Oscillatory movement of a dynein-microtubule complex crosslinked with DNA origami
Curation statements for this article:-
Curated by eLife
Evaluation Summary:
The authors describe the reconstitution of axonemal bending using polymerized microtubules, purified outer-arm dyneins, and synthesized DNA origami to cross-link two microtubules. The work is of interest for the field as it shows that bidirectional sliding and bending of microtubules can be generated by a minimal set of elements.
(This preprint has been reviewed by eLife. We include the public reviews from the reviewers here; the authors also receive private feedback with suggested changes to the manuscript. Reviewer #3 agreed to share their name with the authors.)
This article has been Reviewed by the following groups
Discuss this preprint
Start a discussion What are Sciety discussions?Listed in
- Evaluated articles (eLife)
Abstract
Bending of cilia and flagella occurs when axonemal dynein molecules on one side of the axoneme produce force and move toward the microtubule (MT) minus end. These dyneins are then pulled back when the axoneme bends in the other direction, meaning oscillatory back and forth movement of dynein during repetitive bending of cilia/flagella. There are various factors that may regulate the dynein activity, e.g. the nexin-dynein regulatory complex, radial spokes, and central apparatus. In order to understand the basic mechanism of dynein’s oscillatory movement, we constructed a simple model system composed of MTs, outer-arm dyneins, and crosslinks between the MTs made of DNA origami. Electron microscopy (EM) showed pairs of parallel MTs crossbridged by patches of regularly arranged dynein molecules bound in two different orientations, depending on which of the MTs their tails bind to. The oppositely oriented dyneins are expected to produce opposing forces when the pair of MTs have the same polarity. Optical trapping experiments showed that the dynein-MT-DNA-origami complex actually oscillates back and forth after photolysis of caged ATP. Intriguingly, the complex, when held at one end, showed repetitive bending motions. The results show that a simple system composed of ensembles of oppositely oriented dyneins, MTs, and inter-MT crosslinkers, without any additional regulatory structures, has an intrinsic ability to cause oscillation and repetitive bending motions.
Article activity feed
-

-

Author Response
Reviewer #1 (Public Review):
In this manuscript, Abdellatef et al. describe the reconstitution of axonemal bending using polymerized microtubules (MTs), purified outer-arm dyneins, and synthesized DNA origami. Specifically, the authors purified axonemal dyneins from Chlamydomonas flagella and combined the purified motors with MTs polymerized from purified brain tubulin. Using electron microscopy, the authors demonstrate that patches of dynein motors of the same orientation at both MT ends (i.e., with their tails bound to the same MT) result in pairs of MTs of parallel alignment, while groups of dynein motors of opposite orientation at both MT ends (i.e., with the tails of the dynein motors of both groups bound to different MTs) result in pairs of MTs with anti-parallel alignment. The authors then show that the dynein …
Author Response
Reviewer #1 (Public Review):
In this manuscript, Abdellatef et al. describe the reconstitution of axonemal bending using polymerized microtubules (MTs), purified outer-arm dyneins, and synthesized DNA origami. Specifically, the authors purified axonemal dyneins from Chlamydomonas flagella and combined the purified motors with MTs polymerized from purified brain tubulin. Using electron microscopy, the authors demonstrate that patches of dynein motors of the same orientation at both MT ends (i.e., with their tails bound to the same MT) result in pairs of MTs of parallel alignment, while groups of dynein motors of opposite orientation at both MT ends (i.e., with the tails of the dynein motors of both groups bound to different MTs) result in pairs of MTs with anti-parallel alignment. The authors then show that the dynein motors can slide MTs apart following photolysis of caged ATP, and using optical tweezers, demonstrate active force generation of up to ~30 pN. Finally, the authors show that pairs of anti-parallel MTs exhibit bidirectional motion on the scale of ~50-100 nm when both MTs are cross-linked using DNA origami. The findings should be of interest for the cytoskeletal cell and biophysics communities.
We thank the reviewer for these comments.
We might be misunderstanding this reviewer’s comment, but the complexes with both parallel and anti-parallel MTs had dynein molecules with their tails bound to two different MTs in most cases, as illustrated in Fig.2 – suppl.1. The two groups of dyneins produce opposing forces in a complex with parallel MTs, and majority of our complexes had parallel arrangement of the MTs. To clarify the point, we have modified the Abstract:
“Electron microscopy (EM) showed pairs of parallel MTs crossbridged by patches of regularly arranged dynein molecules bound in two different orientations depending on which of the MTs their tails bind to. The oppositely oriented dyneins are expected to produce opposing forces when the pair of MTs have the same polarity.”
Reviewer #2 (Public Review):
Motile cilia generate rhythmic beating or rotational motion to drive cells or produce extracellular fluid flow. Cilia is made of nine microtubule doublets forming a spoke-like structure and it is known that dynein motor proteins, which connects adjacent microtubule doublet, are the driving force of ciliary motion. However the molecular mechanism to generate motion is still unclear. The authors proved that a pair of microtubules stably linked by DNA-origami and driven by outer dynein arms (ODA) causes beating motion. They employed in vitro motility assay and negative stain TEM to characterize this complex. They demonstrated stable linking of microtubules and ODAs anchored on the both microtubules are essential for oscillatory motion and bending of the microtubules.
Strength
This is an interesting work, addressing an important question in the motile cilia community: what is the minimum system to generate a beating motion? It is an established fact that dynein power stroke on the microtubule doublet is the driving force of the beating motion. It was also known that the radial spoke and the central pair are essential for ciliary motion under the physiological condition, but cilia without radial spokes and the central pair can beat under some special conditions (Yagi and Kamiya, 2000). Therefore in the mechanistic point of view, they are not prerequisite. It is generally thought that fixed connection between adjacent microtubules by nexin converts sliding motion of dyneins to bending, but it was never experimentally investigated. Here the authors successfully enabled a simple system of nexin-like inter-microtubule linkage using DNA origami technique to generate oscillatory and beating motions. This enables an interesting system where ODAs form groups, anchored on two microtubules, orienting oppositely and therefore cause tag-of-war type force generation. The authors demonstrated this system under constraints by DNA origami generates oscillatory and beating motions.
The authors carefully coordinated the experiments to demonstrate oscillations using optical tweezers and sophisticated data analysis (Fourier analysis and a step-finding algorithm). They also proved, using negative stain EM, that this system contains two groups of ODAs forming arrays with opposite polarity on the parallel microtubules. The manuscript is carefully organized with impressive movies. Geometrical and motility analyses of individual ODAs used for statistics are provided in the supplementary source files. They appropriately cited similar past works from Kamiya and Shingyoji groups (they employed systems closer to the physiological axoneme to reproduce beating) and clarify the differences from this study.
We thank the reviewer for these comments.
Weakness
The authors claim this system mimics two pairs of doublets at the opposite sites from 9+2 cilia structure by having two groups of ODAs between two microtubules facing opposite directions within the pair. It is not exactly the case. In the real axoneme, ODA makes continuous array along the entire length of doublets, which means at any point there are ODAs facing opposite directions. In their system, opposite ODAs cannot exist at the same point (therefore the scheme of Dynein-MT complex of Fig.1B is slightly misleading).
Actually, opposite ODAs can exist at the same point in our system as well, and previous work using much higher concentration of dyneins (e.g, Oda et al., J. Cell biol., 2007) showed two continuous arrays of dynein molecules between a pair of microtubules. To observe the structures of individual dynein molecules we used low concentrations of dynein and searched for the areas where dynein could be observed without superposition, but there were some areas where opposite dyneins existed at the same point.
We realize that we did not clearly explain this issue, so we have revised the text accordingly.
In the 1st paragraph of Results: “In the dynein-MT complexes prepared with high concentrations of dynein, a pair of MTs in bundles are crossbridged by two continuous arrays of dynein, so that superposition of two rows of dynein molecules is observed in EM images (Haimo et al., 1979; Oda et al., 2007). On the other hand, when a low concentration of the dynein preparation (6.25–12.5 µg/ml (corresponding to ~3-6 nM outer-arm dynein)) was mixed with 20-25 µg/ml MTs (200-250 nM tubulin dimers), the MTs were only partially decorated with dynein, so that we were able to observe single layers of crossbridges without superposition in many regions.” Legend of Fig. 1(C): “Note that the geometry of dyneins in the dynein-MT complex shown in (B) mimics that of a combination of the dyneins on two opposite sides of the axoneme (cyan boxes), although the dynein arrays in (B) are not continuous.”
If they want to project their result to the ciliary beating model, more insight/explanation would be necessary. For example, arrays of dyneins at certain positions within the long array along one doublet are activated and generate force, while dyneins at different positions are activated on another doublet at the opposite site of the axoneme. This makes the distribution of dyneins and their orientations similar to the system described in this work. Such a localized activation, shown in physiological cilia by Ishikawa and Nicastro groups, may require other regulatory proteins.
We agree that the distributions of activated dyneins in 3D are extremely important in understanding ciliary beating, and that other regulatory proteins would be required to coordinate activation in different places in an axoneme. However, the main goal of this manuscript is to show the minimal components for oscillatory movements, and we feel that discussing the distributions of activated dyneins along the length of the MTs would be too complicated and beyond the scope of this study.
They attempted to reveal conformational change of ODAs induced by power stroke using negative stain EM images, which is less convincing compared to the past cryo-ET works (Ishikawa, Nicastro, Pigino groups) and negative stain EM of sea urchin outer dyneins (Hirose group), where the tail and head parts were clearly defined from the 3D map or 2D averages of two-dynein ODAs. Probably three heavy chains and associated proteins hinder detailed visualization of the tail structure. Because of this, Fig.2C is not clear enough to prove conformational change of ODA. This reviewer imagines refined subaverage (probably with larger datasets) is necessary.
As the reviewer suggests, one of the reasons for less clear averaged images compared to the past images of sea urchin ODA is the three-headed structure of Chlamydomonas ODA. Another and perhaps the bigger reason is the difficulty of obtaining clear images of dynein molecules bound between 2 MTs by negative stain EM: the stain accumulates between MTs that are ~25 nm in diameter and obscures the features of smaller structures. We used cryo-EM with uranyl acetate staining instead of negative staining for the images of sea urchin ODA-MT complexes we previously published (Ueno et al., 2008) in order to visualize dynein stalks. We agree with the reviewer that future work with larger datasets and by cryo-ET is necessary for revealing structural differences.
That having been said, we did not mean to prove structural changes, but rather intended to show that our observation suggests structural changes and thus this system is useful for analyzing structural changes in future. In the revised manuscript, we have extensively modified the parts of the paper discussing structural changes (Please see our response to the next comment).
It is not clear, from the inset of Fig.2 supplement3, how to define the end of the tail for the length measurement, which is the basis for the authors to claim conformational change (Line263-265). The appearance of the tail would be altered, seen from even slightly different view angles. Comparison with 2D projection from apo- and nucleotide-bound 3-headed ODA structures from EM databank will help.
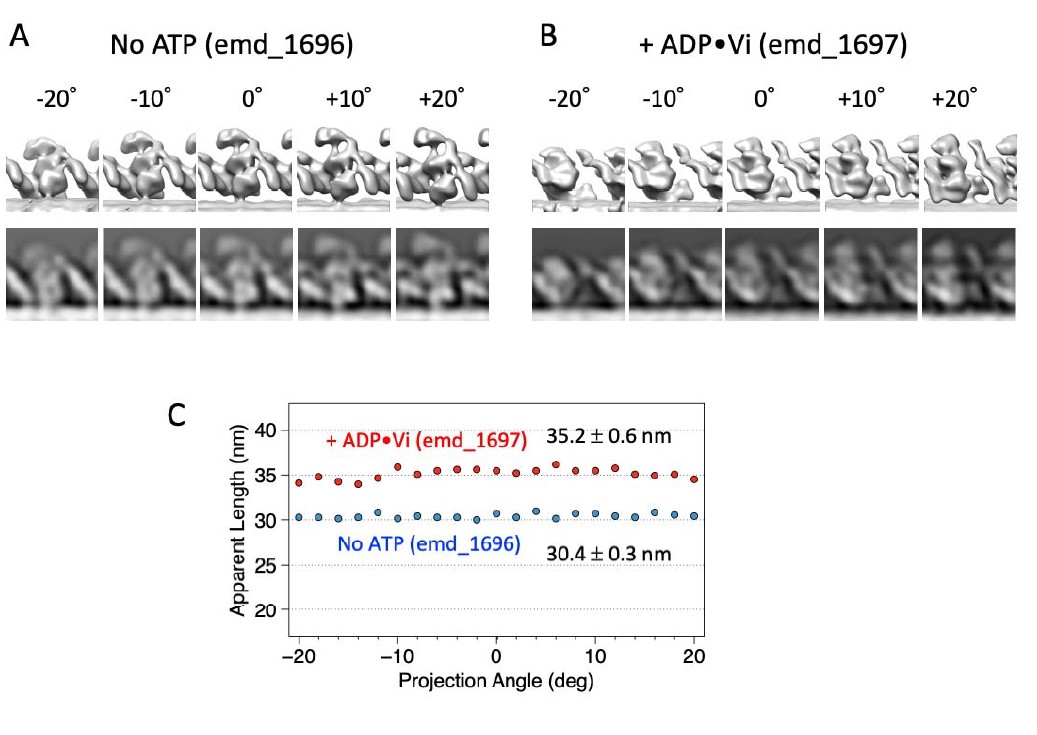
We agree with the reviewer that difference in the viewing angle affects the apparent length of a dynein molecule, although the 2 MTs crossbridged by dyneins lie on the carbon membrane and thus the variation in the viewing angle is expected to be relatively small. To examine how much the apparent length is affected by the view angle, we calculated 2D-projected images of the cryo-ET structures of Chlamydomonas axoneme (emd_1696 and emd_1697; Movassagh et al., 2010) with different view angles, and measured the apparent length of the dynein molecule using the same method we used for our negative-stain images (Author response image 1). As shown in the plot, the effect of view angles on the apparent lengths is smaller than the difference between the two nucleotide states in the range of 40 degrees measured here. Thus, we think that the length difference shown in Fig.2-suppl.4 reflects a real structural difference between no-ATP and ATP states. In addition, it would be reasonable to think that distributions of the view angles in the negative stain images are similar for both absence and presence of ATP, again supporting the conclusion.
Nevertheless, since we agree with the reviewer that we cannot measure the precise length of the molecule using these 2D images, we have revised the corresponding parts of the manuscript, adding description about the effect of view angles on the measured length in the manuscript.
Author response image 1. Effects of viewing angles on apparent length. (A) and (B) 2D-projected images of cryo-electron tomograms of Chlamydomonas outer arm dynein in an axoneme (Movassagh et al., 2010) viewed from different angles. (C) apparent length of the dynein molecule measured in 2D-projected images.
In this manuscript, we discuss two structural changes: 1) a difference in the dynein length between no-nucleotide and +ATP states (Fig.2-suppl.4), and 2) possible structural differences in the arrangement of the dynein heads (Fig.2-suppl.3). Although we realize that extensive analysis using cryo-ET is necessary for revealing the second structural change, we attempted to compare the structures of oppositely oriented dyneins, hoping that it would lead to future research. In the revised manuscript, we have added 2D projection images of emd_1696 and emd_1697 in Fig.2-suppl.3, so that the readers can compare them with our negative stain images. We had an impression that some of our 2D images in the presence of ATP resembled the cryo-ET structure with ADP.Vi, whereas some others appeared to be closer to the no-nucleotide cryo-ET structure. We have also attempted to calculate cross-correlations, but difficulties in removing the effect of MTs sometimes overlapped with a part of dynein, adjusting the magnifications and contrast of different images prevented us from obtaining reliable results.
To address this and the previous comments, we have extensively modified the section titled ‘Structures of dynein in the dynein-MT-DNA-origami complex’.
In Fig.5B (where the oscillation occurs), the microtubule was once driven >150nm unidirectionally and went back to the original position, before oscillation starts. Is it always the case that relatively long unidirectional motion and return precede oscillation? In Fig.7B, where the authors claim no oscillation happened, only one unidirectional motion was shown. Did oscillation not happen after MT returned to the original position?
Long unidirectional movement of ~150 nm was sometimes observed, but not necessarily before the start of oscillation. For example, in Figure 5 – figure supplement 1A, oscillation started soon after the UV flash, and then unidirectional movement occurred.
With the dynein-MT complex in which dyneins are unidirectionally aligned (Fig.7B, Fig.7-suppl.2), the MTs kept moving and escaped from the trap or just stopped moving probably due to depletion of ATP, so we did not see a MT returning to the original position.
Line284-290: More characterization of bending motion will be necessary (and should be possible). How high frequency is it? Do they confirm that other systems (either without DNA-origami or without ODAs arraying oppositely) cannot generate repetitive beating?
The frequencies of the bending motions measured from the movies in Fig.8 and Fig.8-suppl.1 were 0.6 – 1 Hz, and the motions were rather irregular. Even if there were complexes bending at high frequencies, it would not have been possible to detect them due to the low time resolution of these fluorescence microscopy experiments (~0.1 s). Future studies at a higher time resolution will be necessary for further characterization of bending motions.
To observe bending motions, the dynein-MT complex should be fixed to the glass or a bead at one part of the complex while the other end is free in solution. With the dynein-MT-DNA-origami complexes, we looked for such complexes and found some showing bending motions as in Fig. 8. To answer the reviewer’s question asking if we saw repetitive bending in other systems, we checked the movies of the complexes without DNA-origami or without ODAs arraying oppositely but did not notice any repetitive bending motions. However, future studies using the system with a higher temporal resolution and perhaps with an improved method for attaching the complex would be necessary in these cases as well.
-

Evaluation Summary:
The authors describe the reconstitution of axonemal bending using polymerized microtubules, purified outer-arm dyneins, and synthesized DNA origami to cross-link two microtubules. The work is of interest for the field as it shows that bidirectional sliding and bending of microtubules can be generated by a minimal set of elements.
(This preprint has been reviewed by eLife. We include the public reviews from the reviewers here; the authors also receive private feedback with suggested changes to the manuscript. Reviewer #3 agreed to share their name with the authors.)
-

Reviewer #1 (Public Review):
In this manuscript, Abdellatef et al. describe the reconstitution of axonemal bending using polymerized microtubules (MTs), purified outer-arm dyneins, and synthesized DNA origami. Specifically, the authors purified axonemal dyneins from Chlamydomonas flagella and combined the purified motors with MTs polymerized from purified brain tubulin. Using electron microscopy, the authors demonstrate that patches of dynein motors of the same orientation at both MT ends (i.e., with their tails bound to the same MT) result in pairs of MTs of parallel alignment, while groups of dynein motors of opposite orientation at both MT ends (i.e., with the tails of the dynein motors of both groups bound to different MTs) result in pairs of MTs with anti-parallel alignment. The authors then show that the dynein motors can slide …
Reviewer #1 (Public Review):
In this manuscript, Abdellatef et al. describe the reconstitution of axonemal bending using polymerized microtubules (MTs), purified outer-arm dyneins, and synthesized DNA origami. Specifically, the authors purified axonemal dyneins from Chlamydomonas flagella and combined the purified motors with MTs polymerized from purified brain tubulin. Using electron microscopy, the authors demonstrate that patches of dynein motors of the same orientation at both MT ends (i.e., with their tails bound to the same MT) result in pairs of MTs of parallel alignment, while groups of dynein motors of opposite orientation at both MT ends (i.e., with the tails of the dynein motors of both groups bound to different MTs) result in pairs of MTs with anti-parallel alignment. The authors then show that the dynein motors can slide MTs apart following photolysis of caged ATP, and using optical tweezers, demonstrate active force generation of up to ~30 pN. Finally, the authors show that pairs of anti-parallel MTs exhibit bidirectional motion on the scale of ~50-100 nm when both MTs are cross-linked using DNA origami. The findings should be of interest for the cytoskeletal cell and biophysics communities.
-

Reviewer #2 (Public Review):
Motile cilia generate rhythmic beating or rotational motion to drive cells or produce extracellular fluid flow. Cilia is made of nine microtubule doublets forming a spoke-like structure and it is known that dynein motor proteins, which connects adjacent microtubule doublet, are the driving force of ciliary motion. However the molecular mechanism to generate motion is still unclear. The authors proved that a pair of microtubules stably linked by DNA-origami and driven by outer dynein arms (ODA) causes beating motion. They employed in vitro motility assay and negative stain TEM to characterize this complex. They demonstrated stable linking of microtubules and ODAs anchored on the both microtubules are essential for oscillatory motion and bending of the microtubules.
Strength
This is an interesting work, …Reviewer #2 (Public Review):
Motile cilia generate rhythmic beating or rotational motion to drive cells or produce extracellular fluid flow. Cilia is made of nine microtubule doublets forming a spoke-like structure and it is known that dynein motor proteins, which connects adjacent microtubule doublet, are the driving force of ciliary motion. However the molecular mechanism to generate motion is still unclear. The authors proved that a pair of microtubules stably linked by DNA-origami and driven by outer dynein arms (ODA) causes beating motion. They employed in vitro motility assay and negative stain TEM to characterize this complex. They demonstrated stable linking of microtubules and ODAs anchored on the both microtubules are essential for oscillatory motion and bending of the microtubules.
Strength
This is an interesting work, addressing an important question in the motile cilia community: what is the minimum system to generate a beating motion? It is an established fact that dynein power stroke on the microtubule doublet is the driving force of the beating motion. It was also known that the radial spoke and the central pair are essential for ciliary motion under the physiological condition, but cilia without radial spokes and the central pair can beat under some special conditions (Yagi and Kamiya, 2000). Therefore in the mechanistic point of view, they are not prerequisite. It is generally thought that fixed connection between adjacent microtubules by nexin converts sliding motion of dyneins to bending, but it was never experimentally investigated. Here the authors successfully enabled a simple system of nexin-like inter-microtubule linkage using DNA origami technique to generate oscillatory and beating motions. This enables an interesting system where ODAs form groups, anchored on two microtubules, orienting oppositely and therefore cause tag-of-war type force generation. The authors demonstrated this system under constraints by DNA origami generates oscillatory and beating motions.
The authors carefully coordinated the experiments to demonstrate oscillations using optical tweezers and sophisticated data analysis (Fourier analysis and a step-finding algorithm). They also proved, using negative stain EM, that this system contains two groups of ODAs forming arrays with opposite polarity on the parallel microtubules.
The manuscript is carefully organized with impressive movies. Geometrical and motility analyses of individual ODAs used for statistics are provided in the supplementary source files. They appropriately cited similar past works from Kamiya and Shingyoji groups (they employed systems closer to the physiological axoneme to reproduce beating) and clarify the differences from this study.Weakness
The authors claim this system mimics two pairs of doublets at the opposite sites from 9+2 cilia structure by having two groups of ODAs between two microtubules facing opposite directions within the pair. It is not exactly the case. In the real axoneme, ODA makes continuous array along the entire length of doublets, which means at any point there are ODAs facing opposite directions. In their system, opposite ODAs cannot exist at the same point (therefore the scheme of Dynein-MT complex of Fig.1B is slightly misleading). If they want to project their result to the ciliary beating model, more insight/explanation would be necessary. For example, arrays of dyneins at certain positions within the long array along one doublet are activated and generate force, while dyneins at different positions are activated on another doublet at the opposite site of the axoneme. This makes the distribution of dyneins and their orientations similar to the system described in this work. Such a localized activation, shown in physiological cilia by Ishikawa and Nicastro groups, may require other regulatory proteins.
They attempted to reveal conformational change of ODAs induced by power stroke using negative stain EM images, which is less convincing compared to the past cryo-ET works (Ishikawa, Nicastro, Pigino groups) and negative stain EM of sea urchin outer dyneins (Hirose group), where the tail and head parts were clearly defined from the 3D map or 2D averages of two-dynein ODAs. Probably three heavy chains and associated proteins hinder detailed visualization of the tail structure. Because of this, Fig.2C is not clear enough to prove conformational change of ODA. This reviewer imagines refined subaverage (probably with larger datasets) is necessary. It is not clear, from the inset of Fig.2 supplement3, how to define the end of the tail for the length measurement, which is the basis for the authors to claim conformational change (Line263-265). The appearance of the tail would be altered, seen from even slightly different view angles. Comparison with 2D projection from apo- and nucleotide-bound 3-headed ODA structures from EM databank will help.In Fig.5B (where the oscillation occurs), the microtubule was once driven >150nm unidirectionally and went back to the original position, before oscillation starts. Is it always the case that relatively long unidirectional motion and return precede oscillation? In Fig.7B, where the authors claim no oscillation happened, only one unidirectional motion was shown. Did oscillation not happen after MT returned to the original position?
Line284-290: More characterization of bending motion will be necessary (and should be possible). How high frequency is it? Do they confirm that other systems (either without DNA-origami or without ODAs arraying oppositely) cannot generate repetitive beating?
-

Reviewer #3 (Public Review):
The authors goal is to create an in vitro analogue of the in vivo activation-inhibition cycle of flagellar dynein, in order to study its mechanism. In their new model system, two sets of outer arm dyneins link a pair of parallel, taxol-stabilised microtubules and (upon photolysis of caged ATP) slide the microtubules back and forth, switching direction every 40 milliseconds, during which time the microtubules slide a few tens of nanometers. This oscillation depends on the sliding of the two microtubules being restricted by DNA origami cross linkers. At any one time, only one set of dyneins, attached to only one microtubule, appears active. The switch point appears defined by the stretching out of the DNA origami crosslinks.
The authors have studied the structure of their system using electron microscopy and …
Reviewer #3 (Public Review):
The authors goal is to create an in vitro analogue of the in vivo activation-inhibition cycle of flagellar dynein, in order to study its mechanism. In their new model system, two sets of outer arm dyneins link a pair of parallel, taxol-stabilised microtubules and (upon photolysis of caged ATP) slide the microtubules back and forth, switching direction every 40 milliseconds, during which time the microtubules slide a few tens of nanometers. This oscillation depends on the sliding of the two microtubules being restricted by DNA origami cross linkers. At any one time, only one set of dyneins, attached to only one microtubule, appears active. The switch point appears defined by the stretching out of the DNA origami crosslinks.
The authors have studied the structure of their system using electron microscopy and studied its mechanics by optical trapping. By harnessing the intrinsic tendency of outer arm dynein to assemble in patches on microtubules and to link microtubules in parallel, they have recapitulated the in vivo arrangement of outer arm dynein, thereby allowing it to be studied in isolation but in an in vivo-like geometry. By adding in a designed DNA origami as a further crosslinker, they convert the system to an oscillator. To my mind this is a very revealing result. The (relative) structural simplicity allows electron microscopy to be used to inspect dynein conformation. The optical trapping has to be done in a relatively weak/compliant trap, which makes for noisy data, but the data are nonetheless revealing, with single molecular steps discernible at some points, and forces and timings accurately determined.
I think the authors have achieved their aims and that their results, subject to minor clarifications of interpretation, support their conclusions.
This is an exciting advance that will stimulate more use of simplified systems to dissect the beating mechanism of flagella. It seems plausible that there are several regulatory mechanisms for beating at the molecular level that overlie one another. Establishing the mechanical properties of dynein in a more lifelike yet less complex structural context is evidently a good way to go - this will hopefully encourage more work along the same lines.
-