Competitive interactions between culturable bacteria are highly non-additive
Curation statements for this article:-
Curated by eLife
eLife assessment
This important study presents an interesting example of how complexities of communities may be reduced by showing that the joint effects of two or more species on a focal species are generally not additive, but rather dominated by the strongest single effect. The evidence, enabled by over 14,000 measurements using nanodroplet-based microfluidics, is compelling, although the generality of the conclusion awaits further studies. This paper is of interest to microbial ecologists.
This article has been Reviewed by the following groups
Discuss this preprint
Start a discussion What are Sciety discussions?Listed in
- Evaluated articles (eLife)
Abstract
Microorganisms are found in diverse communities whose structure and function are determined by interspecific interactions. Just as single species seldom exist in isolation, communities as a whole are also constantly challenged and affected by external species. Though much work has been done on characterizing how individual species affect each other through pairwise interactions, the joint effects of multiple species on a single (focal) species remain underexplored. As such, it is still unclear how single-species effects combine to a community-level effect on a species of interest. To explore this relationship, we assayed thousands of communities of two, three, and four bacterial species, measuring the effect of single, pairs of, and trios of 61 affecting species on six different focal species. We found that when multiple species each have a negative effect on a focal species, their joint effect is typically not given by the sum of the effects of individual affecting species. Rather, they are dominated by the strongest individual-species effect. Therefore, while joint effects of multiple species are often non-additive, they can still be derived from the effects of individual species, making it plausible to map complex interaction networks based on pairwise measurements. This finding is important for understanding the fate of species introduced into an occupied environment and is relevant for applications in medicine and agriculture, such as probiotics and biocontrol agents, as well as for ecological questions surrounding migrating and invasive species.
Article activity feed
-

-

Author Response
Reviewer #2 (Public Review):
The authors seek to determine how various species combine their effects on the growth of a species of interest when part of the same community.
To this end, the authors carry out an impressive experiment containing what I believe must be one of the largest pairwise + third-order co-culture experiments done to date, using a high-throughput co-culture system they had co-developed in previous work. The unprecedented nature of this data is a major strength of the paper. The authors also discover that species combine their effect through "dominance", i.e. the strongest effect masks the others. This is important as it calls into question the common assumption of additivity that is implicit in the choice of using Lotka-Volterra models.
A stronger claim (i.e. in the abstract) is that joint …
Author Response
Reviewer #2 (Public Review):
The authors seek to determine how various species combine their effects on the growth of a species of interest when part of the same community.
To this end, the authors carry out an impressive experiment containing what I believe must be one of the largest pairwise + third-order co-culture experiments done to date, using a high-throughput co-culture system they had co-developed in previous work. The unprecedented nature of this data is a major strength of the paper. The authors also discover that species combine their effect through "dominance", i.e. the strongest effect masks the others. This is important as it calls into question the common assumption of additivity that is implicit in the choice of using Lotka-Volterra models.
A stronger claim (i.e. in the abstract) is that joint effect of multiple species on the growth of another can be derived from the effect of individual species. Unless I am misunderstanding something, this statement may have to be qualified a little, as the authors show that a model based on pairwise dominance (i.e. the strongest pairwise) does a somewhat better job (lower RMSD, though granted, not by much, 0.57 vs 0.63) than a model based on single species dominance. This is, the effect of the strongest pair predicts better the effect of a trio than the effect of the larger species.
This issue makes one wonder whether, had the authors included higher-order combinations of species (i.e. five-member consortia or higher), the strongest-effect trio would have predicted better than the strongest-effect pair, which in turn is better predictor than the strongest-effect species. This is important, as it would help one determine to what extent the strongest-effect model would work in more diverse communities, such as those one typically finds in nature. Indeed, the authors find that the predictive ability of the strongest effect species is much stronger for pairs than it is for trios (RMSD of 0.28 vs 0.63). Does the predictive ability of the single species model decline faster and faster as diversity grows beyond 4-member consortia?
Thank you for raising this important point. It is true that in our study we see that single species predict pairs better than trios, and that pairs predict trios better than single species. As we did not perform experiments on more diverse communities (n>4), we are not sure if or how these rules will scale up. We explicitly address these caveats in our revised discussion.
Reviewer #3 (Public Review):
A problem in synthetic ecology is that one can't brute-force complex community design because combinatorics make it basically impossible to screen all possible communities from a bank of possible species. Therefore, we need a way to predict phenomena in complex communities from phenomena in simple communities. This paper aims to improve this predictive ability by comparing a few different simple models applied to a large dataset obtained with the use of the author's "kchip" microfluidics device. The main question they ask is whether the effect of two species on a focal species is predicted from the mean, the sum, or the max of the effect of each single "affecting" species on the focal species. They find that the max effect is often the best predictor, in the sense of minimizing the difference between predicted effect and measured effect. They also measure single-species trait data for their library of strains, including resource niche and antibiotic resistance, and then find that Pearson correlations between distance calculations generated from these metrics and the effect of added species are weak and unpredictive. This work is largely well-done, timely and likely to be of high interest to the field, as predicting ecosystem traits from species traits is a major research aim.
My main criticism is that the main take-home from the paper (fig 3B)-that the strongest effect is the best predictor-is oversold. While it is true that, averaged over their six focal species, the "strongest effect" was the best overall predictor, when one looks at the species-specific data (S9), we see that it is not the best predictor for 1/3 of their focal species, and this fraction grows to 1/2 if one considers a difference in nRMSE of 0.01 to be negligible.
As suggested, we have softened our language regarding the take-home message. This matter is addressed in detail above in response to 'Essential Revisions'. Briefly, we see that the strongest model works best when both single species have qualitatively similar effects, but is slightly less accurate when effects are mixed. We also see overall less accurate predictions for positive effects. In light of these findings, we propose that focal species for which the strongest model is not the most accurate is due to the interaction types, and not specific to the focal species.
We made substantial changes to the manuscript, including the first paragraph of the discussion which more accurately describes these findings and emphasizes the relevant caveats:
"By measuring thousands of simplified microbial communities, we quantified the effects of single species, pairs, and trios on multiple focal species. The most accurate model, overall and specifically when both single species effects were negative, was the strongest effect model. This is in stark contrast to models often used in antibiotic compound combinations, despite most effects being negative, where additivity is often the default model (Bollenbach 2015). The additive model performed well for mixed effects (i.e. one negative and one positive), but only slightly better than the strongest model, and poorly when both species had effects of the same sign. When both single species’ effects were positive, the strongest model was also the best, though the difference was less pronounced and all models performed worse for these interactions. This may be due to the small effect size seen with positive effects, as when we limited negative and mixed effects to a similar range of effects strength, their accuracy dropped to similar values (Figure 3–Figure supplement 5). We posit that the difference in accuracy across species is affected mainly by the effect type dominating different focal species' interactions, rather than by inherent species traits (Figure 3–Figure supplement 6)." (Lines 288-304)
The same criticism applies to the result from figure 2-that pairs of affecting species have more negative effects than single species. Considered across all focal species this is true (though minor in effect size, Fig 2A). But there is only a significant effect within two individual species. Again, this points to the effects being focal-species-specific, and perhaps not as generalizable as is currently being claimed.
Upon more rigorous analysis, and with regard to changes in the dataset after filtering, we see that the more accurate statement is that effects become stronger, not necessarily more negative (in line with the accuracy of the strongest model). The overall trend is towards more negative interactions, due to the majority of interactions being negative, but as stated this is not true for each individual focal. As such the following sentence in the manuscript has been changed:
"The median effect on each focal was more negative by 0.28 on average, though the difference was not significant in all cases; additionally, focals with mostly positive single species interactions showed a small increase in median effect (Fig. 2D)" (Lines 151-154)
As well as the title of this section: "Joint effects of species pairs tend to be stronger than those of individual affecting species" (Lines 127-128)
Another thing that points to a focal-species-specific response is Fig 2D, which shows the distributions of responses of each focal species to pairs. Two of these distributions are unimodal, one appears bimodal, and three appear tri-modal. This suggests to me that the focal species respond in categorically different ways to species addition.
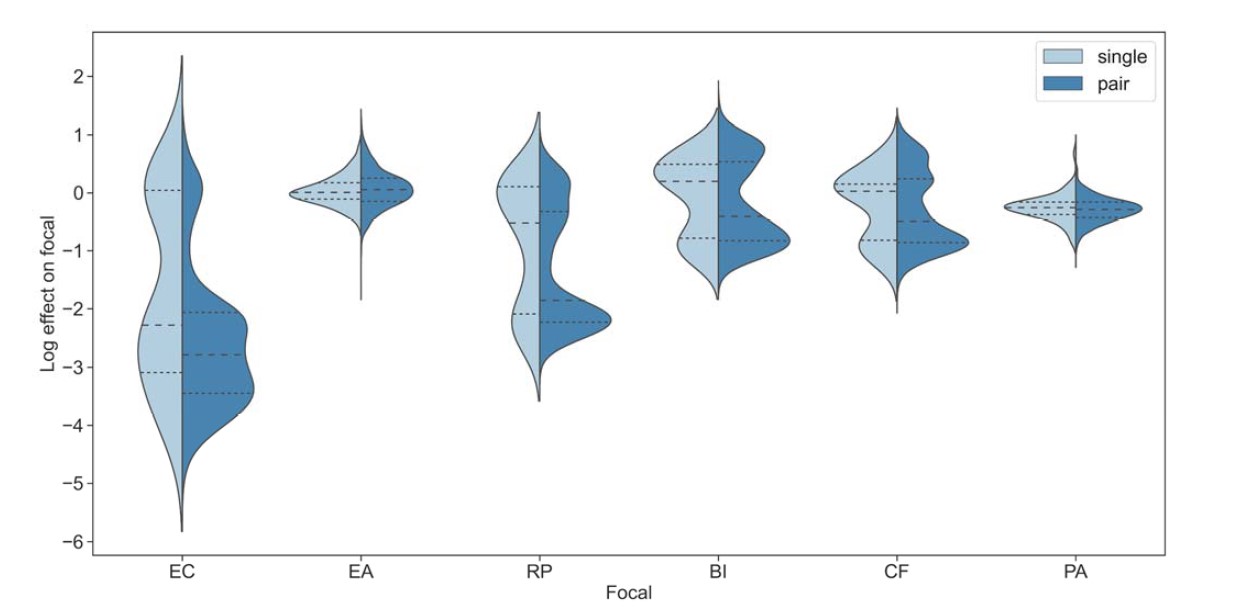
We believe this distribution of pair effects is related to the distribution of single species effects, and not to the way in which different focal species respond to the addition of second species. Though this may be difficult to see from the swarm plots shown in the paper, below is a split violin plot that emphasizes this point.
Fig R1: Distribution of single species and pair effects. Distribution of the effect of single and pairs of affecting species for each focal species individually. Dashed lines represent the median, while dotted lines the interquartile range.
These differences occur even though the focal bacteria are all from the same family. This suggests to me that the generalizability may be even less when a more phylogenetically dispersed set of focal species are used.
We have added the following sentence to the discussion explicitly emphasizing the phylogenetic limitations of our study:
"Lastly, it is important to note that our focal species are all from the same order (Enterobacterales), which may also limit the purview of our findings." (Lines 364-366)
Considering these points together, I argue that the conclusion should be shifted from "strongest effect is the best" to "in 3 of our focal species, strongest effect was the best, but this was not universal, and with only 6 focal species, we can't know if it will always be the best across a set of focal species".
As mentioned above, we have softened our language regarding the take-home message in response to these evaluations.
My second main criticism is that it is hard to understand exactly how the trait data were used to predict effects. It seems like it was just pearson correlation coefficients between interspecies niche distances (or antibiotic distances) and the effect. I'm not very surprised these correlations were unpredictive, because the underlying measurements don't seem to be relevant to the environment tested. What if, rather than using niche data across 20 nutrients, only the growth data on glucose (the carbon source in the experiments) was used? I understand that in a field experiment, for example, one might not know what resources are available, and so measuring niche across 20 resources may be the best thing to do. Here though it seems imperative to test using the most relevant data.
It is true that much of the profiling data is not directly related to the experimental conditions (different carbon sources and antibiotics), but in addition to these we do use measurements from experiments carried out in the same environment as the interactions assays (i.e. growth rate and carrying capacity when growing on glucose), which also showed poor correlation with the effects on focals. Additionally, we believe that these profiles contain relevant information regarding metabolic similarity between species (similar to metabolic models often constructed computationally). To improve clarity, we added the following sentence to the figure legend of Figure 3–Figure supplement 1:
"The growth rate, and maximum OD shown in panel A were measured only in M9 glucose, similar to conditions used in the interaction assays." (Lines 591-592)
Additionally and relatedly, it would be valuable to show the scatterplots leading to the conclusion that trait data were uninformative. Pearson's r only works on an assumption of linearity. But there could be strong relationships between the trait data and effect that are monotonic but not linear, or even that are non-monotonic yet still strong (e.g. U-shaped). For the first case, I recommend switching to Spearman's rho over Pearson's r, because it only assumes monotonicity, not linearity. If there are observable relationships that are not monotonic, a different test should be used.
Per your suggestion, we have changed the measurement of correlation in this analysis from Pearson's r, to Spearman's rho. As we observed similar, and still mostly weak correlations, we did not investigate these relationships further. See Figure 3–Figure supplement 1.
Additionally, we generated heat maps including scatterplots mapping the data leading to these correlations. We found no notable dependency in these plots, and visually they were quite crowded and difficult to interpret. As this is not the central point of our study, we ultimately decided against adding this information to the plots.
In general, I think the analyses using the trait data were too simplistic to conclude that the trait data are not predictive.
We agree that more sophisticated analyses may help connect between species traits and their effects on focal species. In fact, other members of our research group have recently used machine learning to accomplish similar predictions (https://doi.org/10.1101/2022.08.02.502471). As such we have changed the wording in to reflect that this correlation is difficult to find using simple analyses:
"These results indicate that it may be challenging to connect the effects of single and pairs of species on a focal strain to a specific trait of the involved strains, using simple analysis." (Lines 157-159)
-

eLife assessment
This important study presents an interesting example of how complexities of communities may be reduced by showing that the joint effects of two or more species on a focal species are generally not additive, but rather dominated by the strongest single effect. The evidence, enabled by over 14,000 measurements using nanodroplet-based microfluidics, is compelling, although the generality of the conclusion awaits further studies. This paper is of interest to microbial ecologists.
-

Reviewer #1 (Public Review):
The authors used high-throughput nanodroplet-based microfluidics to measure the effect of individual species and the joint effects of species pairs and trios on the growth of six fluorescently labeled strains (E. coli and five soil isolates). In total, over 14,000 bacterial communities composed of subsets of a library of 61 soil and leaf isolates were quantified. They found that the effects of multiple species combine non-additively and are heavily dominated by the strongest single species effect.
Single species showed large variability in the ability to use different carbon sources and survive antibiotics. Interaction assays were carried out in minimal M9 media with 0.5% [w/v] glucose for 24 hours. Authors found that single species' effects on the focal strain is positive in ~1/3 of cases, and is negative …
Reviewer #1 (Public Review):
The authors used high-throughput nanodroplet-based microfluidics to measure the effect of individual species and the joint effects of species pairs and trios on the growth of six fluorescently labeled strains (E. coli and five soil isolates). In total, over 14,000 bacterial communities composed of subsets of a library of 61 soil and leaf isolates were quantified. They found that the effects of multiple species combine non-additively and are heavily dominated by the strongest single species effect.
Single species showed large variability in the ability to use different carbon sources and survive antibiotics. Interaction assays were carried out in minimal M9 media with 0.5% [w/v] glucose for 24 hours. Authors found that single species' effects on the focal strain is positive in ~1/3 of cases, and is negative in 60% of cases. When two species were added, 3/4 of cases became negative. The similarity of phenotypes (e.g. metabolic profile or phylogeny) between the focal and affecting species did not correlate strongly with the effect on focal species. Joint effects of two or more species are not additive, but rather dominated by the strongest single effect.
The paper presents an interesting example of how complexities of communities may be reduced. The assay combines different growth aspects (lag, rate, yield), and these different aspects should ideally be untangled. The mechanism of why joint effects are dominated by the strongest effect is not known. The generality of the findings awaits further studies (e.g. what will happen if environments are changed?) However, these future directions do not diminish the value of this study.
-

Reviewer #2 (Public Review):
The authors seek to determine how various species combine their effects on the growth of a species of interest when part of the same community.
To this end, the authors carry out an impressive experiment containing what I believe must be one of the largest pairwise + third-order co-culture experiments done to date, using a high-throughput co-culture system they had co-developed in previous work. The unprecedented nature of this data is a major strength of the paper. The authors also discover that species combine their effect through "dominance", i.e. the strongest effect masks the others. This is important as it calls into question the common assumption of additivity that is implicit in the choice of using Lotka-Volterra models.
A stronger claim (i.e. in the abstract) is that joint effect of multiple species …
Reviewer #2 (Public Review):
The authors seek to determine how various species combine their effects on the growth of a species of interest when part of the same community.
To this end, the authors carry out an impressive experiment containing what I believe must be one of the largest pairwise + third-order co-culture experiments done to date, using a high-throughput co-culture system they had co-developed in previous work. The unprecedented nature of this data is a major strength of the paper. The authors also discover that species combine their effect through "dominance", i.e. the strongest effect masks the others. This is important as it calls into question the common assumption of additivity that is implicit in the choice of using Lotka-Volterra models.
A stronger claim (i.e. in the abstract) is that joint effect of multiple species on the growth of another can be derived from the effect of individual species. Unless I am misunderstanding something, this statement may have to be qualified a little, as the authors show that a model based on pairwise dominance (i.e. the strongest pairwise) does a somewhat better job (lower RMSD, though granted, not by much, 0.57 vs 0.63) than a model based on single species dominance. This is, the effect of the strongest pair predicts better the effect of a trio than the effect of the larger species.
This issue makes one wonder whether, had the authors included higher-order combinations of species (i.e. five-member consortia or higher), the strongest-effect trio would have predicted better than the strongest-effect pair, which in turn is better predictor than the strongest-effect species. This is important, as it would help one determine to what extent the strongest-effect model would work in more diverse communities, such as those one typically finds in nature. Indeed, the authors find that the predictive ability of the strongest effect species is much stronger for pairs than it is for trios (RMSD of 0.28 vs 0.63). Does the predictive ability of the single species model decline faster and faster as diversity grows beyond 4-member consortia?
While this may limit the scope of the paper somewhat, it does not subtract from its overall impressiveness. This is a very strong paper that combines state of the art methodology, of a kind that will change the field, with an important question and an intriguing finding that will surely motivate further work. Therefore, it will be of wide interest and appeal to the broad audience of microbial ecologists, and it is an important and compelling step forward in the field.
-

Reviewer #3 (Public Review):
A problem in synthetic ecology is that one can't brute-force complex community design because combinatorics make it basically impossible to screen all possible communities from a bank of possible species. Therefore, we need a way to predict phenomena in complex communities from phenomena in simple communities. This paper aims to improve this predictive ability by comparing a few different simple models applied to a large dataset obtained with the use of the author's "kchip" microfluidics device. The main question they ask is whether the effect of two species on a focal species is predicted from the mean, the sum, or the max of the effect of each single "affecting" species on the focal species. They find that the max effect is often the best predictor, in the sense of minimizing the difference between …
Reviewer #3 (Public Review):
A problem in synthetic ecology is that one can't brute-force complex community design because combinatorics make it basically impossible to screen all possible communities from a bank of possible species. Therefore, we need a way to predict phenomena in complex communities from phenomena in simple communities. This paper aims to improve this predictive ability by comparing a few different simple models applied to a large dataset obtained with the use of the author's "kchip" microfluidics device. The main question they ask is whether the effect of two species on a focal species is predicted from the mean, the sum, or the max of the effect of each single "affecting" species on the focal species. They find that the max effect is often the best predictor, in the sense of minimizing the difference between predicted effect and measured effect. They also measure single-species trait data for their library of strains, including resource niche and antibiotic resistance, and then find that Pearson correlations between distance calculations generated from these metrics and the effect of added species are weak and unpredictive. This work is largely well-done, timely and likely to be of high interest to the field, as predicting ecosystem traits from species traits is a major research aim.
My main criticism is that the main take-home from the paper (fig 3B)-that the strongest effect is the best predictor-is oversold. While it is true that, averaged over their six focal species, the "strongest effect" was the best overall predictor, when one looks at the species-specific data (S9), we see that it is not the best predictor for 1/3 of their focal species, and this fraction grows to 1/2 if one considers a difference in nRMSE of 0.01 to be negligible.
The same criticism applies to the result from figure 2-that pairs of affecting species have more negative effects than single species. Considered across all focal species this is true (though minor in effect size, Fig 2A). But there is only a significant effect within two individual species. Again, this points to the effects being focal-species-specific, and perhaps not as generalizable as is currently being claimed.
Another thing that points to a focal-species-specific response is Fig 2D, which shows the distributions of responses of each focal species to pairs. Two of these distributions are unimodal, one appears bimodal, and three appear tri-modal. This suggests to me that the focal species respond in categorically different ways to species addition.
These differences occur even though the focal bacteria are all from the same family. This suggests to me that the generalizability may be even less when a more phylogenetically dispersed set of focal species are used.
Considering these points together, I argue that the conclusion should be shifted from "strongest effect is the best" to "in 3 of our focal species, strongest effect was the best, but this was not universal, and with only 6 focal species, we can't know if it will always be the best across a set of focal species".
My second main criticism is that it is hard to understand exactly how the trait data were used to predict effects. It seems like it was just pearson correlation coefficients between interspecies niche distances (or antibiotic distances) and the effect. I'm not very surprised these correlations were unpredictive, because the underlying measurements don't seem to be relevant to the environment tested. What if, rather than using niche data across 20 nutrients, only the growth data on glucose (the carbon source in the experiments) was used? I understand that in a field experiment, for example, one might not know what resources are available, and so measuring niche across 20 resources may be the best thing to do. Here though it seems imperative to test using the most relevant data.
Additionally and relatedly, it would be valuable to show the scatterplots leading to the conclusion that trait data were uninformative. Pearson's r only works on an assumption of linearity. But there could be strong relationships between the trait data and effect that are monotonic but not linear, or even that are non-monotonic yet still strong (e.g. U-shaped). For the first case, I recommend switching to Spearman's rho over Pearson's r, because it only assumes monotonicity, not linearity. If there are observable relationships that are not monotonic, a different test should be used.
In general, I think the analyses using the trait data were too simplistic to conclude that the trait data are not predictive.
-