T cell receptor convergence is an indicator of antigen-specific T cell response in cancer immunotherapies
Curation statements for this article:-
Curated by eLife
eLife assessment
This manuscript reports an association between TCR convergence and involvement in an antigen-specific response. TCR convergence is assessed as a potential biomarker of response to immune checkpoint blockade (ICB). From jointly analyzing TCR-seq data, single-cell RNA-seq data, and antigen-specific TCR information, the authors provided evidence that convergence is a potential indicator for ongoing T cell antigen-specific response. Overall, the analyses are sound the manuscript is well-written, and the study provides the first evidence that TCRseq alone could be used to predict clinical outcomes.
This article has been Reviewed by the following groups
Discuss this preprint
Start a discussion What are Sciety discussions?Listed in
- Evaluated articles (eLife)
Abstract
T cells are potent at eliminating pathogens and playing a crucial role in the adaptive immune response. T cell receptor (TCR) convergence describes T cells that share identical TCRs with the same amino acid sequences but have different DNA sequences due to codon degeneracy. We conducted a systematic investigation of TCR convergence using single-cell immune profiling and bulk TCRβ-sequence (TCR-seq) data obtained from both mouse and human samples and uncovered a strong link between antigen-specificity and convergence. This association was stronger than T cell expansion, a putative indicator of antigen-specific T cells. By using flow-sorted tetramer + single T cell data, we discovered that convergent T cells were enriched for a neoantigen-specific CD8 + effector phenotype in the tumor microenvironment. Moreover, TCR convergence demonstrated better prediction accuracy for immunotherapy response than the existing TCR repertoire indexes. In conclusion, convergent T cells are likely to be antigen-specific and might be a novel prognostic biomarker for anti-cancer immunotherapy.
Article activity feed
-

-

Author Response
Reviewer #3 (Public Review):
Main results:
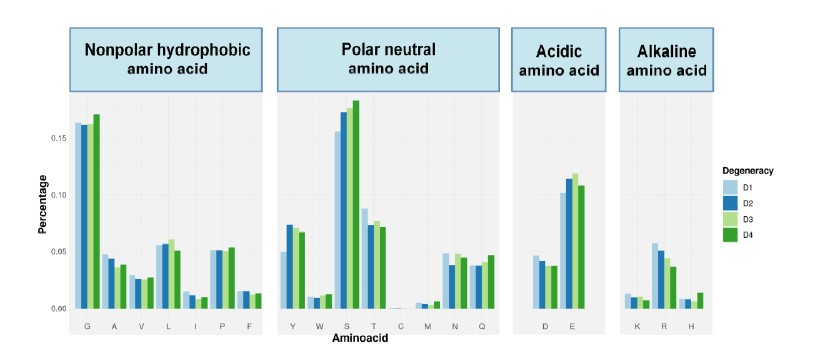
- TCR convergence is different from publicity: The authors look at CDR3 sequence features of convergent TCRs in the large Emerson CMV cohort. Amino usage does not perfectly correlate with codon degeneracy, for example, arginine (which has 6 codons) is less common in convergent TCRs, whereas leucine and serine are elevated. It's argued that there's more to convergence than just recombination biases, which makes sense. (I wonder if the trends for charged amino acids could be explained by the enrichment of convergent TCRs in CD8 T cells, which tend to have more acidic CDR3 loops). There's also a claim that the overlap between convergent and public TCRs is lower in tumors with a high mutational burden (TMB), but this part is sketchy: the definition of public TCRs is murky and …
Author Response
Reviewer #3 (Public Review):
Main results:
- TCR convergence is different from publicity: The authors look at CDR3 sequence features of convergent TCRs in the large Emerson CMV cohort. Amino usage does not perfectly correlate with codon degeneracy, for example, arginine (which has 6 codons) is less common in convergent TCRs, whereas leucine and serine are elevated. It's argued that there's more to convergence than just recombination biases, which makes sense. (I wonder if the trends for charged amino acids could be explained by the enrichment of convergent TCRs in CD8 T cells, which tend to have more acidic CDR3 loops). There's also a claim that the overlap between convergent and public TCRs is lower in tumors with a high mutational burden (TMB), but this part is sketchy: the definition of public TCRs is murky and hard to interpret, and the correlation between TMB and convergence-publicity overlap is modest (two cohorts with low TMB have higher overlap, and the other three have lower, but there is no association over those three, if anything the trend is in the other direction). It's also not clear why the overlap between COVID19 cohort convergent TCRs and public TCRs defined by the pre-2019 Emerson cohort should be high. A confounder here is the potential association between convergence and clonal expansion since expanded clonotypes can spawn apparently convergent TCRs due to sequencing errors. The paper "TCR Convergence in Individuals Treated With Immune Checkpoint Inhibition for Cancer" (Ref#5 here) gives evidence that sequencing errors may be inflating convergence in this specific dataset.
We really appreciate the reviewer’s feedback. We respond to each of the reviewer’s points below:
(1) Amino acid preference of convergent TCRs might be caused by CD8+ T cell enrichment. To test this hypothesis, we performed the same analysis using only CD8+ T cells (using the Cader 2019 lymphoma cohort). The results are shown below. We do not observe significant changes after excluding CD4+ T cells, indicating that this enrichment might be caused by factors other than CD4/CD8 differences.
(2) Definition of public TCRs. We have changed the definition of public TCRs. Instead of mixing the Emerson cohort into each group and using the mixed cohort to define the public TCRs, we just used the 666 samples of the Emerson cohort to define the same set of public TCRs and applied them to each cohort. Both the dataset and the approach used in this manuscript is consistent with a previous study on the same topic (Madi et al., 2014, elife).
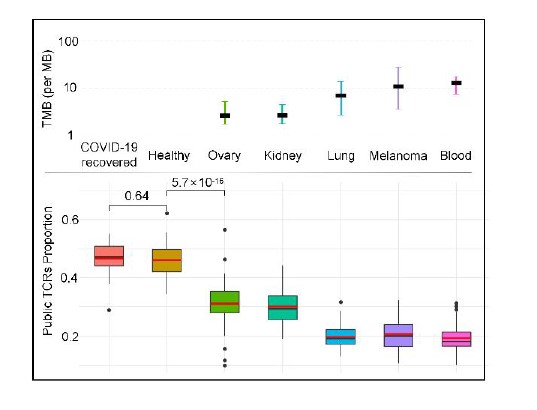
(3) Convergence-publicity overlap: We agree with the reviewer that some high TMB tumors did not show further decrease of convergence-publicity overlap. One potential explanation is that the correlation between the two is not linear. By adding additional cohorts in this revision (healthy and recovered COVID-19 patients), we confirmed the previously observed overall trend between TMB and the overlap, which supported our conclusions (see figure below). On the other hand, we believe that the high overlap of convergent TCRs among healthy cohorts might result from exposure to common antigens. In the cancer patients, while still exposed, private antigens derived from tumor cells are expected to compete for resources, thus reducing the proportion of these public TCRs in the blood repertoire. The above discussion has been added to the revised manuscript:
“Healthy individuals are expected to be exposed to common pathogens, which might induce public T cell responses. On the other hand, cancer patients have more neoantigens due to the accumulative mutation, which drives their antigen-specific T cells to recognize these 'private' antigens. This reduces the proportion of public TCRs in antigen-specific TCRs. Furthermore, a higher tumor mutation burden (TMB) would indicate a higher abundance of neoantigens, resulting in a lower ratio of public TCRs.”
- Convergent TCRs are more likely to be antigen-specific: This is nicely shown on two datasets: the large dextramer dataset from 10x genomics, and the COVID19 datasets from Adaptive biotech. But given previous work on TCR convergence, for example, the Pogorelyy ALICE paper, and many others, this is also not super-surprising.
We thank the reviewer for bringing up this related work. In the Pogorelyy ALICE paper, the authors defined TCR neighbors based on one nucleotide difference of a given CDR3, which included both synonymous and non-synonymous changes. In other words, ALICE combines both convergence and mismatched (with hamming distance 1) sequences as neighbors. Although highly relevant, our approach is different by focusing only on the convergence, as mismatch has been extensively investigated by previous studies. We have now added this paper as Ref 27, and discussed the difference between ALICE and our method in the revised manuscript.
- Convergent T cells exhibit a CD8+ cytotoxic gene signature: This is based on a nice analysis of mouse and human single-cell datasets. One striking finding is that convergent TCRs are WAY more common in CD8+ T cells than in CD4+ T cells. It would be interesting to know how much of this could be explained by greater clonal expansion of CD8+ T cells, together with sequencing errors. A subtle point here is that some of the P values are probably inflated by the presence of expanded clonotypes: a group of cells belonging to the same expanded clonotype will tend to have similar gene expression (and therefore similar cluster membership), and will necessarily all be either convergent or not convergent collectively since they share the same TCR. So it's probably not quite right to treat them as independent for the purposes of assessing associations between gene expression clusters and convergence (or any other TCR-defined feature). You can see evidence for clonal expansion in Figure 3C, where TRAV genes are among the most enriched, suggesting that Cluster 04 may contain expanded clones.
(1) We agree with the reviewer that a possible explanation of the CD8/CD4 difference is the larger cell expansion of CD8+ T cells. We tested this hypothesis by counting the number of T cell clones instead of cell number to remove the effect that would have been caused by CD8 T cell expansion. We first investigated the bulk TCR repertoire sequencing samples as Figure 3 - figure supplement 2C-2D (see figure below). We observed higher convergence levels for the CD8+ T cell clones compared to CD4+ T cells. The additional description of this topic was added at the last paragraph of the result section of “Convergent T cells exhibit a CD8+ cytotoxic gene signature” as follows:
“The results may be explained by larger cell expansions of CD8+ T cells than CD4+ T cells. Therefore, we calculated the number of convergent clones within CD8+ T cells and CD4+ T cells from the above datasets to exclude the effects of cell expansion. As a result, in the scRNA-seq mouse data, while only 1.54% of the CD4+ clones were convergent, 3.76% of the CD8+ clones showed convergence. Likewise, 0.17% of convergent CD4+ T cell clones and 1.03% of convergent CD8+ T cell clones were found in human scRNA-seq data. In the bulk TCR-seq lymphoma data, similar results were also observed, where the gap between the convergent levels of CD4+ and CD8+ T cells narrowed but remained significant (Figure 3—figure supplement 2C-2D). In conclusion, these results suggest that CD8+ T cells show higher levels of convergence than CD4+ T cells, which substantiated our hypothesis that convergent T cells are more likely antigen-experienced. This observation has been tested using multiple datasets with diverse sequencing platforms and sequencing depth to minimize the impact of batch or other technical artifacts.”
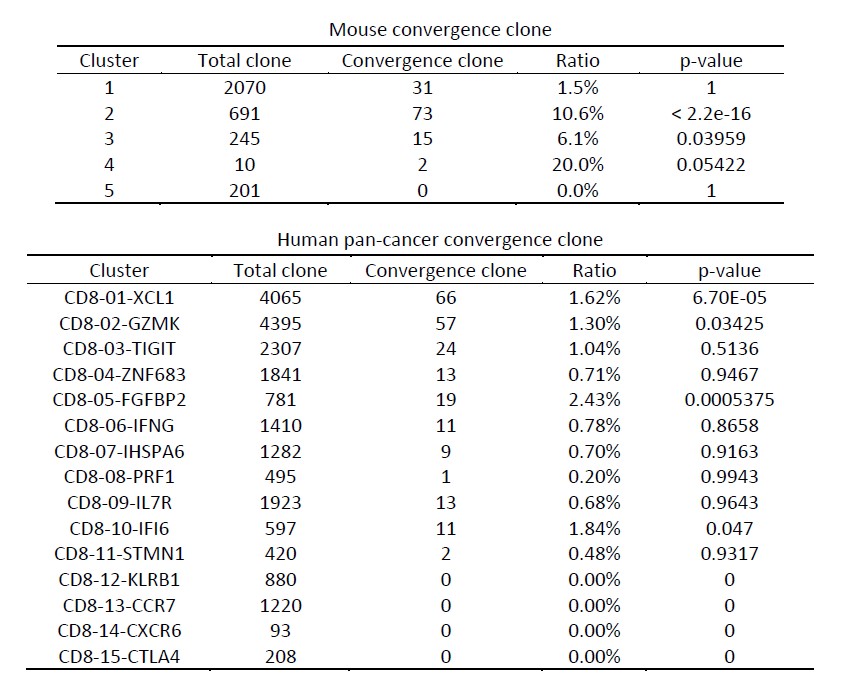
(2) We next investigated the effect of cell expansion in the single cell analysis. We agree with the reviewer that some highly-expanded convergent clones could inflate the p-value. Therefore, we revised the calculation of TCR convergence by using the T cell clone instead of individual cells. We observed that the clusters of interest mentioned in the paper (for both mouse and human data) remain at the top convergent level among all clusters (see table below), with p values estimated using Binomial exact test. These results supported our hypothesis that TCR convergence is enriched for T cell clusters that are more likely antigen-experienced.
- TCR convergence is associated with the clinical outcome of ICB treatment: The associations for the first analysis are described as significant in the text, and they are, but just barely (0.045 and 0.047, but you have to check the figure to see that).
As suggested by the reviewer, we have added the p-value to the test so that it is easier to see. In this revision, we adopted another definition of convergent level, changing from the ratio of convergent TCR to the actual number of convergent T cell clones within each sample. The p-values were more significant using this new indicator (0.02 and 0.00038). To avoid the effect of other variables that might be correlative with convergent levels, especially the sequencing depth, the multivariate Cox model was used for both datasets tested in the paper, correcting for TCR clonality, TCR diversity and sequencing depth (and different treatment methods for melanomas data). As a result, convergence remains significantly prognostic after adjusting for the additional variables.
- Introduction/Discussion: Overall, the authors could do a better job citing previous work on convergence, for example, papers from Venturi on convergent recombination and the work from Mora and Walczak (ALICE, another recombination modeling). They also present the use of convergence as an ICB biomarker as a novel finding, but Ref 5 introduces this concept and validates it in another cohort. Ref 5 also has a careful analysis of the link between sequencing errors and convergence, which could have been more carefully considered here.
We thank the reviewer for this excellent suggestion. We have added the citation of Venturi on convergent recombination as Ref 43 and we cited it at the last paragraph of the result selection:
“Convergent recombination was claimed to be the mechanistic basis for public TCR response in many previous studies(Quigley et al., 2010; Venturi et al., 2006).”
We also included work from Mora and Walczak in the fourth paragraph of the introduction and the third paragraph of the discussion as Ref 27 to introduce this TCR similarity-based clustering method as well as its application in predicting ICB response:
“This idea has led several TCR similarity-based clustering algorithms, such as ALICE (Pogorelyy et al., 2019), TCRdist (Dash et al., 2017), GLIPH2 (Huang et al., 2020), iSMART (Zhang et al., 2020), and GIANA (Zhang et al., 2021), to be developed for studying antigen-driven T cell expansion during viral infection or tumorigenesis.”
“In addition, the potential prognostic value of TCR convergence and TCR similarity-based clustering was testified in other studies(Looney et al., 2019; Pogorelyy et al., 2019).”
Ref 5 was recited while discussing the effect of sequencing error on TCR convergence in the fourth paragraph of discussion:
“Improper handling of sequencing errors may result in the overestimation of TCR convergence (Looney et al., 2019).”
-

eLife assessment
This manuscript reports an association between TCR convergence and involvement in an antigen-specific response. TCR convergence is assessed as a potential biomarker of response to immune checkpoint blockade (ICB). From jointly analyzing TCR-seq data, single-cell RNA-seq data, and antigen-specific TCR information, the authors provided evidence that convergence is a potential indicator for ongoing T cell antigen-specific response. Overall, the analyses are sound the manuscript is well-written, and the study provides the first evidence that TCRseq alone could be used to predict clinical outcomes.
-

Reviewer #1 (Public Review):
This is a very interesting manuscript by Pan et al. The authors focused their analysis on TCR degeneracy and systematically characterized it in single TCR antigen sequencing data, bulk TCR sequencing data, and single-cell RNA sequencing data and obtained unexpected results. Later, authors also used TCR degeneracy as a metric to predict the immunotherapy outcomes. The analyses are sound, and the manuscript is well written.
-

Reviewer #2 (Public Review):
As the first comprehensive integrative analysis on TCR convergence, this study provided several interesting insights:
- Convergence might be induced by an ongoing immune response against viral infection or tumor; 2) in the tumor, there is a positive association between TCR convergence and tumor mutation load, and neoantigen-specific T cells are enriched for convergent TCRs, both observations further supporting the tumor-reactive hypothesis; 3) a potentially new diagnostic predictor for ICB treatment. Given these strengths, this work is of general interest to a broad audience.
-

Reviewer #3 (Public Review):
This manuscript explores the concept of TCR convergence, defined here as the presence of TCRs with the same amino acid sequence but distinct nucleotide sequences. The central premise is that TCR convergence is a sign of antigen-driven selection. TCR convergence as a biomarker for immune checkpoint blockade (ICB) response is also investigated. Although both these ideas have been put forward in the literature, this manuscript provides some new analyses and a new perspective on these topics.
Main results:
- TCR convergence is different from publicity: The authors look at CDR3 sequence features of convergent TCRs in the large Emerson CMV cohort. Amino usage does not perfectly correlate with codon degeneracy, for example, arginine (which has 6 codons) is less common in convergent TCRs, whereas leucine and serine …
Reviewer #3 (Public Review):
This manuscript explores the concept of TCR convergence, defined here as the presence of TCRs with the same amino acid sequence but distinct nucleotide sequences. The central premise is that TCR convergence is a sign of antigen-driven selection. TCR convergence as a biomarker for immune checkpoint blockade (ICB) response is also investigated. Although both these ideas have been put forward in the literature, this manuscript provides some new analyses and a new perspective on these topics.
Main results:
- TCR convergence is different from publicity: The authors look at CDR3 sequence features of convergent TCRs in the large Emerson CMV cohort. Amino usage does not perfectly correlate with codon degeneracy, for example, arginine (which has 6 codons) is less common in convergent TCRs, whereas leucine and serine are elevated. It's argued that there's more to convergence than just recombination biases, which makes sense. (I wonder if the trends for charged amino acids could be explained by the enrichment of convergent TCRs in CD8 T cells, which tend to have more acidic CDR3 loops). There's also a claim that the overlap between convergent and public TCRs is lower in tumors with a high mutational burden (TMB), but this part is sketchy: the definition of public TCRs is murky and hard to interpret, and the correlation between TMB and convergence-publicity overlap is modest (two cohorts with low TMB have higher overlap, and the other three have lower, but there is no association over those three, if anything the trend is in the other direction). It's also not clear why the overlap between COVID19 cohort convergent TCRs and public TCRs defined by the pre-2019 Emerson cohort should be high. A confounder here is the potential association between convergence and clonal expansion since expanded clonotypes can spawn apparently convergent TCRs due to sequencing errors. The paper "TCR Convergence in Individuals Treated With Immune Checkpoint Inhibition for Cancer" (Ref#5 here) gives evidence that sequencing errors may be inflating convergence in this specific dataset.
- Convergent TCRs are more likely to be antigen-specific: This is nicely shown on two datasets: the large dextramer dataset from 10x genomics, and the COVID19 datasets from Adaptive biotech. But given previous work on TCR convergence, for example, the Pogorelyy ALICE paper, and many others, this is also not super-surprising.
- Convergent T cells exhibit a CD8+ cytotoxic gene signature: This is based on a nice analysis of mouse and human single-cell datasets. One striking finding is that convergent TCRs are WAY more common in CD8+ T cells than in CD4+ T cells. It would be interesting to know how much of this could be explained by greater clonal expansion of CD8+ T cells, together with sequencing errors. A subtle point here is that some of the P values are probably inflated by the presence of expanded clonotypes: a group of cells belonging to the same expanded clonotype will tend to have similar gene expression (and therefore similar cluster membership), and will necessarily all be either convergent or not convergent collectively since they share the same TCR. So it's probably not quite right to treat them as independent for the purposes of assessing associations between gene expression clusters and convergence (or any other TCR-defined feature). You can see evidence for clonal expansion in Figure 3C, where TRAV genes are among the most enriched, suggesting that Cluster 04 may contain expanded clones.
- TCR convergence is associated with the clinical outcome of ICB treatment: The associations for the first analysis are described as significant in the text, and they are, but just barely (0.045 and 0.047, but you have to check the figure to see that).
- Introduction/Discussion: Overall, the authors could do a better job citing previous work on convergence, for example, papers from Venturi on convergent recombination and the work from Mora and Walczak (ALICE, another recombination modeling). They also present the use of convergence as an ICB biomarker as a novel finding, but Ref 5 introduces this concept and validates it in another cohort. Ref 5 also has a careful analysis of the link between sequencing errors and convergence, which could have been more carefully considered here.
-