Human influenza virus infection elicits distinct patterns of monocyte and dendritic cell mobilization in blood and the nasopharynx
Curation statements for this article:-
Curated by eLife
eLife assessment
This study presents a careful evaluation of the distribution of monocytes and dendritic cells in the the blood and nasopharyngeal aspirates of patients with mild respiratory tract infections. There are some interesting differences between monocytes and dendritic cells and variations with patient age. This is an important contribution to understanding monocyte and DC subset specific functions in dependency on the tissue microenvironment.
This article has been Reviewed by the following groups
Discuss this preprint
Start a discussion What are Sciety discussions?Listed in
- Evaluated articles (eLife)
Abstract
During respiratory viral infections, the precise roles of monocytes and dendritic cells (DCs) in the nasopharynx in limiting infection and influencing disease severity are incompletely described. We studied circulating and nasopharyngeal monocytes and DCs in healthy controls (HCs) and in patients with mild to moderate infections (primarily influenza A virus [IAV]). As compared to HCs, patients with acute IAV infection displayed reduced DC but increased intermediate monocytes frequencies in blood, and an accumulation of most monocyte and DC subsets in the nasopharynx. IAV patients had more mature monocytes and DCs in the nasopharynx, and higher levels of TNFα, IL-6, and IFNα in plasma and the nasopharynx than HCs. In blood, monocytes were the most frequent cellular source of TNFα during IAV infection and remained responsive to additional stimulation with TLR7/8L. Immune responses in older patients skewed towards increased monocyte frequencies rather than DCs, suggesting a contributory role for monocytes in disease severity. In patients with other respiratory virus infections, we observed changes in monocyte and DC frequencies in the nasopharynx distinct from IAV patients, while differences in blood were more similar across infection groups. Using SomaScan, a high-throughput aptamer-based assay to study proteomic changes between patients and HCs, we found differential expression of innate immunity-related proteins in plasma and nasopharyngeal secretions of IAV and SARS-CoV-2 patients. Together, our findings demonstrate tissue-specific and pathogen-specific patterns of monocyte and DC function during human respiratory viral infections and highlight the importance of comparative investigations in blood and the nasopharynx.
Article activity feed
-

-

Author Response
Reviewer #1 (Public Review):
Strength: The study is summarizing a large cohort of human samples of blood, nasal swabs and nasopharyngeal aspirates. This is very uncommon as most of the time studies focus on the blood and serum of patients. Within the study, 3 monocyte and 3 DC subsets have been followed in healthy and Influenza A virus-infected persons. The study also includes functional data on the responsiveness of Influenza A virus-infected DC and monocyte populations. The authors achieved their aims in that they were able to show that the tissue microenvironment is important to understand subset specific migration and activation behavior in Influenza A virus infection and in addition that it matters with which kind of agent a person is infected. Thus, this study also impacts a better understanding of vaccine …
Author Response
Reviewer #1 (Public Review):
Strength: The study is summarizing a large cohort of human samples of blood, nasal swabs and nasopharyngeal aspirates. This is very uncommon as most of the time studies focus on the blood and serum of patients. Within the study, 3 monocyte and 3 DC subsets have been followed in healthy and Influenza A virus-infected persons. The study also includes functional data on the responsiveness of Influenza A virus-infected DC and monocyte populations. The authors achieved their aims in that they were able to show that the tissue microenvironment is important to understand subset specific migration and activation behavior in Influenza A virus infection and in addition that it matters with which kind of agent a person is infected. Thus, this study also impacts a better understanding of vaccine design for respiratory viruses.
We thank Reviewer 1 for highlighting what we believe to be the greatest strengths of our study. The key feature of this study was to generate a comprehensive description of monocytes and dendritic cells (DC) in the human nasopharynx during influenza A virus infection, and to provide a comparison with healthy and convalescent individuals. Further, we wished to emphasize the value of studying the nasopharynx during respiratory viral infections, particularly in light of the ongoing COVID-19 pandemic. We describe a non-invasive method to (longitudinally) sample this anatomical compartment that allows retrieval of intact immune cells as well as mucosal fluid for soluble marker analysis. We also believe that the addition of proteomic profiles in the different compartments (new Figure 7) further highlights the importance of the tissue microenvironment.
Weakness: In the described study, the authors used a different nomenclature to introduce the DC subsets. This is confusing and the authors should stick to the nomenclature introduced by Guilliams et al., 2014 (doi.org/10.1038/nri3712) and commented in Ginhoux et al., 2022 (DOI: 10.1038/s41577-022-00675-7 ) or at least should introduce the alternative names (cDC1, cDC2, expression markers XCR1, CD172a/Sirpa). Further, Segura et al., 2013 (doi: 10.1084/jem.20121103) showed that all three DC subpopulations were able to perform cross-presentation when directly isolated. Overall, a more up-to-date introduction would be useful.
Reviewer 1 commented on the DC nomenclature used in the manuscript. We agree that our manuscript would benefit from appropriately updating the DC nomenclature. We therefore revised the text, and now we refer to the subsets previously described as CD1c+ and CD141+ myeloid DCs (MDC) as cDC2 and CDC1 subsets, respectively. We have also modified the text in the Introduction of the revised manuscript to reflect the same and give a more up-to-date introduction of DC subsets (marked-up version lines 75-81).
As the data of this was already obtained in 2016-2018 it is clear that the FACS panel was not developed to study DC3. If possible, the authors might be able to speculate about the role of this subset in their data set. Moreover, there were other studies on SARS-CoV-2 infection and DC subset analyses in blood (line 87, and line 489) e.g. Winheim et al., (DOI: 10.1371/journal.ppat.1009742 ), which the authors should introduce and discuss in regard to their own data.
As reviewer 1 accurately pointed out, the flow cytometry panel used in this study was indeed not developed to study the DC3 subset. The data was obtained in 2016-2018, and lack the typical markers used to identify the DC3 subset, such as CD163, BTLA and CD5 (Cytlak et al, https://doi.org/10.1016/j.immuni.2020.07.003, Villani et al, https://doi.org/10.1126/science.aah4573). Due to the constraints of the panel, we would not be able to accurately identify DC3s. However, in an attempt to dig deeper into the data that is available, we re-analyzed the data to identify CD14+CD1c+ cells among the lineage–HLADR+CD16–CD14+ cells, here collectively called “mo-DC”. This population is likely a combination of monocytes upregulating CD1c and bona fide DC3 expressing CD14. Accordingly, the gating strategy was updated in Supplementary figure 1 (marked-up version lines 192-194), and new data plot in Figure 2H (marked-up version lines 208-220) summarizes the changes observed in mo-DC numbers in IAV patients between blood and the nasopharynx. Parallel to the pattern seen in other DC subsets, mo-DC frequencies are reduced in blood and we observed an increase (not significant) in the nasopharynx.
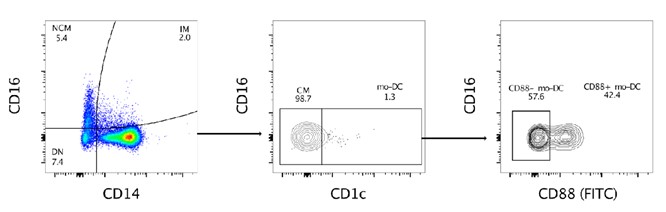
As CD88 was not included in the original panel, it was not possible to discriminate between bona fide monocytes and DC3s. We performed a staining of PBMCs (buffy coat) with CD88 (FITC) added to the original flow panel used in the study, to assess if CD88 can be helpful for future studies (Reviewer figure 1). The staining showed that some cells in the mo-DC population are CD88 positive, indicating a bona fide monocyte origin, whereas some are negative, indicating that they are bona fide DC3 expressing CD14. (Bourdely et al, https://doi.org/10.1016/j.immuni.2020.06.002).
Reviewer figure 1. Expression of CD88 in the “mo-DC” population. Cells from a buffy coat were stained with the flow cytometry panel used in the manuscript, with the addition of CD88 (FITC). Within the CD14+CD1c+ population, the “mo-DC” population, we identified both CD88+ and CD88- cells.
Reviewer 1 also suggested citing Winheim et al (https://doi.org/10.1371/journal.ppat.1009742), and we thank them for their suggestion. We have now cited Winheim et al, and two additional reports (Kvedaraite et al, https://doi.org/10.1073/pnas.2018587118 and Affandi et al, https://doi.org/10.3389/fimmu.2021.697840) describing a depletion of DC3s (and other DC subsets) from circulation, and functional impairment of DCs following SARS-CoV-2 infection. Further, Winheim et al observed an increased frequency of a CD163+CD14+ subpopulation within the DC3s, which correlated with systemic inflammatory responses in SARS-CoV-2 infection. We speculate that perhaps in IAV infection too, DC3s may follow the trend of other DC subsets and be found in increased numbers in the nasopharynx (marked-up version lines 75-81 and 543-552).
Taken together, although the data are very important and very interesting, my overall impression of the manuscript is that in the era of RNA seq and scRNA seq analyses the study lacks a bit of comprehensiveness.
The final comment from reviewer 1 is well taken, in that our study does not include RNA-seq analyses. Again, we ask Reviewer 1 to take into consideration the challenging material we worked with in our study in combination with the COVID-19 pandemic that subsequently has excluded recruitment of new influenza patients to the study. The cell numbers and viability in the nasopharyngeal aspirates limit what experimental approaches can be done simultaneously, and flow cytometry seemed to be the best approach for the study. However, we agree that in future studies, both our own and those of others in the field, will greatly benefit from single cell analysis of nasopharyngeal immune cells, and from generating transcriptomic or epigenetic profiles of these cells. Unfortunately, it is a limitation that we are currently unable to overcome within the scope of this revision. Despite this weakness, we agree with Reviewer 1 that the methods we developed and the data we generated are important and interesting.
Moreover, we have added additional proteomics data from both NPA and plasma from influenza and COVID-19 patients, using the SomaScan platform (new Figure 7) (marked-up version lines 472-511, 738-755 and 768-792). We also included a supplementary table listing enriched pathway data from gProfiler. Briefly, our data showed sizeable changes within the blood and nasopharyngeal proteome during respiratory virus infection (IAV or SARS-CoV-2), as compared to healthy controls. Importantly, we found several differentially expressed proteins unique to the nasopharynx that were not seen in blood, and pathway analysis highlighted “host immune responses” and “innate immunity” pathways, containing TNF, IL-6, ISG15, IL-18R, CCL7, CXCL10 (IP-10), CXCL11, GZMB, SEMA4A, S100A8, S100A9. These findings are in line with our flow cytometry data, and support our hypothesis that the immunological response to viral infection in the upper airways differ from that in matching plasma samples. One of the main messages in this manuscript is the importance of looking at the site of infection, and not only at systemic immune responses to better understand respiratory viral infections in humans. We believe that the addition of the proteomics data serves to further highlight this point.
Reviewer #2 (Public Review):
This study aims to describe the distribution and functional status of monocytes and dendritic cells in the blood and nasopharyngeal aspirate (NPA) after respiratory viral infection in more than 50 patients affected by influenza A, B, RSV and SARS-CoV2. The authors use flow cytometry to define HLA-DR+ lineage negative cells, and within this gate, classical, intermediate and non-classical monocytes and CD1c+, CD141+, and CD123+ dendritic cells (DC). They show a large increase in classical monocytes in NPA and an increase in intermediate monocytes in blood and NPA, with more subtle changes in non-classical monocytes. Changes in intermediate monocytes were age-dependent and resolution was seen with convalescence. While blood monocytes tended to increase in blood and NPA, DC frequency was reduced in blood but also increased in NPA. There were signs of maturation in monocytes and DC in NPA compared with blood as judged by expression of HLA-DR and CD86. Cytokine levels in NPA were increased in infection in association with enrichment of cytokine-producing cells. Various patterns were observed in different viral infections suggesting some specificity of pathogen response. The work did not fully document the diversity of human myeloid cells that have arisen from single-cell transcriptomics over the last 5 years, notably the classification of monocytes which shows only two distinct subsets (intermediate cannot be distinguished from classical), distinct populations of DC1, DC2 and DC3 (DC2 and 3 both having CD1c, but different levels of monocyte antigens), and the lack of distinction provided by CD123 which also includes a precursor population of AXL+SIGLEC6+ myeloid cells in addition to plasmacytoid DC. Furthermore, some greater precision of the gating could have been achieved for the subsets presented. Specifically, CD34+ cells were not excluded from the HLA-DR+ lineage- gate, and the threshold of CD11c may have excluded some DC1 owing to the low expression of this antigen. Overall, the work shows that interesting results can be obtained by comparing myeloid populations of blood and NPA during viral infection and that lineage, viral and age-specific patterns are observed. However, the mechanistic insights for host defense provided by these observations remain relatively modest.
We thank Reviewer 2 for their assessment of our manuscript and summarizing our key findings in their public review. As reviewer 2 noted, our study describes changes in frequencies of monocytes and DCs during acute IAV infection, in blood and in the nasopharynx. Additionally, we also demonstrate pathogen-specific changes in both compartments. Reviewer 2 also highlighted a drawback of our study- that the approach did not fully capture the breadth of monocyte and DC diversity as it currently stands. Despite this, the findings we presented here laid the groundwork for continued research and led to significant progress, including mechanistic insights (Falck-Jones et al, https://doi.org/10.1172/JCI144734 and Cagigi et al, https://doi.org/10.1172/jci.insight.151463, Havervall et al. https://doi.org/10.1056/nejmc2209651 and Marking et al. Lancet Infectious Diseases in press), in understanding the role of myeloid cells in the human airways during viral infections.
-

eLife assessment
This study presents a careful evaluation of the distribution of monocytes and dendritic cells in the the blood and nasopharyngeal aspirates of patients with mild respiratory tract infections. There are some interesting differences between monocytes and dendritic cells and variations with patient age. This is an important contribution to understanding monocyte and DC subset specific functions in dependency on the tissue microenvironment.
-

Reviewer #1 (Public Review):
The manuscript is of importance for vaccine design and understanding tissue microenvironmental influence on the functionality of monocytes and DCs to respiratory viruses such as Influenza A virus or SARS-CoV-2 virus. The methods used were mainly flow cytometry, ELISA, Luminex as well as TNFα secretion Assay Detection. The authors wanted to evaluate if and how the tissue microenvironment might impact DC and monocyte subset presence and functionality during Influenza A virus infection.
Strength:
The study is summarizing a large cohort of human samples of blood, nasal swabs and nasopharyngeal aspirates. This is very uncommon as most of the time studies focus on the blood and serum of patients. Within the study, 3 monocyte and 3 DC subsets have been followed in healthy and Influenza A virus-infected persons. The …Reviewer #1 (Public Review):
The manuscript is of importance for vaccine design and understanding tissue microenvironmental influence on the functionality of monocytes and DCs to respiratory viruses such as Influenza A virus or SARS-CoV-2 virus. The methods used were mainly flow cytometry, ELISA, Luminex as well as TNFα secretion Assay Detection. The authors wanted to evaluate if and how the tissue microenvironment might impact DC and monocyte subset presence and functionality during Influenza A virus infection.
Strength:
The study is summarizing a large cohort of human samples of blood, nasal swabs and nasopharyngeal aspirates. This is very uncommon as most of the time studies focus on the blood and serum of patients. Within the study, 3 monocyte and 3 DC subsets have been followed in healthy and Influenza A virus-infected persons. The study also includes functional data on the responsiveness of Influenza A virus-infected DC and monocyte populations. The authors achieved their aims in that they were able to show that the tissue microenvironment is important to understand subset specific migration and activation behavior in Influenza A virus infection and in addition that it matters with which kind of agent a person is infected. Thus, this study also impacts a better understanding of vaccine design for respiratory viruses.Weakness:
In the described study, the authors used a different nomenclature to introduce the DC subsets. This is confusing and the authors should stick to the nomenclature introduced by Guilliams et al., 2014 (doi.org/10.1038/nri3712) and commented in Ginhoux et al., 2022 (DOI: 10.1038/s41577-022-00675-7 ) or at least should introduce the alternative names (cDC1, cDC2, expression markers XCR1, CD172a/Sirpa). Further, Segura et al., 2013 (doi: 10.1084/jem.20121103) showed that all three DC subpopulations were able to perform cross-presentation when directly isolated. Overall, a more up-to-date introduction would be useful.
As the data of this was already obtained in 2016-2018 it is clear that the FACS panel was not developed to study DC3. If possible, the authors might be able to speculate about the role of this subset in their data set. Moreover, there were other studies on SARS-CoV-2 infection and DC subset analyses in blood (line 87, and line 489) e.g. Winheim et al., (DOI: 10.1371/journal.ppat.1009742 ), which the authors should introduce and discuss in regard to their own data. Taken together, although the data are very important and very interesting, my overall impression of the manuscript is that in the era of RNA seq and scRNA seq analyses the study lacks a bit of comprehensiveness. -

Reviewer #2 (Public Review):
This study aims to describe the distribution and functional status of monocytes and dendritic cells in the blood and nasopharyngeal aspirate (NPA) after respiratory viral infection in more than 50 patients affected by influenza A, B, RSV and SARS-CoV2. The authors use flow cytometry to define HLA-DR+ lineage negative cells, and within this gate, classical, intermediate and non-classical monocytes and CD1c+, CD141+, and CD123+ dendritic cells (DC). They show a large increase in classical monocytes in NPA and an increase in intermediate monocytes in blood and NPA, with more subtle changes in non-classical monocytes. Changes in intermediate monocytes were age-dependent and resolution was seen with convalescence. While blood monocytes tended to increase in blood and NPA, DC frequency was reduced in blood but …
Reviewer #2 (Public Review):
This study aims to describe the distribution and functional status of monocytes and dendritic cells in the blood and nasopharyngeal aspirate (NPA) after respiratory viral infection in more than 50 patients affected by influenza A, B, RSV and SARS-CoV2. The authors use flow cytometry to define HLA-DR+ lineage negative cells, and within this gate, classical, intermediate and non-classical monocytes and CD1c+, CD141+, and CD123+ dendritic cells (DC). They show a large increase in classical monocytes in NPA and an increase in intermediate monocytes in blood and NPA, with more subtle changes in non-classical monocytes. Changes in intermediate monocytes were age-dependent and resolution was seen with convalescence. While blood monocytes tended to increase in blood and NPA, DC frequency was reduced in blood but also increased in NPA. There were signs of maturation in monocytes and DC in NPA compared with blood as judged by expression of HLA-DR and CD86. Cytokine levels in NPA were increased in infection in association with enrichment of cytokine-producing cells. Various patterns were observed in different viral infections suggesting some specificity of pathogen response. The work did not fully document the diversity of human myeloid cells that have arisen from single-cell transcriptomics over the last 5 years, notably the classification of monocytes which shows only two distinct subsets (intermediate cannot be distinguished from classical), distinct populations of DC1, DC2 and DC3 (DC2 and 3 both having CD1c, but different levels of monocyte antigens), and the lack of distinction provided by CD123 which also includes a precursor population of AXL+SIGLEC6+ myeloid cells in addition to plasmacytoid DC. Furthermore, some greater precision of the gating could have been achieved for the subsets presented. Specifically, CD34+ cells were not excluded from the HLA-DR+ lineage- gate, and the threshold of CD11c may have excluded some DC1 owing to the low expression of this antigen. Overall, the work shows that interesting results can be obtained by comparing myeloid populations of blood and NPA during viral infection and that lineage, viral and age-specific patterns are observed. However, the mechanistic insights for host defense provided by these observations remain relatively modest.
-