Little skate genome provides insights into genetic programs essential for limb-based locomotion
Curation statements for this article:-
Curated by eLife
eLife assessment
This manuscript provides an improved version of the little skate genome, which will be of great interest to the field of comparative genomics and evolutionary biology. The authors use the genome to compare gene expression and chromatin accessibility profiles in motor neurons of the little skate and other species (mouse, chicken), aiming to predict conserved and divergent gene regulatory mechanisms underlying motor neuron development. While the manuscript contributes a valuable resource to the field, more rigorous analyses and experimental validation are needed to support the major claims of this study.
This article has been Reviewed by the following groups
Discuss this preprint
Start a discussion What are Sciety discussions?Listed in
- Evaluated articles (eLife)
Abstract
The little skate Leucoraja erinacea , a cartilaginous fish, displays pelvic fin driven walking-like behavior using genetic programs and neuronal subtypes similar to those of land vertebrates. However, mechanistic studies on little skate motor circuit development have been limited, due to a lack of high-quality reference genome. Here, we generated an assembly of the little skate genome, with precise gene annotation and structures, which allowed post-genome analysis of spinal motor neurons (MNs) essential for locomotion. Through interspecies comparison of mouse, skate and chicken MN transcriptomes, shared and divergent gene expression profiles were identified. Comparison of accessible chromatin regions between mouse and skate MNs predicted shared transcription factor (TF) motifs with divergent ones, which could be used for achieving differential regulation of MN-expressed genes. A greater number of TF motif predictions were observed in MN-expressed genes in mouse than in little skate. These findings suggest conserved and divergent molecular mechanisms controlling MN development of vertebrates during evolution, which might contribute to intricate gene regulatory networks in the emergence of a more sophisticated motor system in tetrapods.
Article activity feed
-

-

Author Response:
Reviewer #1 (Public Review):
This manuscript investigates the gene regulatory mechanisms that are involved in the development and evolution of motor neurons, utilizing cross-species comparison of RNA-sequencing and ATAC-sequencing data from little skate, chick and mouse. The authors suggest that both conserved and divergent mechanisms contribute to motor neuron specification in each species. They also claim that more complex regulatory mechanisms have evolved in tetrapods to accommodate sophisticated motor behaviors. While this is strongly suggested by the authors' ATAC-seq data, some additional validation would be required to thoroughly support this claim.
Strengths of the manuscript:
- The manuscript provides a valuable resource to the field by generating an assembly of the little skate genome, containing precise …
Author Response:
Reviewer #1 (Public Review):
This manuscript investigates the gene regulatory mechanisms that are involved in the development and evolution of motor neurons, utilizing cross-species comparison of RNA-sequencing and ATAC-sequencing data from little skate, chick and mouse. The authors suggest that both conserved and divergent mechanisms contribute to motor neuron specification in each species. They also claim that more complex regulatory mechanisms have evolved in tetrapods to accommodate sophisticated motor behaviors. While this is strongly suggested by the authors' ATAC-seq data, some additional validation would be required to thoroughly support this claim.
Strengths of the manuscript:
- The manuscript provides a valuable resource to the field by generating an assembly of the little skate genome, containing precise gene annotations that can now be utilized to perform gene expression and epigenetic analyses. The authors take advantage of this novel resource to identify novel gene expression programs and regulatory modules in little skate motor neurons.
- Cross-species RNA-seq and ATAC-seq data comparisons are combined in a powerful approach to identify novel mechanisms that control motor neuron development and evolution.
Weaknesses:
- It is surprising that the analysis of RNA-seq datasets between mouse, chick, and little skate only identified 5 genes that are common between the 3 species, especially given the authors' previous work identifying highly conserved molecular programs between little skate and mouse motor neurons, including core transcription factors (Isl1, Hb9, Lhx3), Hox genes and cholinergic transmission genes. This raises some questions about the robustness of the sequencing data and whether the genes identified represent the full transcriptome of these motor neurons.
To address reviewer #1’s questions, we have generated RNA sequencing data with mouse forelimb MNs and re-analyzed the RNA-seq data using only the homologous MN populations (Figure 3) among different species. As a result, many genes (1038 genes) are commonly expressed in MNs in different species, including many known MN marker genes. In the result section, we have added the following:
“The evolution of genetic programs in MNs was investigated unbiasedly by comparing highly expressed genes in pec-MNs (percentile expression > 70) of little skate with the ones from MNs of mouse and chick, two well-studied tetrapod species. In order to compare gene expression with homologous cell types from each species, we performed RNA sequencing on forelimb MNs of mouse embryos at embryonic day 13.5 (e13.5) and wing level MNs of chick embryos at Hamburger-Hamilton (HH) stage 26–27…”
We have also compared our re-analysis with previous results in Figure 2–figure supplement 1, shown above. Most of the fin MN genes (21/24) are highly expressed in pecMNs (percentile > 70), consistent with the previous in situ experiments. In the Results we have added the following:
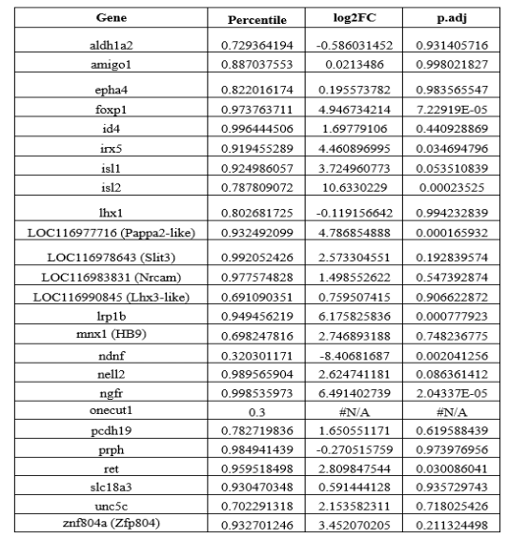
“Although the total number of DEGs are different from the previous data (592 vs. 135 genes in pec-MN DEGs), which might be caused by different statistical analysis with different reference genome, previous RNA-seq data based on de novo assembly and annotation using zebrafish was mostly recapitulated in our DEG analysis based on our new skate genome (21 out of 24 previous fin MN marker genes have the expression level ranked above 70th percentile in Pec-MNs; Figure 2‒figure supplement 1).”
- The authors suggest based on analysis of binding motifs in their ATAC-seq data that the greater number of putative binding sites in the mouse MNs allows for a higher complexity of regulation and specialization of putative motor pools. This could certainly be true in theory but needs to be further validated. The authors show FoxP1 as an example, which seems to be more heavily regulated in the mouse, but there is no evidence that FoxP1 expression profile is different between mouse and skate. It is suggested in Fig.5 that FoxP1 might be differentially regulated by SnaiI in mouse and skate but the expression of SnaiI in MNs in either species is not shown.
We have added further discussion and data about differential expression of Foxp1 in mouse and little skate in Figure 5–figure supplement 16 and have discussed as follows:
“Foxp1, the major limb/fin MN determinant appears to be differentially regulated in tetrapod and little skate. Although Foxp1 is expressed in and required for the specification of all limb MNs in tetrapods, Foxp1 is downregulated in Pea3 positive MN pools during maturation in mice (Catela et al., 2016; Dasen et al., 2008). In addition, preganglionic motor column neurons (PGC MNs) in the thoracic spinal cord of mouse and chick express half the level of Foxp1 expression than limb MNs. Although PGC neurons have not yet been identified in little skate, we tested the expression level of Foxp1 using a previously characterized tetrapod PGC marker, pSmad. We observed that Foxp1 is not expressed in MNs that express pSmad (Figure 5‒figure supplement 3). Since there is currently no known marker for PGC MNs in little skate, our conclusion should be taken with caution.”
As for Snai1, in the revision we performed a motif enrichment analysis with an unbiased gene list where Snai1 didn’t show up. However, when we performed an RNA in situ hybridization experiment for Snai1 (Figure 5–figure supplement 3), we found that Snai1 is expressed in MNs of both mouse and little skate, but not in chick, which has been shown previously (Cheung et al., 2005). In order to examine the function of Snai1 in the regulation of Foxp1 expression, we ectopically expressed Snai1 in chick spinal cord by performing in ovo electroporation. However, we did not detect any changes in Foxp1. Instead we observed an increase in the number of neurons and abnormal MN exits from the spinal cord, which is the reminiscent of a previous observation (Zander et al., 2014). Although we did not detect any changes in Foxp1 expression, we cannot rule out the possibility that Snai1 regulates Foxp1 in mouse and little skate, which may require a gene knock out experiment. Because binding sites of Snai1 were not enriched in the new gene sets that we analyzed in the revision, we have not further discussed the Snai1 in the text.
- In their discussion section the authors state that they found both conserved and divergent molecular markers across multiple species but they do not validate the expression of novel markers in either category beyond RNA-seq, for example by in situ or antibody staining.
We have added RNA in situ hybridization results in Figure 3C and Figure 3–figure supplement 1 and 2. Most of the genes were expressed in tissues in accordance with the sequencing results (6 out of 9 common MN genes; 4 out of 6 mouse specific genes; 5 out of 7 skate specific genes). Specifcally, Uchl1, Slc5a7, Alcam, and Serinc1 are expressed in MNs of all three species; Coch, Ppp1rc, Ctxn1, and Clmp are expressed in MNs of mouse but not in MNs of other species; Eya1, Etv5, Dnmbp, and Spint1 are expressed in MNs of skate but not in MNs of other species. In the result section, we have summarized the results as follow:
“These results were validated by performing RNA in situ hybridization in tissue sections on a subset of species-specific genes …”
-

eLife assessment
This manuscript provides an improved version of the little skate genome, which will be of great interest to the field of comparative genomics and evolutionary biology. The authors use the genome to compare gene expression and chromatin accessibility profiles in motor neurons of the little skate and other species (mouse, chicken), aiming to predict conserved and divergent gene regulatory mechanisms underlying motor neuron development. While the manuscript contributes a valuable resource to the field, more rigorous analyses and experimental validation are needed to support the major claims of this study.
-

Reviewer #1 (Public Review):
This manuscript investigates the gene regulatory mechanisms that are involved in the development and evolution of motor neurons, utilizing cross-species comparison of RNA-sequencing and ATAC-sequencing data from little skate, chick and mouse. The authors suggest that both conserved and divergent mechanisms contribute to motor neuron specification in each species. They also claim that more complex regulatory mechanisms have evolved in tetrapods to accommodate sophisticated motor behaviors. While this is strongly suggested by the authors' ATAC-seq data, some additional validation would be required to thoroughly support this claim.
Strengths of the manuscript:
The manuscript provides a valuable resource to the field by generating an assembly of the little skate genome, containing precise gene annotations that …
Reviewer #1 (Public Review):
This manuscript investigates the gene regulatory mechanisms that are involved in the development and evolution of motor neurons, utilizing cross-species comparison of RNA-sequencing and ATAC-sequencing data from little skate, chick and mouse. The authors suggest that both conserved and divergent mechanisms contribute to motor neuron specification in each species. They also claim that more complex regulatory mechanisms have evolved in tetrapods to accommodate sophisticated motor behaviors. While this is strongly suggested by the authors' ATAC-seq data, some additional validation would be required to thoroughly support this claim.
Strengths of the manuscript:
The manuscript provides a valuable resource to the field by generating an assembly of the little skate genome, containing precise gene annotations that can now be utilized to perform gene expression and epigenetic analyses. The authors take advantage of this novel resource to identify novel gene expression programs and regulatory modules in little skate motor neurons.
Cross-species RNA-seq and ATAC-seq data comparisons are combined in a powerful approach to identify novel mechanisms that control motor neuron development and evolution.
Weaknesses:
It is surprising that the analysis of RNA-seq datasets between mouse, chick, and little skate only identified 5 genes that are common between the 3 species, especially given the authors' previous work identifying highly conserved molecular programs between little skate and mouse motor neurons, including core transcription factors (Isl1, Hb9, Lhx3), Hox genes and cholinergic transmission genes. This raises some questions about the robustness of the sequencing data and whether the genes identified represent the full transcriptome of these motor neurons.
The authors suggest based on analysis of binding motifs in their ATAC-seq data that the greater number of putative binding sites in the mouse MNs allows for a higher complexity of regulation and specialization of putative motor pools. This could certainly be true in theory but needs to be further validated. The authors show FoxP1 as an example, which seems to be more heavily regulated in the mouse, but there is no evidence that FoxP1 expression profile is different between mouse and skate. It is suggested in Fig.5 that FoxP1 might be differentially regulated by SnaiI in mouse and skate but the expression of SnaiI in MNs in either species is not shown.
In their discussion section the authors state that they found both conserved and divergent molecular markers across multiple species but they do not validate the expression of novel markers in either category beyond RNA-seq, for example by in situ or antibody staining.
-

Reviewer #2 (Public Review):
The cartilaginous fish Leucoraja erinacea (little skate) exhibits core features of tetrapod locomotion, thus it is a key species to study conserved principles of tetrapod motor neuron development. Baek et al. provide a new and improved version of the little skate genome, which will be of great interest to the field of comparative genomics and evolutionary biology. In addition, the manuscript uses already published RNA-seq data from skate, mouse and chicken, as well as newly generated ATAC-seq data in little skate to try to reach a better understanding of the regulatory networks underlying motor neuron specification in these different species. While the question is of key importance, the bioinformatics comparisons followed by the authors seem inadequate and deeply biased. All comparative analyses are …
Reviewer #2 (Public Review):
The cartilaginous fish Leucoraja erinacea (little skate) exhibits core features of tetrapod locomotion, thus it is a key species to study conserved principles of tetrapod motor neuron development. Baek et al. provide a new and improved version of the little skate genome, which will be of great interest to the field of comparative genomics and evolutionary biology. In addition, the manuscript uses already published RNA-seq data from skate, mouse and chicken, as well as newly generated ATAC-seq data in little skate to try to reach a better understanding of the regulatory networks underlying motor neuron specification in these different species. While the question is of key importance, the bioinformatics comparisons followed by the authors seem inadequate and deeply biased. All comparative analyses are performed with lists of genes that for each species are selected following different criteria or compared with different neuronal populations, introducing important biases that will later limit the conclusions driven by the authors. Moreover, additional key aspects of evolution, such as paralog substitution or expression of species-specific genes should also be studied. Finally, the lack of experimental validations also reduces the impact of the conclusions, which at this point are highly speculative.
-

Reviewer #3 (Public Review):
Yoo et al. present a greatly improved assembly and annotation of the little skate genome. Using this new assembly and annotation, the authors re-analyze previously published gene expression data from little skate motor neurons, which were initially analyzed using instead zebrafish gene models. New in this paper is the ATAC-seq showing regions of chromatin accessibility, which was made possible by the improved assembly. Finally, the authors search for predicted transcription factor binding motifs in the vicinity of little skate motor neuron-specific genes to arrive at a model for gene regulatory networks operating in this species. They compare this gene expression and accessibility data and predicted network connections to those observed or predicted in other vertebrates (i.e. tetrapods).
The improved …
Reviewer #3 (Public Review):
Yoo et al. present a greatly improved assembly and annotation of the little skate genome. Using this new assembly and annotation, the authors re-analyze previously published gene expression data from little skate motor neurons, which were initially analyzed using instead zebrafish gene models. New in this paper is the ATAC-seq showing regions of chromatin accessibility, which was made possible by the improved assembly. Finally, the authors search for predicted transcription factor binding motifs in the vicinity of little skate motor neuron-specific genes to arrive at a model for gene regulatory networks operating in this species. They compare this gene expression and accessibility data and predicted network connections to those observed or predicted in other vertebrates (i.e. tetrapods).
The improved assembly and reanalysis of gene expression are of great use for the study of vertebrate motor neuron development and evolution. The ATAC-seq data are new and highly valuable. The thorough analysis of predicted binding sites is impressive and hints at differences in gene regulatory network architecture between cartilaginous fish and tetrapods.
A major weakness of this paper is the fact that the transcription factor binding site analysis is entirely dependent on bioinformatic predictions, as pointed out by the paper's limitations statement. The authors recognize that there is no actual binding site data obtained using little skate proteins, cells, or DNA (e.g. no ChIP-seq, no knockdowns, no cis-regulatory DNA reporters or mutations, etc). Unfortunately, this results in several unsubstantiated claims made throughout the paper, in which the presence of predicted binding sites is taken as a regulatory connection between genes.
-