsify the hypotheses both in qualitative- and quantitative metabolomic profiles. …
Author Response
Reviewer #1 (Public Review):
While the mechanism about arm-races between plant and specialist herbivores has been studied, such as detoxification of specific secondary metabolites, the mechanism of the wider diet breadth, so-called generalist herbivores have been less studied. Since the heterogeneity of host plant species, the experimental validation of phylogenetic generalism of herbivores seemed as hard to be conducted. The authors declared the two major hypotheses about the large diet breadth ("metabolic generalism" and "multi-host metabolic specialism"), and carefully designed the experiment using Drosophila suzukii as a model herbivore species.
By an untargeted metabolomics approach using UHPLC-MS, authors attempted to falsify the hypotheses both in qualitative- and quantitative metabolomic profiles. Intersections of four fruit (puree) samples and each diet-based fly individual samples from the qualitative data revealed that there were few ions that occur as the specific metabolite in each diet-based fly group, which could reject the "multi-host metabolic specialism" hypothesis. Quantitative data also showed results that could support the "metabolic generalism" hypothesis. Therefore, the wide diet breadth of D. suzukii seemed to be derived from the general metabolism rather than the adaptive traits of the diverse host plant species. On the other hand, the reduction of the metabolites (ions) set using GLM seemed logical and 2-D clustering from the reduced ions set showed that quantitative aspects of diet-associated ions could classify "what the flies ate". These interesting results could enhance the understanding of the diet breadth (niche) of herbivorous insects.
The authors' approach seemed clear to falsify the hypotheses based on the appropriate data processing. The intersection of shared ions from the qualitative dataset could distinguish the diet-specific metabolites in flies and commonly occurring metabolites among flies and/or fruits. Also, filtering on the diet-specific ions seemed to be a logical and appropriate way. Meanwhile, the discussion about the results seemed to be focused on different points regarding the research hypotheses which were raised in the introduction part. Discussion about the results mainly focused on the metabolism of D. suzukii itself, rather than the research hypotheses and questions that were raised from the evolution of the wide diet breadth of generalist herbivores. In particular, the conclusion seems to be far from the main context of the authors' research; e.g. frugivory. It makes the implication of the study weaker.
We wish to thank Reviewer #1 for their appreciation of our study. As recommended, we now focus our discussion more on the general aspect of our findings (relevant to insects, herbivores, or frugivores), and less on the peculiarities of the metabolism of D. suzukii itself. Specifically, we now only mention D. suzukii in one section (two sentences) of our Discussion, to serve as an example (l.387-396). Thanks to this comment, the Discussion may interest a broader readership, on the evolution of diet breadth in generalist herbivorous species and offers a better understanding of the general implications of our findings.
Reviewer #2 (Public Review):
The manuscript: "Metabolic consequences of various fruit-based diets in a generalist insect species" by Olazcuaga et al., addresses an interesting question. Using an untargeted metabolomics approach, the authors study how diet generalism may have evolved versus diet specialization which is generally more commonly observed, at least in drosophila species. Using the phytophagous species Drosophila suzukii, and by directly comparing the metabolomes of fruit purees and the flies that fed on them, the authors found evidence for "metabolic generalism". Metabolic generalism means that individuals of a generalist species process all types of diet in a similar way, which is in contrast to "multi-host metabolic specialism" which entails the use of specific pathways to metabolize unique compounds of different diets. The authors find strong evidence for the first hypothesis, as they could easily detect the signature of each fruit diet in the flies. The authors then go on to speculate on the evolutionary ramifications of this for how potentially diet specializations may have evolved from diet generalism. Overall, the paper is well written, the experiments well documented, and the conclusions convincing.
We thank Reviewer #2 for their comments and appreciation of our work.
Reviewer #3 (Public Review):
Laure Olazcuaga et al. investigated the metabolomes of four fruit-based diets and corresponding individuals of Drosophila suzukii that reared on them using comparative metabolomics analysis. They observed that the four fruit-based diets are metabolically dissimilar. On the contrary, flies that fed on them are mostly similar in their metabolic response. From a quantitative point of view, they find that part of the fly metabolomes correlates well with that of the corresponding diet metabolomes, which is indicative of insect ingestive history. By further focusing on 71 metabolites derived from diet-specific fly ions and highly abundant fruit ions, the authors show that D. suzukii differentially accumulates diet metabolism in a compound-specific manner. The authors claim that the data support the metabolic generalism hypothesis while rejecting the multi-host metabolic specialism hypothesis. This study provides a valuable global chemical comparison of how diverse diet metabolites are processed by a generalist insect species.
Strengths:
The rapid advances in high-resolution mass spectrometry have recently accelerated the discovery of many novel post-ingestive compounds through comparative metabolomics analysis of insect/frass and plant samples. Untargeted metabolomics is thus a very powerful approach for the systematic comparison of global chemical shifts when diverse plant-derived specialized metabolites are further modified or quantitatively metabolized after ingestion by insects. The technique can be readily extended to a larger micro- or macro-evolutionary context for both generalist and specialist insects to systematically investigate how plant chemical diversity contributes to dietary generalism and specialism.
We would like to thank Reviewer #3 for their insightful comments on the power of untargeted metabolomics to evaluate the fate of plant metabolites and their use by herbivores. We also agree that these techniques can be used to tackle eco-evolutionary issues, such as the origin and maintenance of dietary generalism and specialism here. We hope that our study will inspire other researchers to explore such techniques and experiments to gain a global overview of biochemistry fluxes and their evolution. We now mention it in the conclusion (L454-459).
Weaknesses:
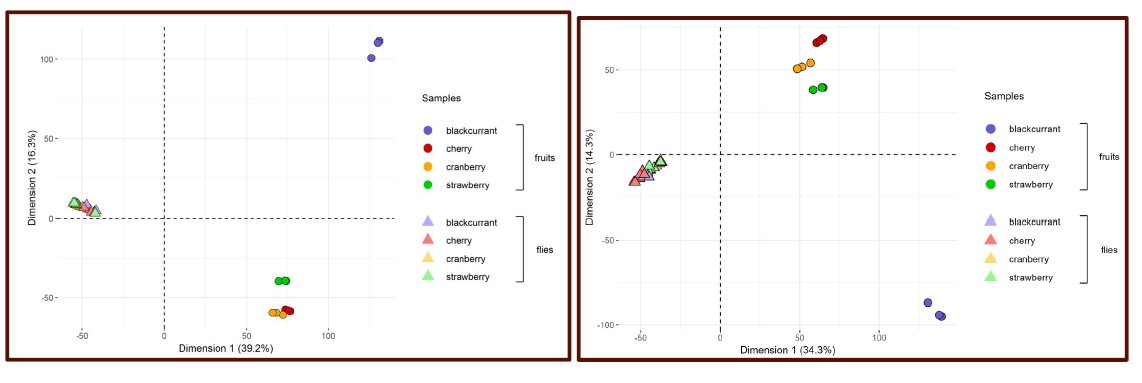
The authors claim that their data support the hypothesis of metabolic generalism, however, a total analysis of insect metabolism may not generate a clean dataset for direct comparison of fruit-derived metabolites with those metabolized by D. suzukii, given that much of these metabolites would be "diluted" proportionally by insect-derived metabolites. If the insect-derived metabolites predominate, then, as the authors observed, a tight clustering of D. suzukii metabolomes in the PCA plot would be expected. It is therefore very difficult to interpret these patterns.
We agree with Reviewer #3 that a careful examination of the different possible origins of metabolites should take place to distinguish between our two competing hypotheses.
The only source of metabolites for insects in our experimental setup is a mixture of (i) a large proportion of fruit purees and (ii) a minor proportion of artificial medium consisting mainly of yeast. Our goal is thus to understand the fate of (i) “fruit-derived” metabolites (transformed and untransformed), while controlling for (ii) “artificial media-derived” metabolites, that constitute a nuisance signal but are necessary for a complete development in our system.
By “fruit-derived” and “insect-derived” metabolites, it is our understanding that Reviewer #3 means “fruit” metabolites (when in insects, untransformed “fruit-derived” metabolites) and “artificial medium-derived” metabolites. It is true that we do wish to avoid a predominance of “artificial medium-derived” metabolites and focus on “fruit-derived” metabolites in insects. We also want to note that it is of primary importance in our study to distinguish between “fruit” metabolites that are carried as is (“fruit” metabolites present in insects, ie untransformed “fruit-derived” metabolites), and “fruit” metabolites that are used after transformation by the insect (i.e., transformed “fruit-derived” metabolites).
We agree with Reviewer #3 that the presence of “artificial medium-derived” metabolites could be problematic in direct comparisons of fruits and insects (and not among fruits or among insects’ comparisons).
However, we took some steps to avoid such problems:
We included control fly samples in our experiment: at each experimental generation, flies developed only on artificial medium (without fruit puree) were collected and processed simultaneously with flies that developed on fruit media. Results using these artificial medium-reared flies as controls (by subtracting their ions levels and removing ions that were similar, respective of their generation) were similar to results using raw data and conclusions were identical (see below).
We lowered the proportion of artificial medium in our fruit media so that it was kept to a minimum, compatible with larval development and adult survival.
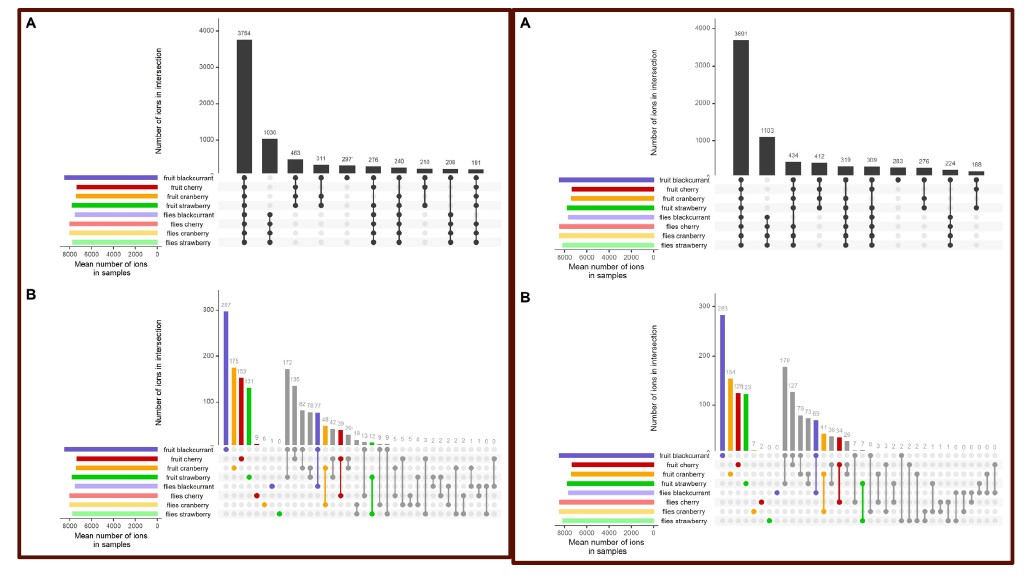
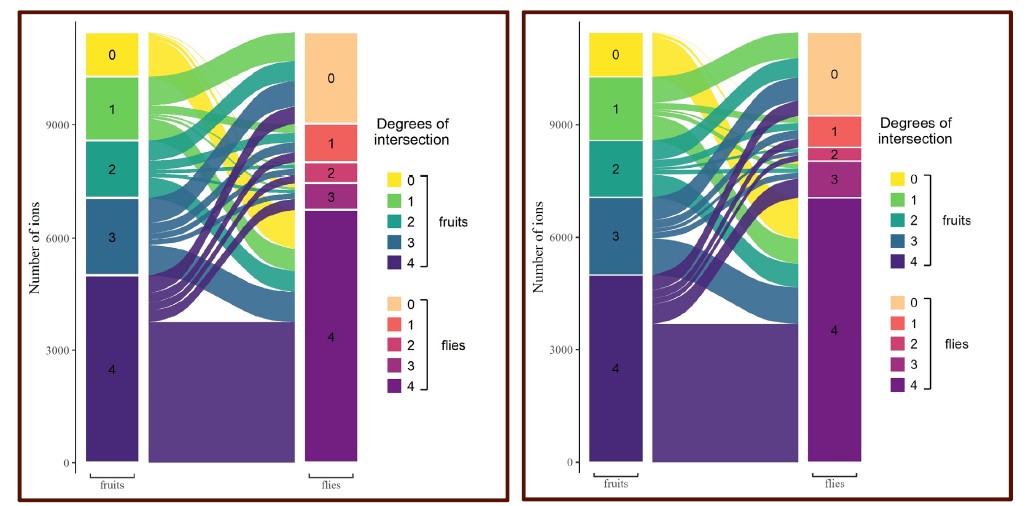
Consistent with the low impact of this “artificial medium” component on our conclusions, we also wish to point out the presence pattern of metabolites found only in flies and never in fruits when using raw data (Figure 3, yellow stack). Even in the most conservative hypothesis of 100% of these metabolites originating from our artificial medium (which is probably not the case), we observe that it constitutes only a minor proportion of metabolites common to all flies (15.7%).
For your consideration, we include below the main Figures, using both raw data and artificial medium-controlled:
Figure 2, left = raw data; right = artificial-media controlled:
Figure 3, left = raw data; right = artificial-media controlled:
Figure 3S1, left = raw data; right = artificial-media controlled:
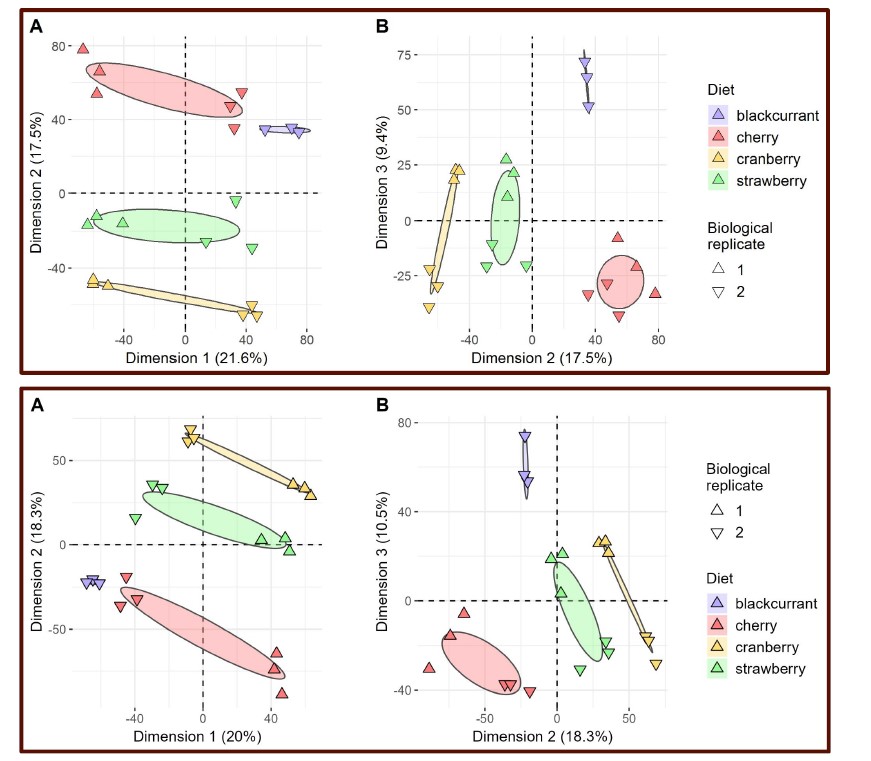
Figure 4, above = raw data; below = artificial-media controlled:
We hope that we convinced the Editor/Reviewers that raw data and artificial-medium controlled data provide a single and same answer to all our analyses. We chose to present only raw data, to simplify the Materials & Methods section.
We however modified the current version of the manuscript to inform the reader that proper controls were done and that their inclusion do not modify any of our conclusions (l.110-113 and l.583-589).
We also wish to point out two additional comments:
As Reviewer #1 also recommended, we modified the expectations drawn in Fig1G to better consider the general comment of “insect derived” metabolites being fundamentally different from plant metabolites (even if we do show in our study that only approx. 9% of metabolites are private to flies).
The main part of our care in the use of this global PCA analysis is that it follows two other analyses (global intersection and comparison of intersections among fruits and among flies) and precedes another one (fly-focused PCA). We hope that all these analyses help the readers get a comprehensive overview of the dataset and associated results, avoiding reliance on a single analysis.
We also help readers to explore and visualize all analyses presented in our manuscript by setting up a shiny application (in addition to our available dataset and R code), at https://fruitfliesmetabo.shinyapps.io/shiny/. This is now mentioned in the main text (l.588-589).
We thank the Reviewer for their comment that greatly improved the manuscript.
The authors generated a qualitative dataset using the peak list produced by XCMS which contains quantitative peak areas, it is unclear how the threshold was selected to determine if a peak is present or absent in a given sample. The qualitative dataset would influence the output of their data analysis.
The referee is right in pointing out that the threshold used to determine if a peak is present or absent in a given sample was not clearly specified. This has now been corrected in the “Host use” section of the Materials & Methods (l.513-516). Briefly, a given replicate of a compound was considered present if the corresponding peak area following XCMS quantification was > 1000. This threshold was selected to be close to the practical quantification threshold of the Thermo Exactive mass spectrometer used in this study. This threshold was selected in order to allow the quantification of low-abundance compounds, as many plant-derived diet compounds were expected to be present in trace amounts in flies. We additionally applied a stringent rule for presence of any given compound (presence in at least 3 biological replicates).
The authors reply on in-source fragmentation for peak annotation when authentic standards are not available. The accuracy of the annotation thus requires further validation.
The Supplementary Table 1 was unfortunately omitted in the first submission of the manuscript. This oversight has been now corrected and the Supplementary Table 1 details all information used for metabolite annotation. In particular, MS/MS data comparison with mass spectral databases as well as with published literature have been added to substantiate metabolite identifications. This MS/MS data was produced thanks to the comment of the Reviewer. We also provide four more annotations from standards to attain 30 / 71 identifications validated through chemical standards.
-

eLife assessment
This useful study uses untargeted metabolomics to help us understand how some herbivores are able to be generalists, rather than specializing in the metabolism of specific plant species. This is an important area, since little is known about how generalist insect species metabolize their food. In its current form, the study lacks ecological relevance due to the exclusive use of refined sampling procedures, and the metabolomic analysis is incomplete.
-

Reviewer #1 (Public Review):
While the mechanism about arm-races between plant and specialist herbivores has been studied, such as detoxification of specific secondary metabolites, the mechanism of the wider diet breadth, so-called generalist herbivores have been less studied. Since the heterogeneity of host plant species, the experimental validation of phylogenetic generalism of herbivores seemed as hard to be conducted. The authors declared the two major hypotheses about the large diet breadth ("metabolic generalism" and "multi-host metabolic specialism"), and carefully designed the experiment using Drosophila suzukii as a model herbivore species.
By an untargeted metabolomics approach using UHPLC-MS, authors attempted to falsify the hypotheses both in qualitative- and quantitative metabolomic profiles. Intersections of four fruit …
Reviewer #1 (Public Review):
While the mechanism about arm-races between plant and specialist herbivores has been studied, such as detoxification of specific secondary metabolites, the mechanism of the wider diet breadth, so-called generalist herbivores have been less studied. Since the heterogeneity of host plant species, the experimental validation of phylogenetic generalism of herbivores seemed as hard to be conducted. The authors declared the two major hypotheses about the large diet breadth ("metabolic generalism" and "multi-host metabolic specialism"), and carefully designed the experiment using Drosophila suzukii as a model herbivore species.
By an untargeted metabolomics approach using UHPLC-MS, authors attempted to falsify the hypotheses both in qualitative- and quantitative metabolomic profiles. Intersections of four fruit (puree) samples and each diet-based fly individual samples from the qualitative data revealed that there were few ions that occur as the specific metabolite in each diet-based fly group, which could reject the "multi-host metabolic specialism" hypothesis. Quantitative data also showed results that could support the "metabolic generalism" hypothesis. Therefore, the wide diet breadth of D. suzukii seemed to be derived from the general metabolism rather than the adaptive traits of the diverse host plant species. On the other hand, the reduction of the metabolites (ions) set using GLM seemed logical and 2-D clustering from the reduced ions set showed that quantitative aspects of diet-associated ions could classify "what the flies ate". These interesting results could enhance the understanding of the diet breadth (niche) of herbivorous insects.
The authors' approach seemed clear to falsify the hypotheses based on the appropriate data processing. The intersection of shared ions from the qualitative dataset could distinguish the diet-specific metabolites in flies and commonly occurring metabolites among flies and/or fruits. Also, filtering on the diet-specific ions seemed to be a logical and appropriate way. Meanwhile, the discussion about the results seemed to be focused on different points regarding the research hypotheses which were raised in the introduction part. Discussion about the results mainly focused on the metabolism of D. suzukii itself, rather than the research hypotheses and questions that were raised from the evolution of the wide diet breadth of generalist herbivores. In particular, the conclusion seems to be far from the main context of the authors' research; e.g. frugivory. It makes the implication of the study weaker.
-

Reviewer #2 (Public Review):
The manuscript: "Metabolic consequences of various fruit-based diets in a generalist insect species" by Olazcuaga et al., addresses an interesting question. Using an untargeted metabolomics approach, the authors study how diet generalism may have evolved versus diet specialization which is generally more commonly observed, at least in drosophila species. Using the phytophagous species Drosophila suzukii, and by directly comparing the metabolomes of fruit purees and the flies that fed on them, the authors found evidence for "metabolic generalism". Metabolic generalism means that individuals of a generalist species process all types of diet in a similar way, which is in contrast to "multi-host metabolic specialism" which entails the use of specific pathways to metabolize unique compounds of different diets. …
Reviewer #2 (Public Review):
The manuscript: "Metabolic consequences of various fruit-based diets in a generalist insect species" by Olazcuaga et al., addresses an interesting question. Using an untargeted metabolomics approach, the authors study how diet generalism may have evolved versus diet specialization which is generally more commonly observed, at least in drosophila species. Using the phytophagous species Drosophila suzukii, and by directly comparing the metabolomes of fruit purees and the flies that fed on them, the authors found evidence for "metabolic generalism". Metabolic generalism means that individuals of a generalist species process all types of diet in a similar way, which is in contrast to "multi-host metabolic specialism" which entails the use of specific pathways to metabolize unique compounds of different diets. The authors find strong evidence for the first hypothesis, as they could easily detect the signature of each fruit diet in the flies. The authors then go on to speculate on the evolutionary ramifications of this for how potentially diet specializations may have evolved from diet generalism. Overall, the paper is well written, the experiments well documented, and the conclusions convincing.
-

Reviewer #3 (Public Review):
Laure Olazcuaga et al. investigated the metabolomes of four fruit-based diets and corresponding individuals of Drosophila suzukii that reared on them using comparative metabolomics analysis. They observed that the four fruit-based diets are metabolically dissimilar. On the contrary, flies that fed on them are mostly similar in their metabolic response. From a quantitative point of view, they find that part of the fly metabolomes correlates well with that of the corresponding diet metabolomes, which is indicative of insect ingestive history. By further focusing on 71 metabolites derived from diet-specific fly ions and highly abundant fruit ions, the authors show that D. suzukii differentially accumulates diet metabolism in a compound-specific manner. The authors claim that the data support the metabolic …
Reviewer #3 (Public Review):
Laure Olazcuaga et al. investigated the metabolomes of four fruit-based diets and corresponding individuals of Drosophila suzukii that reared on them using comparative metabolomics analysis. They observed that the four fruit-based diets are metabolically dissimilar. On the contrary, flies that fed on them are mostly similar in their metabolic response. From a quantitative point of view, they find that part of the fly metabolomes correlates well with that of the corresponding diet metabolomes, which is indicative of insect ingestive history. By further focusing on 71 metabolites derived from diet-specific fly ions and highly abundant fruit ions, the authors show that D. suzukii differentially accumulates diet metabolism in a compound-specific manner. The authors claim that the data support the metabolic generalism hypothesis while rejecting the multi-host metabolic specialism hypothesis. This study provides a valuable global chemical comparison of how diverse diet metabolites are processed by a generalist insect species.
Strengths:
The rapid advances in high-resolution mass spectrometry have recently accelerated the discovery of many novel post-ingestive compounds through comparative metabolomics analysis of insect/frass and plant samples. Untargeted metabolomics is thus a very powerful approach for the systematic comparison of global chemical shifts when diverse plant-derived specialized metabolites are further modified or quantitatively metabolized after ingestion by insects. The technique can be readily extended to a larger micro- or macro-evolutionary context for both generalist and specialist insects to systematically investigate how plant chemical diversity contributes to dietary generalism and specialism.Weaknesses:
The authors claim that their data support the hypothesis of metabolic generalism, however, a total analysis of insect metabolism may not generate a clean dataset for direct comparison of fruit-derived metabolites with those metabolized by D. suzukii, given that much of these metabolites would be "diluted" proportionally by insect-derived metabolites. If the insect-derived metabolites predominate, then, as the authors observed, a tight clustering of D. suzukii metabolomes in the PCA plot would be expected. It is therefore very difficult to interpret these patterns.The authors generated a qualitative dataset using the peak list produced by XCMS which contains quantitative peak areas, it is unclear how the threshold was selected to determine if a peak is present or absent in a given sample. The qualitative dataset would influence the output of their data analysis.
The authors reply on in-source fragmentation for peak annotation when authentic standards are not available. The accuracy of the annotation thus requires further validation.
-